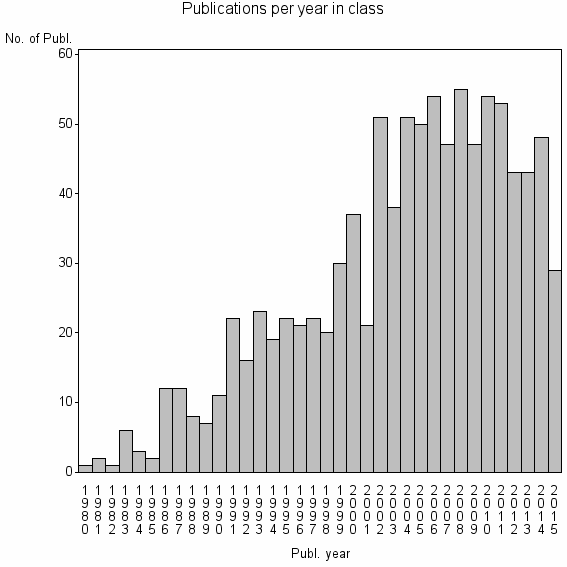
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 10621 | 981 | 43.0 | 84% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 238 | 19563 | BIOINFORMATICS//BMC BIOINFORMATICS//GENOME RESEARCH |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | ISOCHORES | Author keyword | 66 | 60% | 7% | 72 |
| 2 | EVOLUZ MOL | Address | 26 | 59% | 3% | 29 |
| 3 | ISOCHORE | Author keyword | 23 | 47% | 4% | 36 |
| 4 | BIASED GENE CONVERSION | Author keyword | 22 | 62% | 2% | 23 |
| 5 | GC BIASED GENE CONVERSION | Author keyword | 13 | 69% | 1% | 11 |
| 6 | MALE DRIVEN EVOLUTION | Author keyword | 13 | 71% | 1% | 10 |
| 7 | MALE MUTATION BIAS | Author keyword | 11 | 67% | 1% | 10 |
| 8 | GC CONTENT | Author keyword | 9 | 16% | 5% | 50 |
| 9 | BASE COMPOSITION | Author keyword | 7 | 20% | 3% | 30 |
| 10 | CHROMOSOMAL BANDS | Author keyword | 6 | 80% | 0% | 4 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | ISOCHORES | 66 | 60% | 7% | 72 | Search ISOCHORES | Search ISOCHORES |
| 2 | ISOCHORE | 23 | 47% | 4% | 36 | Search ISOCHORE | Search ISOCHORE |
| 3 | BIASED GENE CONVERSION | 22 | 62% | 2% | 23 | Search BIASED+GENE+CONVERSION | Search BIASED+GENE+CONVERSION |
| 4 | GC BIASED GENE CONVERSION | 13 | 69% | 1% | 11 | Search GC+BIASED+GENE+CONVERSION | Search GC+BIASED+GENE+CONVERSION |
| 5 | MALE DRIVEN EVOLUTION | 13 | 71% | 1% | 10 | Search MALE+DRIVEN+EVOLUTION | Search MALE+DRIVEN+EVOLUTION |
| 6 | MALE MUTATION BIAS | 11 | 67% | 1% | 10 | Search MALE+MUTATION+BIAS | Search MALE+MUTATION+BIAS |
| 7 | GC CONTENT | 9 | 16% | 5% | 50 | Search GC+CONTENT | Search GC+CONTENT |
| 8 | BASE COMPOSITION | 7 | 20% | 3% | 30 | Search BASE+COMPOSITION | Search BASE+COMPOSITION |
| 9 | CHROMOSOMAL BANDS | 6 | 80% | 0% | 4 | Search CHROMOSOMAL+BANDS | Search CHROMOSOMAL+BANDS |
| 10 | COMPOSITIONAL HOMOGENEITY | 6 | 58% | 1% | 7 | Search COMPOSITIONAL+HOMOGENEITY | Search COMPOSITIONAL+HOMOGENEITY |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ISOCHORES | 76 | 56% | 9% | 92 |
| 2 | COLD BLOODED VERTEBRATES | 42 | 70% | 4% | 35 |
| 3 | BIASED GENE CONVERSION | 29 | 42% | 5% | 53 |
| 4 | GC CONTENT | 29 | 28% | 9% | 88 |
| 5 | PRDM9 | 21 | 54% | 3% | 27 |
| 6 | GENE DISTRIBUTION | 20 | 62% | 2% | 21 |
| 7 | LOCAL SIMILARITY | 19 | 63% | 2% | 19 |
| 8 | RICH ISOCHORES | 18 | 89% | 1% | 8 |
| 9 | COMPOSITIONAL PATTERNS | 17 | 79% | 1% | 11 |
