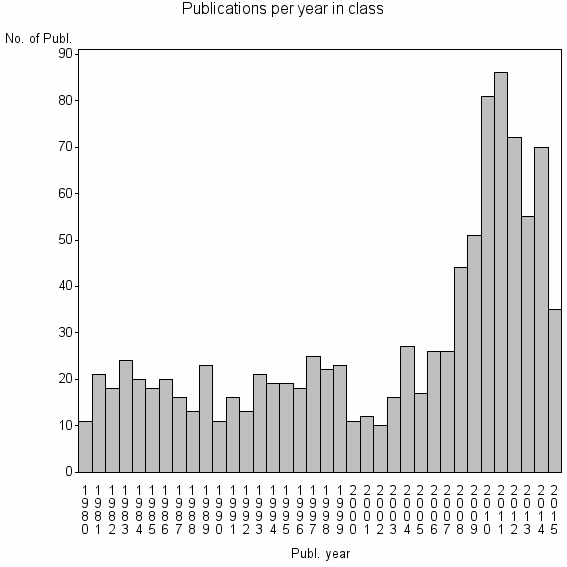
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 10294 | 1010 | 45.4 | 84% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | NUCLEOSOME POSITIONING | Author keyword | 49 | 46% | 8% | 80 |
| 2 | NUCLEOSOME | Author keyword | 9 | 11% | 8% | 76 |
| 3 | NUCLEOSOME MAPPING | Author keyword | 8 | 75% | 1% | 6 |
| 4 | NUCLEOSOME POSITION | Author keyword | 8 | 75% | 1% | 6 |
| 5 | MNASE SEQ | Author keyword | 6 | 80% | 0% | 4 |
| 6 | CHROMATIN CODE | Author keyword | 6 | 100% | 0% | 4 |
| 7 | NUCLEOSOME DNA | Author keyword | 6 | 100% | 0% | 4 |
| 8 | STRONG NUCLEOSOME | Author keyword | 6 | 100% | 0% | 4 |
| 9 | GENOME ERS | Address | 5 | 16% | 3% | 26 |
| 10 | FDN IST PASTEUR | Address | 4 | 38% | 1% | 9 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | NUCLEOSOME POSITIONING | 49 | 46% | 8% | 80 | Search NUCLEOSOME+POSITIONING | Search NUCLEOSOME+POSITIONING |
| 2 | NUCLEOSOME | 9 | 11% | 8% | 76 | Search NUCLEOSOME | Search NUCLEOSOME |
| 3 | NUCLEOSOME MAPPING | 8 | 75% | 1% | 6 | Search NUCLEOSOME+MAPPING | Search NUCLEOSOME+MAPPING |
| 4 | NUCLEOSOME POSITION | 8 | 75% | 1% | 6 | Search NUCLEOSOME+POSITION | Search NUCLEOSOME+POSITION |
| 5 | MNASE SEQ | 6 | 80% | 0% | 4 | Search MNASE+SEQ | Search MNASE+SEQ |
| 6 | CHROMATIN CODE | 6 | 100% | 0% | 4 | Search CHROMATIN+CODE | Search CHROMATIN+CODE |
| 7 | NUCLEOSOME DNA | 6 | 100% | 0% | 4 | Search NUCLEOSOME+DNA | Search NUCLEOSOME+DNA |
| 8 | STRONG NUCLEOSOME | 6 | 100% | 0% | 4 | Search STRONG+NUCLEOSOME | Search STRONG+NUCLEOSOME |
| 9 | HISTONE DNA INTERACTION | 4 | 75% | 0% | 3 | Search HISTONE+DNA+INTERACTION | Search HISTONE+DNA+INTERACTION |
| 10 | BENDABILITY MATRIX | 3 | 100% | 0% | 3 | Search BENDABILITY+MATRIX | Search BENDABILITY+MATRIX |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | EUKARYOTIC GENOME | 34 | 35% | 8% | 79 |
| 2 | HISTONE DNA INTERACTIONS | 32 | 60% | 3% | 35 |
| 3 | MICROCOCCAL NUCLEASE | 21 | 35% | 5% | 49 |
| 4 | DIFFERENTIAL COFACTOR REQUIREMENTS | 18 | 89% | 1% | 8 |
| 5 | POSITIONS IN VIVO | 18 | 89% | 1% | 8 |
| 6 | PHO5 PROMOTER | 15 | 28% | 4% | 45 |
| 7 | POSITIONED NUCLEOSOMES | 12 | 26% | 4% | 41 |
