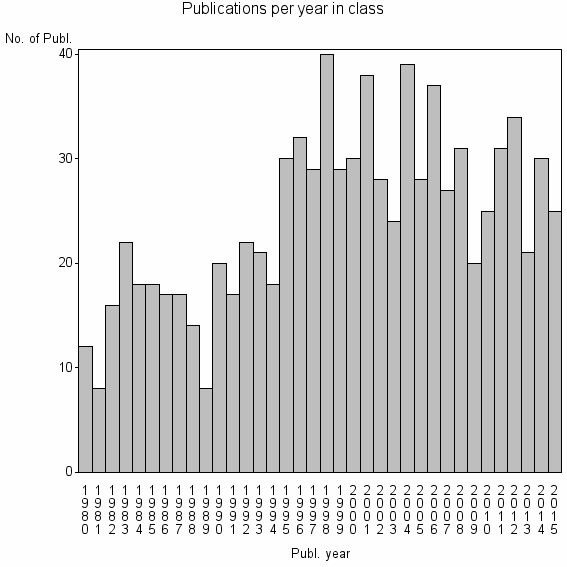
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 11979 | 876 | 35.8 | 76% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | DNA LIGASE | Author keyword | 48 | 53% | 7% | 64 |
| 2 | NAD DEPENDENT DNA LIGASE | Author keyword | 20 | 100% | 1% | 9 |
| 3 | DNA LIGASE I | Author keyword | 12 | 54% | 2% | 15 |
| 4 | DNA LIGASES | Author keyword | 11 | 78% | 1% | 7 |
| 5 | RNA TRIPHOSPHATASE | Author keyword | 8 | 75% | 1% | 6 |
| 6 | BACTERIAL DNA LIGASE | Author keyword | 3 | 100% | 0% | 3 |
| 7 | FUNCTIONAL ORGANIZATION OF THE NUCLEUS | Author keyword | 3 | 100% | 0% | 3 |
| 8 | LIGASE INHIBITOR | Author keyword | 3 | 100% | 0% | 3 |
| 9 | MRNA CAPPING ENZYME | Author keyword | 3 | 60% | 0% | 3 |
| 10 | 7 METHYLGUANOSINE | Author keyword | 3 | 42% | 1% | 5 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | DNA LIGASE | 48 | 53% | 7% | 64 | Search DNA+LIGASE | Search DNA+LIGASE |
| 2 | NAD DEPENDENT DNA LIGASE | 20 | 100% | 1% | 9 | Search NAD+DEPENDENT+DNA+LIGASE | Search NAD+DEPENDENT+DNA+LIGASE |
| 3 | DNA LIGASE I | 12 | 54% | 2% | 15 | Search DNA+LIGASE+I | Search DNA+LIGASE+I |
| 4 | DNA LIGASES | 11 | 78% | 1% | 7 | Search DNA+LIGASES | Search DNA+LIGASES |
| 5 | RNA TRIPHOSPHATASE | 8 | 75% | 1% | 6 | Search RNA+TRIPHOSPHATASE | Search RNA+TRIPHOSPHATASE |
| 6 | BACTERIAL DNA LIGASE | 3 | 100% | 0% | 3 | Search BACTERIAL+DNA+LIGASE | Search BACTERIAL+DNA+LIGASE |
| 7 | FUNCTIONAL ORGANIZATION OF THE NUCLEUS | 3 | 100% | 0% | 3 | Search FUNCTIONAL+ORGANIZATION+OF+THE+NUCLEUS | Search FUNCTIONAL+ORGANIZATION+OF+THE+NUCLEUS |
| 8 | LIGASE INHIBITOR | 3 | 100% | 0% | 3 | Search LIGASE+INHIBITOR | Search LIGASE+INHIBITOR |
| 9 | MRNA CAPPING ENZYME | 3 | 60% | 0% | 3 | Search MRNA+CAPPING+ENZYME | Search MRNA+CAPPING+ENZYME |
| 10 | 7 METHYLGUANOSINE | 3 | 42% | 1% | 5 | Search 7+METHYLGUANOSINE | Search 7+METHYLGUANOSINE |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | RNA CAPPING ENZYME | 47 | 60% | 6% | 51 |
| 2 | 5 TRIPHOSPHATASE | 33 | 66% | 4% | 31 |
| 3 | END JOINING FUNCTION | 31 | 92% | 1% | 12 |
| 4 | NICK RECOGNITION | 27 | 76% | 2% | 19 |
| 5 | ADENYLYLATION SITE | 27 | 83% | 2% | 15 |
| 6 | GUANYLYLTRANSFERASE | 23 | 40% | 5% | 45 |
| 7 | NICK | 23 | 70% | 2% | 19 |