| 10 | WARM BLOODED VERTEBRATES | 17 | 75% | 1% | 12 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Biased Gene Conversion and the Evolution of Mammalian Genomic Landscapes | 2009 | 194 | 135 | 61% |
| Adaptation or biased gene conversion? Extending the null hypothesis of molecular evolution | 2007 | 105 | 26 | 73% |
| Isochores and the evolutionary genomics of vertebrates | 2000 | 332 | 77 | 73% |
| Variation in the mutation rate across mammalian genomes | 2011 | 78 | 108 | 41% |
| The neoselectionist theory of genome evolution | 2007 | 51 | 68 | 56% |
| What are the genomic drivers of the rapid evolution of PRDM9? | 2011 | 30 | 35 | 51% |
| Mammalian meiotic recombination hot spots | 2007 | 52 | 172 | 52% |
| Characteristics, causes and evolutionary consequences of male-biased mutation | 2007 | 73 | 101 | 39% |
| An evolutionary view of human recombination | 2007 | 102 | 122 | 30% |
| Direct and indirect consequences of meiotic recombination: implications for genome evolution | 2012 | 27 | 98 | 43% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | EVOLUZ MOL | 26 | 59% | 3.0% | 29 |
| 2 | DIPARTIMENTO BIOL ANIM M LA GRECA | 1 | 38% | 0.3% | 3 |
| 3 | EVOL MOL | 1 | 100% | 0.2% | 2 |
| 4 | MATH METHODS BIOL | 1 | 30% | 0.3% | 3 |
| 5 | COURSE MED SCI TECHNOL | 1 | 40% | 0.2% | 2 |
| 6 | SECC BIOMATEMAT | 1 | 15% | 0.6% | 6 |
| 7 | POPULAT CONSERVAT BIOL | 1 | 33% | 0.2% | 2 |
| 8 | ANIM EVOLUT PHYSIOL | 1 | 50% | 0.1% | 1 |
| 9 | BASIN PROGRAM | 1 | 50% | 0.1% | 1 |
| 10 | BIOL MOL OKOL | 1 | 50% | 0.1% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000164836 | TRANSLATIONAL SELECTION//CODON USAGE//SYNONYMOUS CODON USAGE |
| 2 | 0.0000129644 | MCDONALD KREITMAN TEST//COALESCENT//SELECTIVE SWEEPS |
| 3 | 0.0000125254 | K MAXIMUM SUMS PROBLEM//MAXIMUM SUM PROBLEM//MAXIMUM SUM SEGMENT |
| 4 | 0.0000117623 | ASCOBOLUS//EVOLUTION OF PLOIDY//MULTIGENE ARRAYS |
| 5 | 0.0000111619 | SYNAPTONEMAL COMPLEX//MEIOSIS//SYNAPSIS |
| 6 | 0.0000096850 | IROQUOIS//CONSERVED NONCODING ELEMENTS//ENERGY ENVIRONM BIOL COMP |
| 7 | 0.0000089680 | LATENT PERIODICITY//3 BASE PERIODICITY//DNA WALK |
| 8 | 0.0000086719 | MOLECULAR CLOCK//RELAXED CLOCK//PHYLODYNAMICS |
| 9 | 0.0000078759 | BIDIRECTIONAL PROMOTER//NEIGHBORING GENES//GENOME NETWORK PROJECT CORE GRPGENOM SCI |
| 10 | 0.0000077339 | PROCESSED PSEUDOGENES//PHYSICAL AND PSYCHOLOGICAL STRESSES//DUPLICATED PSEUDOGENES |