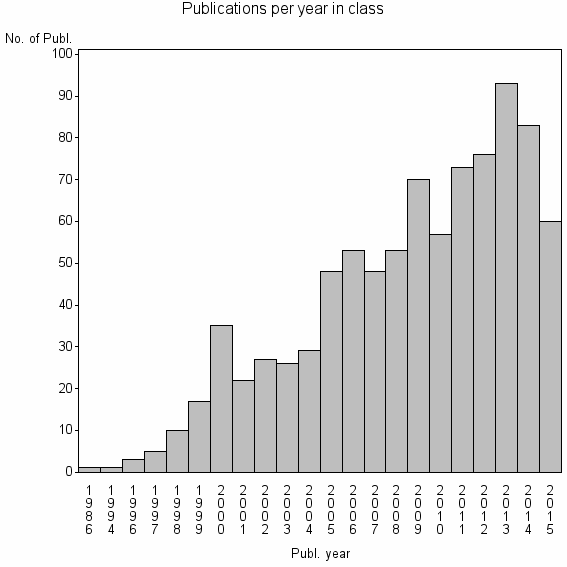
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 11788 | 890 | 41.9 | 86% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 42 | 31225 | PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN STRUCTURE PREDICTION//PROTEIN FOLDING |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | NSSNP | Author keyword | 13 | 62% | 1% | 13 |
| 2 | POLYPHEN | Author keyword | 12 | 63% | 1% | 12 |
| 3 | NSSNPS | Author keyword | 9 | 59% | 1% | 10 |
| 4 | MUTATION DATABASE | Author keyword | 6 | 29% | 2% | 19 |
| 5 | POLYPHEN 2 | Author keyword | 6 | 71% | 1% | 5 |
| 6 | LSDB | Author keyword | 5 | 40% | 1% | 10 |
| 7 | MODELED STRUCTURE | Author keyword | 5 | 63% | 1% | 5 |
| 8 | BRISTOL SYST BIOMED | Address | 4 | 75% | 0% | 3 |
| 9 | HGMD | Author keyword | 4 | 75% | 0% | 3 |
| 10 | BIOTECHNOL CHEM BIOMED ENGN | Address | 3 | 20% | 2% | 15 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | NSSNP | 13 | 62% | 1% | 13 | Search NSSNP | Search NSSNP |
| 2 | POLYPHEN | 12 | 63% | 1% | 12 | Search POLYPHEN | Search POLYPHEN |
| 3 | NSSNPS | 9 | 59% | 1% | 10 | Search NSSNPS | Search NSSNPS |
| 4 | MUTATION DATABASE | 6 | 29% | 2% | 19 | Search MUTATION+DATABASE | Search MUTATION+DATABASE |
| 5 | POLYPHEN 2 | 6 | 71% | 1% | 5 | Search POLYPHEN+2 | Search POLYPHEN+2 |
| 6 | LSDB | 5 | 40% | 1% | 10 | Search LSDB | Search LSDB |
| 7 | MODELED STRUCTURE | 5 | 63% | 1% | 5 | Search MODELED+STRUCTURE | Search MODELED+STRUCTURE |
| 8 | HGMD | 4 | 75% | 0% | 3 | Search HGMD | Search HGMD |
| 9 | FATHMM | 3 | 100% | 0% | 3 | Search FATHMM | Search FATHMM |
| 10 | SNPSGO | 3 | 100% | 0% | 3 | Search SNPSGO | Search SNPSGO |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | NON SYNONYMOUS SNPS | 103 | 77% | 8% | 70 |
| 2 | PROTEIN STABILITY CHANGES | 18 | 52% | 3% | 24 |
| 3 | SYNONYMOUS CODING SNPS | 18 | 89% | 1% | 8 |
| 4 | AFFECT PROTEIN FUNCTION | 16 | 42% | 3% | 30 |
| 5 | MUTATION DATABASES | 14 | 55% | 2% | 17 |
| 6 | GENOTYPING PURPOSES | 12 | 86% | 1% | 6 |
| 7 | STABILITY CHANGES | 11 | 31% | 3% | 30 |
| 8 | PATHOLOGICAL MUTATIONS | 11 | 69% | 1% | 9 |
| 9 | HIV 1 PROTEASE MUTANTS | 10 | 73% | 1% | 8 |
| 10 | DISEASE RELATED MUTATIONS | 8 | 100% | 1% | 5 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Predicting the effects of amino acid substitutions on protein function | 2006 | 353 | 77 | 45% |