| 8 | GENOMIC CODE | 12 | 86% | 1% | 6 |
| 9 | YEAST PHO5 PROMOTER | 11 | 41% | 2% | 21 |
| 10 | PREFERENTIAL ACCESSIBILITY | 11 | 67% | 1% | 10 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Determinants of nucleosome positioning | 2013 | 96 | 68 | 76% |
| Nucleosome positioning in yeasts: methods, maps, and mechanisms | 2015 | 1 | 172 | 70% |
| Gene regulation by nucleosome positioning | 2010 | 64 | 82 | 62% |
| Mechanisms Underlying Nucleosome Positioning In Vivo | 2014 | 7 | 120 | 53% |
| Nucleosome positioning: How is it established, and why does it matter? | 2010 | 80 | 99 | 46% |
| Nucleosome positioning and gene regulation: advances through genomics | 2009 | 353 | 126 | 34% |
| Nucleosome positioning: bringing order to the eukaryotic genome | 2012 | 21 | 65 | 62% |
| Genome-wide "Re"-Modeling of Nucleosome Positions | 2011 | 31 | 21 | 67% |
| The logic of chromatin architecture and remodelling at promoters | 2009 | 209 | 62 | 42% |
| What controls nucleosome positions? | 2009 | 181 | 91 | 46% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | GENOME ERS | 5 | 16% | 2.6% | 26 |
| 2 | FDN IST PASTEUR | 4 | 38% | 0.9% | 9 |
| 3 | CAGT PL GENOTYPING | 3 | 100% | 0.3% | 3 |
| 4 | FDN PIERRE GILLES GENNES RECH | 1 | 50% | 0.2% | 2 |
| 5 | ALTHOUSE 308 | 1 | 40% | 0.2% | 2 |
| 6 | THEORET GENET | 1 | 21% | 0.4% | 4 |
| 7 | STUDIO ACIDI NUCL | 1 | 17% | 0.5% | 5 |
| 8 | BEREICH GENET | 1 | 19% | 0.4% | 4 |
| 9 | AGR BIOINFORMAT UNIT | 1 | 10% | 0.7% | 7 |
| 10 | PROGRAM CANC GENET EPIGENET TUMOR VIROL | 1 | 21% | 0.3% | 3 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000202522 | LINKER HISTONE//HISTONE H1//NUCLEOSOME |
| 2 | 0.0000181678 | SWI SNF//BRG1//ISWI |
| 3 | 0.0000159400 | P I TRANSPORTER//PHO81//PHO89 |
| 4 | 0.0000146453 | NUCLEAR FACTOR 1//NUCLEAR FACTOR I//DEV GENOM GRP |
| 5 | 0.0000144658 | HISTONE CHAPERONE//ASF1//CAF 1 |
| 6 | 0.0000136719 | CHIP SEQ//PEAK CALLING//DNASE SEQ |
| 7 | 0.0000136053 | DNA CURVATURE//A TRACT//DNA FLEXIBILITY |
| 8 | 0.0000109835 | SIR COMPLEX//SIR3//SIR PROTEINS |
| 9 | 0.0000089473 | GAL4P//GAL80//GAL GENES |
| 10 | 0.0000076512 | ANTI THY 12 ANTIBODY//CELL TYPE IDENTITY//ENHANCER CORE |