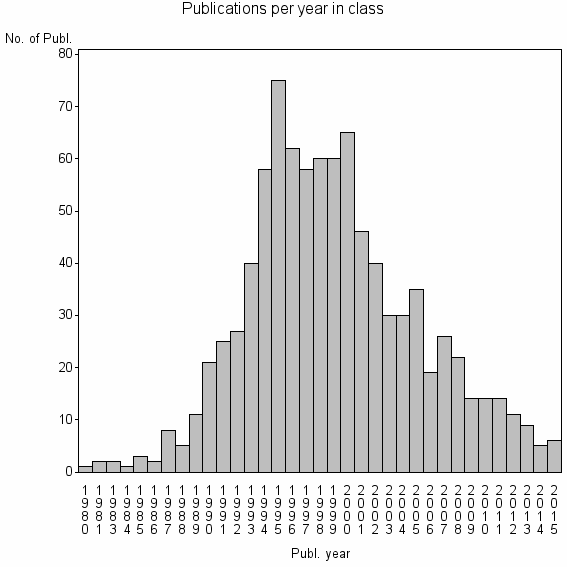
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 11546 | 907 | 37.8 | 87% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 3331 | 1286 | CHROMATIN SUPRAORGANIZATION//ZNF703//NUCLEAR PHENOTYPES |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | CLIN BREAST CARE PROJECT | Address | 5 | 24% | 2% | 17 |
| 2 | 16Q243 | Author keyword | 4 | 75% | 0% | 3 |
| 3 | CHROMOSOMAL SCANNING | Author keyword | 3 | 100% | 0% | 3 |
| 4 | HSRBC | Author keyword | 3 | 100% | 0% | 3 |
| 5 | LOH11CR2A | Author keyword | 3 | 100% | 0% | 3 |
| 6 | EI24 | Author keyword | 3 | 60% | 0% | 3 |
| 7 | LEA135 | Author keyword | 2 | 67% | 0% | 2 |
| 8 | EQUIPE GENOME CANC | Address | 2 | 40% | 0% | 4 |
| 9 | P ANIKOLAOU | Address | 2 | 40% | 0% | 4 |
| 10 | CHROMOSOME 11Q | Author keyword | 1 | 21% | 1% | 6 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | 16Q243 | 4 | 75% | 0% | 3 | Search 16Q243 | Search 16Q243 |
| 2 | CHROMOSOMAL SCANNING | 3 | 100% | 0% | 3 | Search CHROMOSOMAL+SCANNING | Search CHROMOSOMAL+SCANNING |
| 3 | HSRBC | 3 | 100% | 0% | 3 | Search HSRBC | Search HSRBC |
| 4 | LOH11CR2A | 3 | 100% | 0% | 3 | Search LOH11CR2A | Search LOH11CR2A |
| 5 | EI24 | 3 | 60% | 0% | 3 | Search EI24 | Search EI24 |
| 6 | LEA135 | 2 | 67% | 0% | 2 | Search LEA135 | Search LEA135 |
| 7 | CHROMOSOME 11Q | 1 | 21% | 1% | 6 | Search CHROMOSOME+11Q | Search CHROMOSOME+11Q |
| 8 | BLID | 1 | 100% | 0% | 2 | Search BLID | Search BLID |
| 9 | BREAST BENIGN TUMOR | 1 | 100% | 0% | 2 | Search BREAST+BENIGN+TUMOR | Search BREAST+BENIGN+TUMOR |
| 10 | LEA 135 | 1 | 100% | 0% | 2 | Search LEA+135 | Search LEA+135 |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | 2 CM INTERVAL | 9 | 83% | 1% | 5 |
| 2 | ALLELIC IMBALANCE | 7 | 16% | 5% | 42 |
| 3 | 16Q243 | 5 | 63% | 1% | 5 |
| 4 | SHORT TERM CULTURE | 5 | 33% | 1% | 12 |
| 5 | CLONAL CHROMOSOME ABNORMALITIES | 4 | 28% | 1% | 11 |
| 6 | KARYOTYPIC PATTERN | 3 | 57% | 0% | 4 |
| 7 | HETEROZYGOSITY REGION | 3 | 43% | 1% | 6 |
| 8 | CHROMOSOME ARM 16Q | 3 | 24% | 1% | 11 |
| 9 | HSRBC | 2 | 67% | 0% | 2 |
| 10 | HOMOGENEOUSLY STAINING REGIONS | 2 | 12% | 1% | 13 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| The molecular genetics of breast cancer: The contribution of comparative genomic hybridization | 2005 | 57 | 101 | 41% |
