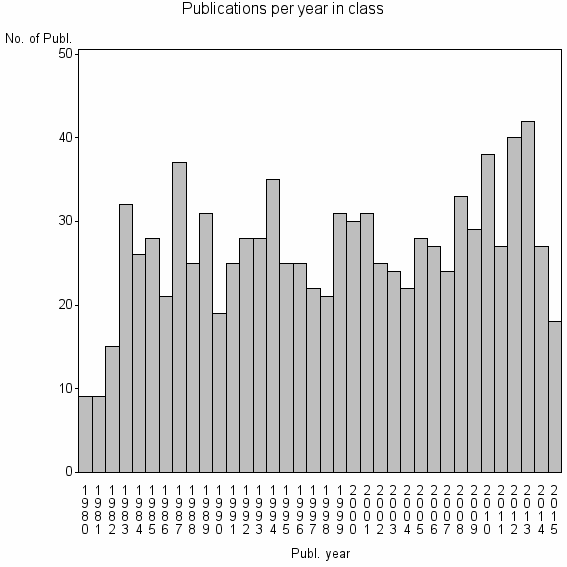
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 10905 | 957 | 37.4 | 71% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 2974 | 2151 | ARGININE METHYLATION//CARM1//ISOASPARTATE |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | ISOASPARTATE | Author keyword | 50 | 72% | 4% | 39 |
| 2 | PROTEIN L ISOASPARTYL METHYLTRANSFERASE | Author keyword | 38 | 89% | 2% | 17 |
| 3 | DEAMIDATION | Author keyword | 30 | 27% | 10% | 96 |
| 4 | BETA ASPARTIC ACID | Author keyword | 24 | 91% | 1% | 10 |
| 5 | PROTEIN REPAIR | Author keyword | 20 | 52% | 3% | 27 |
| 6 | ASPARAGINE DEAMIDATION | Author keyword | 18 | 83% | 1% | 10 |
| 7 | ISOASPARTIC ACID | Author keyword | 18 | 89% | 1% | 8 |
| 8 | ISOASPARTYL | Author keyword | 14 | 100% | 1% | 7 |
| 9 | PIMT | Author keyword | 9 | 64% | 1% | 9 |
| 10 | ASPARAGINYL | Author keyword | 9 | 83% | 1% | 5 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | ISOASPARTATE | 50 | 72% | 4% | 39 | Search ISOASPARTATE | Search ISOASPARTATE |
| 2 | PROTEIN L ISOASPARTYL METHYLTRANSFERASE | 38 | 89% | 2% | 17 | Search PROTEIN+L+ISOASPARTYL+METHYLTRANSFERASE | Search PROTEIN+L+ISOASPARTYL+METHYLTRANSFERASE |
| 3 | DEAMIDATION | 30 | 27% | 10% | 96 | Search DEAMIDATION | Search DEAMIDATION |
| 4 | BETA ASPARTIC ACID | 24 | 91% | 1% | 10 | Search BETA+ASPARTIC+ACID | Search BETA+ASPARTIC+ACID |
| 5 | PROTEIN REPAIR | 20 | 52% | 3% | 27 | Search PROTEIN+REPAIR | Search PROTEIN+REPAIR |
| 6 | ASPARAGINE DEAMIDATION | 18 | 83% | 1% | 10 | Search ASPARAGINE+DEAMIDATION | Search ASPARAGINE+DEAMIDATION |
| 7 | ISOASPARTIC ACID | 18 | 89% | 1% | 8 | Search ISOASPARTIC+ACID | Search ISOASPARTIC+ACID |
| 8 | ISOASPARTYL | 14 | 100% | 1% | 7 | Search ISOASPARTYL | Search ISOASPARTYL |
| 9 | PIMT | 9 | 64% | 1% | 9 | Search PIMT | Search PIMT |
| 10 | ASPARAGINYL | 9 | 83% | 1% | 5 | Search ASPARAGINYL | Search ASPARAGINYL |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ASPARTYL RESIDUES | 184 | 83% | 11% | 105 |
| 2 | CARBOXYL METHYLTRANSFERASE | 73 | 52% | 10% | 99 |
| 3 | SPONTANEOUS DEGRADATION | 70 | 82% | 4% | 41 |
| 4 | DEAMIDATION | 67 | 30% | 20% | 189 |
| 5 | ASPARAGINYL RESIDUES | 59 | 63% | 6% | 60 |
| 6 | CHEMICAL PATHWAYS | 58 | 63% | 6% | 59 |
| 7 | ASPARAGINYL | 51 | 49% | 8% | 75 |
| 8 | SUCCINIMIDE FORMATION | 47 | 60% | 5% | 51 |
| 9 | PROTEIN CARBOXYL METHYLTRANSFERASE | 46 | 70% | 4% | 38 |
| 10 | ASPARTYL | 46 | 53% | 6% | 61 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Mass spectrometric analysis of asparagine deamidation and aspartate isomerization in polypeptides | 2010 | 34 | 45 | 73% |
