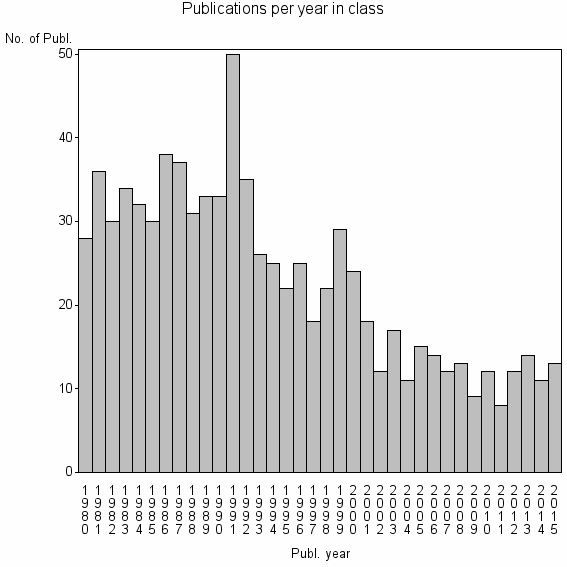
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 12630 | 829 | 34.8 | 56% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | MEIOTIC DRIVE | Author keyword | 42 | 48% | 8% | 64 |
| 2 | T HAPLOTYPE | Author keyword | 21 | 85% | 1% | 11 |
| 3 | T COMPLEX | Author keyword | 12 | 40% | 3% | 23 |
| 4 | TRANSMISSION RATIO DISTORTION | Author keyword | 5 | 32% | 1% | 12 |
| 5 | DDK SYNDROME | Author keyword | 4 | 75% | 0% | 3 |
| 6 | T HAPLOTYPES | Author keyword | 3 | 100% | 0% | 3 |
| 7 | DROSOPHILA MEDIOPUNCTATA | Author keyword | 3 | 50% | 0% | 4 |
| 8 | SRSNR | Address | 3 | 60% | 0% | 3 |
| 9 | SEGREGATION DISTORTION | Author keyword | 3 | 11% | 3% | 23 |
| 10 | TRANSMISSION DISTORTION | Author keyword | 2 | 28% | 1% | 7 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | MEIOTIC DRIVE | 42 | 48% | 8% | 64 | Search MEIOTIC+DRIVE | Search MEIOTIC+DRIVE |
| 2 | T HAPLOTYPE | 21 | 85% | 1% | 11 | Search T+HAPLOTYPE | Search T+HAPLOTYPE |
| 3 | T COMPLEX | 12 | 40% | 3% | 23 | Search T+COMPLEX | Search T+COMPLEX |
| 4 | TRANSMISSION RATIO DISTORTION | 5 | 32% | 1% | 12 | Search TRANSMISSION+RATIO+DISTORTION | Search TRANSMISSION+RATIO+DISTORTION |
| 5 | DDK SYNDROME | 4 | 75% | 0% | 3 | Search DDK+SYNDROME | Search DDK+SYNDROME |
| 6 | T HAPLOTYPES | 3 | 100% | 0% | 3 | Search T+HAPLOTYPES | Search T+HAPLOTYPES |
| 7 | DROSOPHILA MEDIOPUNCTATA | 3 | 50% | 0% | 4 | Search DROSOPHILA+MEDIOPUNCTATA | Search DROSOPHILA+MEDIOPUNCTATA |
| 8 | SEGREGATION DISTORTION | 3 | 11% | 3% | 23 | Search SEGREGATION+DISTORTION | Search SEGREGATION+DISTORTION |
| 9 | TRANSMISSION DISTORTION | 2 | 28% | 1% | 7 | Search TRANSMISSION+DISTORTION | Search TRANSMISSION+DISTORTION |
| 10 | DISTORTER | 2 | 67% | 0% | 2 | Search DISTORTER | Search DISTORTER |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | TERT HAPLOTYPES | 69 | 93% | 3% | 26 |
| 2 | TRANSMISSION RATIO DISTORTION | 56 | 48% | 10% | 87 |
| 3 | CANDIDATE GENE FAMILY | 37 | 100% | 2% | 14 |
| 4 | VIRILITY DEFICIENCY | 35 | 89% | 2% | 16 |
| 5 | MEIOTIC DRIVE | 29 | 25% | 12% | 98 |
| 6 | T T COMPLEX | 23 | 69% | 2% | 20 |
| 7 | DDK SYNDROME | 23 | 86% | 1% | 12 |
| 8 | SD SYSTEM | 23 | 100% | 1% | 10 |
| 9 | FOREIGN SPERM | 20 | 100% | 1% | 9 |