| Genomic analysis: Toward a new approach in breast cancer management | 2012 | 6 | 130 | 29% |
| Cytogenetic clues to breast carcinogenesis | 2002 | 43 | 92 | 51% |
| Chromosome 16 tumor-suppressor genes in breast cancer | 2006 | 43 | 105 | 41% |
| Genome-wide copy number analysis in primary breast cancer | 2012 | 4 | 45 | 33% |
| GENETIC ALTERATIONS IN BREAST-CANCER | 1995 | 242 | 187 | 28% |
| Allelic loss on chromosome band 18p11.3 occurs early and reveals heterogeneity in breast cancer progression | 2001 | 18 | 21 | 67% |
| SOMATIC GENETIC CHANGES IN HUMAN BREAST-CANCER | 1994 | 204 | 159 | 36% |
| Pooled analysis of loss of heterozygosity in breast cancer: a genome scan provides comparative evidence for multiple tumor suppressors and identifies novel candidate regions | 2003 | 69 | 97 | 26% |
| Molecular genetics of breast cancer progression | 1999 | 68 | 144 | 33% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CLIN BREAST CARE PROJECT | 5 | 24% | 1.9% | 17 |
| 2 | EQUIPE GENOME CANC | 2 | 40% | 0.4% | 4 |
| 3 | P ANIKOLAOU | 2 | 40% | 0.4% | 4 |
| 4 | IMMUNOCYTOCHEM MOL PATHOL | 1 | 40% | 0.2% | 2 |
| 5 | OTOLARYNGOL BIOCHEM | 1 | 40% | 0.2% | 2 |
| 6 | CANC GENET EPIGENET PROGRAM | 1 | 23% | 0.3% | 3 |
| 7 | FRE 2584 | 1 | 21% | 0.3% | 3 |
| 8 | BIOMED NE | 1 | 50% | 0.1% | 1 |
| 9 | BREAST CANC S | 1 | 50% | 0.1% | 1 |
| 10 | BREAST SURG PATHOL | 1 | 50% | 0.1% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000229436 | ZNF703//C8ORF4//8P12 |
| 2 | 0.0000214012 | LIFESPAN MED//CYTOGENET FISH GENOTOXICOL//BUCCAL SMEARS |
| 3 | 0.0000157697 | INTER SIMPLE SEQUENCE REPEAT PCR//BIOPHYS CYTOMETRY//HOSP UNIV SALAMANCA IBSAL |
| 4 | 0.0000139186 | CHROMATIN SUPRAORGANIZATION//NUCLEAR PHENOTYPES//HUMAN BREAST EPITHELIAL CELLS |
| 5 | 0.0000124327 | OVARIAN CARCINOMAS//CANC MONTREAL//6Q27 |
| 6 | 0.0000124220 | SPECTRAL SIMILARITY MAPPING//FAST FISH//CHROMOSOME SPREADING |
| 7 | 0.0000113644 | DUCTAL CARCINOMA IN SITU//LOBULAR CARCINOMA IN SITU//FLAT EPITHELIAL ATYPIA |
| 8 | 0.0000085760 | LZTS1//UROGENE//NUCLEAR ROUNDNESS |
| 9 | 0.0000083110 | RBM5//HUMAN CHROMOSOME 3//LUCA 15 |
| 10 | 0.0000081356 | CHROMOSOME 3Q26//PAN RAS//HUMAN TELOMERASE GENE |