| 8 | LIGASE D | 22 | 66% | 2% | 21 |
| 9 | GUANINE 7 METHYLTRANSFERASE COMPLEX | 21 | 75% | 2% | 15 |
| 10 | CAP METHYLTRANSFERASE | 20 | 66% | 2% | 19 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| DNA Ligases: Progress and Prospects | 2009 | 42 | 30 | 90% |
| Bacterial DNA ligases | 2001 | 122 | 27 | 89% |
| DNA and RNA ligases: structural variations and shared mechanisms | 2008 | 39 | 34 | 94% |
| The polynucleotide ligase and RNA capping enzyme superfamily of covalent nucleotidyltransferases | 2004 | 105 | 30 | 87% |
| Eukaryotic DNA ligases: Structural and functional insights | 2008 | 107 | 148 | 36% |
| Myc and mRNA capping | 2015 | 3 | 55 | 16% |
| Structure, mechanism, and evolution of the mRNA capping apparatus | 2001 | 129 | 82 | 72% |
| DNA ligases: Structure, reaction mechanism, and function | 2006 | 110 | 113 | 37% |
| Bacterial DNA repair by non-homologous end joining | 2007 | 54 | 47 | 49% |
| Enzymology of RNA cap synthesis | 2010 | 29 | 124 | 39% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | RADIOCYTOGENET | 2 | 67% | 0.2% | 2 |
| 2 | MOL BIOL SECTOR | 1 | 50% | 0.2% | 2 |
| 3 | BIOMACROMOL PROTE PLATFORM | 1 | 50% | 0.1% | 1 |
| 4 | DNA ENZYMES | 1 | 50% | 0.1% | 1 |
| 5 | GEN DAMAGE STABIL | 1 | 50% | 0.1% | 1 |
| 6 | GENET MICROBIOLIFR115 | 1 | 50% | 0.1% | 1 |
| 7 | GENOME SEQUENCING ANAL IL | 1 | 50% | 0.1% | 1 |
| 8 | HEARST MICROBIOL STRANG CANC PREVENT | 1 | 50% | 0.1% | 1 |
| 9 | INFECT DIS TAU | 1 | 50% | 0.1% | 1 |
| 10 | MOL CELLULAR BIOL PROGAM | 1 | 50% | 0.1% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000163078 | TRNA SPLICING//POLYNUCLEOTIDE KINASE//SPLICING ENDONUCLEASE |
| 2 | 0.0000088943 | DNA PK//DNA DEPENDENT PROTEIN KINASE//KU |
| 3 | 0.0000065516 | CLAMP LOADER//REPLICATION FACTOR C//DNA REPLICAT |
| 4 | 0.0000065040 | TRIVALINE//TRIPLE RINGS//NADP ANALOGS |
| 5 | 0.0000059173 | P TEFB//CDK9//HEXIM1 |
| 6 | 0.0000050936 | APE1//AP ENDONUCLEASE//APE1 REF 1 |
| 7 | 0.0000050484 | ROLLING CIRCLE AMPLIFICATION//HYBRIDIZATION CHAIN REACTION//DNAZYME |
| 8 | 0.0000049168 | DNA POLYMERASE ALPHA//DNA SYNTHESOME//DNA POLYMERASE DELTA |
| 9 | 0.0000048795 | FEN1//BRAUN S//CREAT INITIAT CELL CYCLE CONTROL |
| 10 | 0.0000045331 | DNA POOLING//DNA POOL//PYROSEQUENCING TECHNOLOGY |