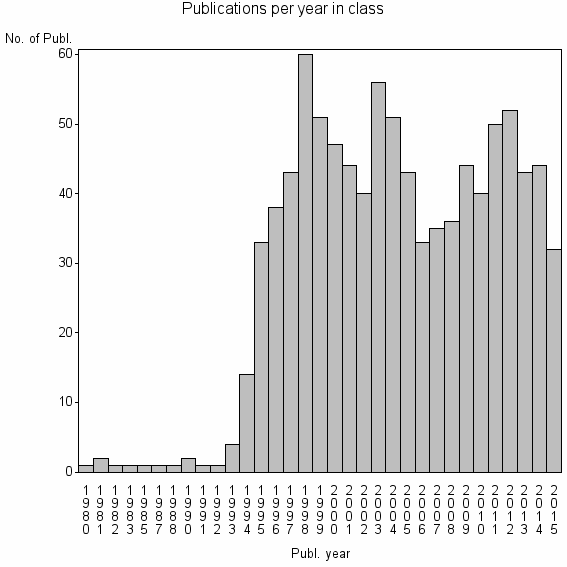
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 11057 | 945 | 52.7 | 90% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 302 | 17871 | TRANSLESION SYNTHESIS//DNA POLYMERASE//RECA PROTEIN |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | FEN1 | Author keyword | 21 | 55% | 3% | 26 |
| 2 | BRAUN S | Address | 18 | 59% | 2% | 20 |
| 3 | CREAT INITIAT CELL CYCLE CONTROL | Address | 17 | 72% | 1% | 13 |
| 4 | DNA REPLICAT GENOME ABIL | Address | 11 | 78% | 1% | 7 |
| 5 | OKAZAKI FRAGMENT PROCESSING | Author keyword | 11 | 78% | 1% | 7 |
| 6 | RAD27 | Author keyword | 11 | 78% | 1% | 7 |
| 7 | FEN 1 | Author keyword | 10 | 63% | 1% | 10 |
| 8 | FLAP ENDONUCLEASE | Author keyword | 10 | 63% | 1% | 10 |
| 9 | DNA2 | Author keyword | 10 | 58% | 1% | 11 |
| 10 | FLAP ENDONUCLEASE 1 | Author keyword | 7 | 53% | 1% | 10 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | FEN1 | 21 | 55% | 3% | 26 | Search FEN1 | Search FEN1 |
| 2 | OKAZAKI FRAGMENT PROCESSING | 11 | 78% | 1% | 7 | Search OKAZAKI+FRAGMENT+PROCESSING | Search OKAZAKI+FRAGMENT+PROCESSING |
| 3 | RAD27 | 11 | 78% | 1% | 7 | Search RAD27 | Search RAD27 |
| 4 | FEN 1 | 10 | 63% | 1% | 10 | Search FEN+1 | Search FEN+1 |
| 5 | FLAP ENDONUCLEASE | 10 | 63% | 1% | 10 | Search FLAP+ENDONUCLEASE | Search FLAP+ENDONUCLEASE |
| 6 | DNA2 | 10 | 58% | 1% | 11 | Search DNA2 | Search DNA2 |
| 7 | FLAP ENDONUCLEASE 1 | 7 | 53% | 1% | 10 | Search FLAP+ENDONUCLEASE+1 | Search FLAP+ENDONUCLEASE+1 |
| 8 | OKAZAKI FRAGMENT MATURATION | 7 | 67% | 1% | 6 | Search OKAZAKI+FRAGMENT+MATURATION | Search OKAZAKI+FRAGMENT+MATURATION |
| 9 | MUTS BETA | 6 | 80% | 0% | 4 | Search MUTS+BETA | Search MUTS+BETA |
| 10 | TRIPLET REPEATS | 5 | 24% | 2% | 18 | Search TRIPLET+REPEATS | Search TRIPLET+REPEATS |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | TRIPLET REPEATS | 56 | 43% | 11% | 100 |
| 2 | FEN 1 | 46 | 70% | 4% | 39 |
| 3 | HEREDITARY DISEASE GENES | 38 | 89% | 2% | 17 |
| 4 | ORIENTATION DEPENDENT MANNER | 38 | 93% | 1% | 14 |
| 5 | FLAP ENDONUCLEASE 1 | 37 | 35% | 9% | 87 |
| 6 | ESSENTIAL IN VIVO | 35 | 72% | 3% | 28 |
| 7 | CENTER DOT CAG | 35 | 86% | 2% | 18 |
| 8 | TRINUCLEOTIDE REPEATS | 35 | 13% | 26% | 245 |
| 9 | CALF RTH 1 NUCLEASE | 34 | 93% | 1% | 13 |