| Aging as war between chemical and biochemical processes: Protein methylation and the recognition of age-damaged proteins for repair | 2003 | 153 | 83 | 86% |
| Deamidation and isoaspartate formation in proteins: unwanted alterations or surreptitious signals? | 2003 | 127 | 82 | 72% |
| The Role of Protein L-Isoaspartyl/D-Aspartyl O-Methyltransferase (PIMT) in Intracellular Signal Transduction | 2010 | 14 | 42 | 83% |
| Formulation considerations for proteins susceptible to asparagine deamidation and aspartate isomerization | 2006 | 85 | 70 | 61% |
| Secondary structure and protein deamidation | 1999 | 67 | 21 | 81% |
| Protein Repair L-Isoaspartyl Methyltransferase1 Is Involved in Both Seed Longevity and Germination Vigor in Arabidopsis | 2008 | 52 | 80 | 45% |
| Collapse of Homochirality of Amino Acids in Proteins from Various Tissues during Aging | 2010 | 6 | 25 | 88% |
| Racemization of aspartic acid in human proteins | 2002 | 64 | 104 | 70% |
| Early Engineering Approaches to Improve Peptide Developability and Manufacturability | 2015 | 1 | 67 | 13% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | OPEN PROT SCI | 2 | 67% | 0.2% | 2 |
| 2 | BION DESIGN | 2 | 43% | 0.3% | 3 |
| 3 | RADIAT LIFE SCI RADIAT MED SCI | 2 | 33% | 0.4% | 4 |
| 4 | AGR BIOTECHNOL UNDER PROGRAM | 1 | 100% | 0.2% | 2 |
| 5 | AGR SCI N220C | 1 | 100% | 0.2% | 2 |
| 6 | LIFE SCI NAT OURCES GENET ENGN | 1 | 100% | 0.2% | 2 |
| 7 | SEED BIOL GRP | 1 | 30% | 0.3% | 3 |
| 8 | ANALYT SCI IMMUNOL DIS | 1 | 50% | 0.1% | 1 |
| 9 | CELL BIOPHYS UNIT | 1 | 50% | 0.1% | 1 |
| 10 | CNRSU382 | 1 | 50% | 0.1% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000103133 | AGALACTOSYL IGG//PROT ANALYT CHEM//IGG GLYCOSYLATION |
| 2 | 0.0000070169 | PROTEIN FORMULATION//SUBVISIBLE PARTICLES//PROTEIN AGGREGATION |
| 3 | 0.0000065990 | FAMILIAL DANISH DEMENTIA//BRI2//GLUTAMINYL CYCLASE |
| 4 | 0.0000053808 | D AMINO ACID OXIDASE//D SERINE//SERINE RACEMASE |
| 5 | 0.0000053798 | TRIOSEPHOSPHATE ISOMERASE//TRIOSEPHOSPHATE ISOMERASE DEFICIENCY//TRIOSEPHOSPHATE ISOMERASE TIM |
| 6 | 0.0000046990 | ARGININE METHYLATION//CARM1//PROTEIN ARGININE METHYLTRANSFERASE |
| 7 | 0.0000046562 | ALPHA CRYSTALLIN//SMALL HEAT SHOCK PROTEIN//ALPHA B CRYSTALLIN |
| 8 | 0.0000045308 | FORENSIC AGE ESTIMATION//AGE ESTIMATION//AGE DETERMINATION BY TEETH |
| 9 | 0.0000039614 | ASPARTYLGLUCOSAMINURIA//GLYCOSYLASPARAGINASE//ASPARTYLGLUCOSAMINIDASE |
| 10 | 0.0000035663 | LYSINOALANINE//CROSS LINKED AMINO ACIDS//HISTIDINOALANINE |