| 10 | X CHROMOSOME DRIVE | 18 | 89% | 1% | 8 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references |
% act. ref. to same field |
|---|---|---|---|---|
| SEGREGATION DISTORTERS | 1991 | 391 | 89 | 71% |
| Transmission ratio distortion in mice | 2003 | 70 | 55 | 93% |
| THE PECULIAR JOURNEY OF A SELFISH CHROMOSOME - MOUSE T-HAPLOTYPES AND MEIOTIC DRIVE | 1993 | 171 | 24 | 92% |
| Sex chromosome meiotic drive | 2001 | 141 | 75 | 56% |
| CHEATERS SOMETIMES PROSPER - DISTORTION OF MENDELIAN SEGREGATION BY MEIOTIC DRIVE | 1993 | 126 | 29 | 90% |
| The role of meiotic drive in hybrid male sterility | 2010 | 29 | 70 | 43% |
| Closing the (Ran)GAP on segregation distortion in Drosophila | 2003 | 45 | 24 | 58% |
| Putting the brake on drive: meiotic drive of t haplotypes in natural populations of mice | 1998 | 34 | 29 | 86% |
| MOUSE T-HAPLOTYPES | 1985 | 289 | 67 | 73% |
| Nonrandom segregation during meiosis: the unfairness of females | 2001 | 91 | 46 | 37% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | SRSNR | 3 | 60% | 0.4% | 3 |
| 2 | DEV TEAM MAMMALIAN CELLULAR DYNAM | 2 | 67% | 0.2% | 2 |
| 3 | BIOL GENET EVOLUCAO | 2 | 43% | 0.4% | 3 |
| 4 | MAX PLANCK INDEPENDENT GRP POPULAT GEN | 0 | 33% | 0.1% | 1 |
| 5 | U 579 | 0 | 33% | 0.1% | 1 |
| 6 | CNRSINA PG | 0 | 20% | 0.1% | 1 |
| 7 | DST NRF EXCELLENCE BIOMED TB HLTH SCI | 0 | 20% | 0.1% | 1 |
| 8 | MED GENET CBF | 0 | 20% | 0.1% | 1 |
| 9 | MOL GENET H5 | 0 | 20% | 0.1% | 1 |
| 10 | PL GENET PHYSIOLCHIKUSA KU | 0 | 20% | 0.1% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000130433 | MUS MUSCULUS DOMESTICUS//HOUSE MOUSE//RB FUSION |
| 2 | 0.0000103158 | ANIM GENET BIOINFORMAT//BASIC PROGRAMMAMMALIAN GENET//FREDERICK CANC ILBIONET INC |
| 3 | 0.0000086066 | POKEY//LAMPBRUSH LOOPS//FERTILITY GENES |
| 4 | 0.0000075297 | MOL IMMUNOL RUMENTAT//GENOMIC MATCHING TECHNIQUE//HLA A ALLELES |
| 5 | 0.0000072371 | HN1//CANC 6401 E THOMAS RD//HORMONAL PROFILING |
| 6 | 0.0000058489 | ZENTRUM ANAT ABT 5//CLASS II MHC SEQUENCE//IMMUNGENET ABT GOSSLERSTR 12D |
| 7 | 0.0000057240 | HYBRID ZONE//REPRODUCTIVE ISOLATION//HYBRID INVIABILITY |
| 8 | 0.0000053195 | MAMMALIAN MOL GENET GRP//ADRENAL X ZONE MORPHOLOGY//DDD MOUSE |
| 9 | 0.0000051893 | ECOL MODELLING GRP//GEOGRAPHICAL STRUCTURE//HARDY WEINBERG EQUATION |
| 10 | 0.0000049998 | SCHLAFEN//SURG SERV 11S//SCHLAFEN 3 |