| 10 | HUMAN FLAP ENDONUCLEASE 1 | 34 | 63% | 4% | 34 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Repeat instability: Mechanisms of dynamic mutations | 2005 | 380 | 153 | 52% |
| Expandable DNA repeats and human disease | 2007 | 290 | 91 | 53% |
| Repeat instability during DNA repair: Insights from model systems | 2015 | 1 | 295 | 55% |
| Repeat instability as the basis for human diseases and as a potential target for therapy | 2010 | 99 | 48 | 67% |
| Flap endonuclease 1: A central component of DNA metabolism | 2004 | 205 | 177 | 65% |
| Flap Endonuclease 1 | 2013 | 24 | 118 | 59% |
| Mechanisms of trinucleotide repeat instability during human development | 2010 | 124 | 119 | 46% |
| DNA base excision repair: a mechanism of trinucleotide repeat expansion | 2012 | 26 | 70 | 70% |
| Polymerase Dynamics at the Eukaryotic DNA Replication Fork | 2009 | 156 | 37 | 43% |
| Okazaki fragment maturation: nucleases take centre stage | 2011 | 43 | 66 | 65% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BRAUN S | 18 | 59% | 2.1% | 20 |
| 2 | CREAT INITIAT CELL CYCLE CONTROL | 17 | 72% | 1.4% | 13 |
| 3 | DNA REPLICAT GENOME ABIL | 11 | 78% | 0.7% | 7 |
| 4 | SECT GENE STRUCT DIS | 3 | 39% | 0.7% | 7 |
| 5 | DNA STRUCT MUTAGENESIS | 3 | 57% | 0.4% | 4 |
| 6 | GENOME DNA STRUCT MUTAGENESIS | 3 | 57% | 0.4% | 4 |
| 7 | GENOME BIOSCI TECHNOL | 3 | 50% | 0.4% | 4 |
| 8 | PROGRAMME TRANSLAT MED NEUROGENET | 2 | 67% | 0.2% | 2 |
| 9 | SAMSUNG BIOMED CHANGAN KU | 2 | 67% | 0.2% | 2 |
| 10 | BRAUN S 147 75 | 2 | 36% | 0.4% | 4 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000150428 | EMANUEL SYNDROME//PARTIAL TRISOMY 11Q//DUPLICATION 11Q |
| 2 | 0.0000113839 | MICROSATELLITE EVOLUTION//COMPOUND MICROSATELLITE//STEPWISE MUTATION MODEL |
| 3 | 0.0000109786 | MUTS//MISMATCH REPAIR//DNA MISMATCH REPAIR |
| 4 | 0.0000101949 | CLAMP LOADER//REPLICATION FACTOR C//DNA REPLICAT |
| 5 | 0.0000098722 | FRIEDREICH ATAXIA//FRIEDREICHS ATAXIA//FRATAXIN |
| 6 | 0.0000097736 | MYOTONIC DYSTROPHY//MYOTONIC DYSTROPHY TYPE 1//MYOTONIC DYSTROPHY TYPE 2 |
| 7 | 0.0000084187 | AICARDI GOUTIERES SYNDROME//TREX1//RIBONUCLEASE H |
| 8 | 0.0000083347 | APE1//AP ENDONUCLEASE//APE1 REF 1 |
| 9 | 0.0000074732 | DNA POLYMERASE//DNA POLYMERASE BETA//DNA POLYMERASE LAMBDA |
| 10 | 0.0000070427 | DNA POLYMERASE ALPHA//DNA SYNTHESOME//DNA POLYMERASE DELTA |