| Computational SNP Analysis: Current Approaches and Future Prospects | 2014 | 3 | 49 | 90% |
| Databases of genomic variation and phenotypes: existing resources and future needs | 2013 | 9 | 7 | 71% |
| Prediction of Deleterious Nonsynonymous Single-Nucleotide Polymorphism for Human Diseases | 2013 | 11 | 30 | 60% |
| Needles in stacks of needles: finding disease-causal variants in a wealth of genomic data | 2011 | 180 | 103 | 18% |
| PI3K-AKT-mTOR-Signaling and beyond: the Complex Network in Gastroenteropancreatic Neuroendocrine Neoplasms | 2014 | 6 | 81 | 43% |
| Towards Precision Medicine: Advances in Computational Approaches for the Analysis of Human Variants | 2013 | 14 | 149 | 40% |
| Human allelic variation: perspective from protein function, structure, and evolution | 2010 | 31 | 71 | 46% |
| The Effects of Non-Synonymous Single Nucleotide Polymorphisms (nsSNPs) on Protein Protein Interactions | 2013 | 24 | 97 | 25% |
| In silico SNP analysis and bioinformatics tools: a review of the state of the art to aid drug discovery | 2011 | 15 | 29 | 86% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BRISTOL SYST BIOMED | 4 | 75% | 0.3% | 3 |
| 2 | BIOTECHNOL CHEM BIOMED ENGN | 3 | 20% | 1.7% | 15 |
| 3 | CELL BIOL GENETMGC | 2 | 67% | 0.2% | 2 |
| 4 | NETWORK BIOMED RARE DIS CIBERER | 2 | 67% | 0.2% | 2 |
| 5 | PL INFORMAT MANAGEMENT ECON | 2 | 67% | 0.2% | 2 |
| 6 | GENOM DISORDERS | 2 | 22% | 0.7% | 6 |
| 7 | NGC | 1 | 50% | 0.2% | 2 |
| 8 | NGRL MANCHESTER | 1 | 50% | 0.2% | 2 |
| 9 | BREAKTHROUGH CANC UNIT | 1 | 40% | 0.2% | 2 |
| 10 | CIRB BIOL | 1 | 40% | 0.2% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000100808 | GENOME ASSEMBLY//VARIANT CALLING//TARGET ENRICHMENT |
| 2 | 0.0000091842 | EVOLUT CANC//PETOS PARADOX//TRANSLAT CANC THER EUT |
| 3 | 0.0000081674 | HAPLOTYPE INFERENCE//PURE PARSIMONY//HAPLOTYPE ASSEMBLY |
| 4 | 0.0000079227 | STAPHYLOCOCCAL NUCLEASE//THERMODYNAMIC MOLECULAR SWITCH//PROTEIN THERMOSTABILITY |
| 5 | 0.0000064827 | JOURNAL OF BIOMEDICAL SEMANTICS//ZFIN//ZEBRAFISH INFORMAT NETWORK |
| 6 | 0.0000058879 | COMPLEX SCI SOFTWARE//3D ZERNIKE DESCRIPTOR//PROTEIN SURFACE SHAPE |
| 7 | 0.0000053201 | RARE VARIANTS//GENETIC EPIDEMIOLOGY//RARE VARIANT ANALYSIS |
| 8 | 0.0000046014 | EMBL OUTSTN//PROT INFORMAT OURCE//HLTH SCI TECHNOL ICIR |
| 9 | 0.0000045647 | CAPRI//PROTEIN PROTEIN DOCKING//PROTEIN DOCKING |
| 10 | 0.0000045338 | DNA POOLING//DNA POOL//PYROSEQUENCING TECHNOLOGY |