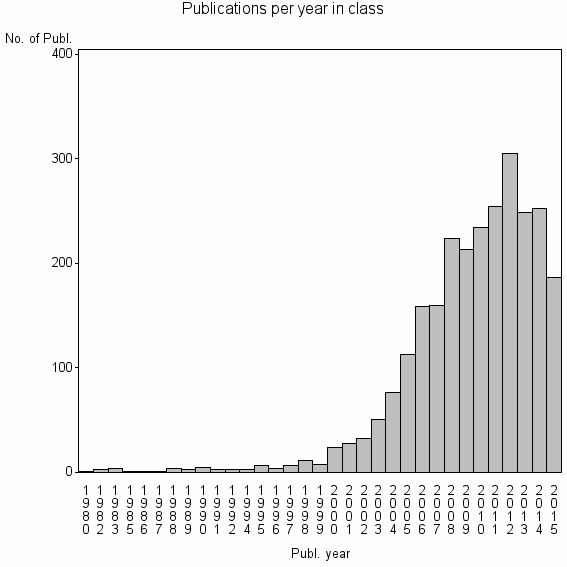
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 1332 | 2615 | 41.3 | 76% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 584 | 13517 | BMC SYSTEMS BIOLOGY//FLUX BALANCE ANALYSIS//SYNTHETIC BIOLOGY |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | LINEAR NOISE APPROXIMATION | Author keyword | 49 | 94% | 1% | 17 |
| 2 | STOCHASTIC GENE EXPRESSION | Author keyword | 37 | 68% | 1% | 32 |
| 3 | CHEMICAL MASTER EQUATION | Author keyword | 35 | 58% | 2% | 41 |
| 4 | TAU LEAPING | Author keyword | 32 | 85% | 1% | 17 |
| 5 | BIOSYST DYNAM | Address | 21 | 71% | 1% | 17 |
| 6 | STOCHASTIC SIMULATION ALGORITHM | Author keyword | 20 | 58% | 1% | 23 |
| 7 | STOCHASTIC CHEMICAL KINETICS | Author keyword | 17 | 59% | 1% | 19 |
| 8 | GENE EXPRESSION NOISE | Author keyword | 17 | 63% | 1% | 17 |
| 9 | INTRINSIC NOISE | Author keyword | 15 | 49% | 1% | 23 |
| 10 | INTEGRAT SYST BIOL UNIT | Address | 10 | 73% | 0% | 8 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | LINEAR NOISE APPROXIMATION | 49 | 94% | 1% | 17 | Search LINEAR+NOISE+APPROXIMATION | Search LINEAR+NOISE+APPROXIMATION |
| 2 | STOCHASTIC GENE EXPRESSION | 37 | 68% | 1% | 32 | Search STOCHASTIC+GENE+EXPRESSION | Search STOCHASTIC+GENE+EXPRESSION |
| 3 | CHEMICAL MASTER EQUATION | 35 | 58% | 2% | 41 | Search CHEMICAL+MASTER+EQUATION | Search CHEMICAL+MASTER+EQUATION |
| 4 | TAU LEAPING | 32 | 85% | 1% | 17 | Search TAU+LEAPING | Search TAU+LEAPING |
| 5 | STOCHASTIC SIMULATION ALGORITHM | 20 | 58% | 1% | 23 | Search STOCHASTIC+SIMULATION+ALGORITHM | Search STOCHASTIC+SIMULATION+ALGORITHM |
| 6 | STOCHASTIC CHEMICAL KINETICS | 17 | 59% | 1% | 19 | Search STOCHASTIC+CHEMICAL+KINETICS | Search STOCHASTIC+CHEMICAL+KINETICS |
| 7 | GENE EXPRESSION NOISE | 17 | 63% | 1% | 17 | Search GENE+EXPRESSION+NOISE | Search GENE+EXPRESSION+NOISE |
| 8 | INTRINSIC NOISE | 15 | 49% | 1% | 23 | Search INTRINSIC+NOISE | Search INTRINSIC+NOISE |
| 9 | GILLESPIE ALGORITHM | 9 | 38% | 1% | 19 | Search GILLESPIE+ALGORITHM | Search GILLESPIE+ALGORITHM |
| 10 | TAU LEAP | 9 | 83% | 0% | 5 | Search TAU+LEAP | Search TAU+LEAP |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | STOCHASTIC GENE EXPRESSION | 138 | 51% | 7% | 194 |
| 2 | SINGLE CELL | 68 | 20% | 11% | 300 |
| 3 | FLUCTUATING ENVIRONMENTS | 64 | 41% | 5% | 122 |
| 4 | REACTING SYSTEMS | 49 | 60% | 2% | 54 |
| 5 | CHEMICALLY REACTING SYSTEMS | 47 | 47% | 3% | 75 |
| 6 | EUKARYOTIC GENE EXPRESSION | 45 | 50% | 3% | 66 |
| 7 | EXACT STOCHASTIC SIMULATION | 44 | 45% | 3% | 74 |
| 8 | LINEAR NOISE APPROXIMATION | 44 | 88% | 1% | 21 |
| 9 | STOCHASTICITY | 41 | 25% | 6% | 146 |
| 10 | DISSIPATION COST | 41 | 100% | 1% | 15 |
Journals |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | IET SYSTEMS BIOLOGY | 7 | 15% | 2% | 42 |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Using Gene Expression Noise to Understand Gene Regulation | 2012 | 133 | 33 | 73% |
| Functional roles for noise in genetic circuits | 2010 | 387 | 75 | 63% |
| Stochastic simulation of chemical kinetics | 2007 | 513 | 31 | 94% |
| Nature, Nurture, or Chance: Stochastic Gene Expression and Its Consequences | 2008 | 612 | 98 | 65% |
| Network motifs: theory and experimental approaches | 2007 | 943 | 90 | 41% |
| Cellular Decision Making and Biological Noise: From Microbes to Mammals | 2011 | 204 | 93 | 49% |
| Genetic Determinants and Cellular Constraints in Noisy Gene Expression | 2013 | 41 | 40 | 80% |
| Stochasticity in gene expression: From theories to phenotypes | 2005 | 977 | 123 | 57% |
| Regulation of Noise in Gene Expression | 2013 | 23 | 57 | 75% |
| Noise in biology | 2014 | 14 | 198 | 44% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BIOSYST DYNAM | 21 | 71% | 0.7% | 17 |
| 2 | INTEGRAT SYST BIOL UNIT | 10 | 73% | 0.3% | 8 |
| 3 | CHEM PHYS PL MATH | 6 | 40% | 0.5% | 12 |
| 4 | COMPUTAT SYST BIOL GRP | 6 | 19% | 1.0% | 27 |
| 5 | SYNTHSYS EDINBURGH | 4 | 56% | 0.2% | 5 |
| 6 | ADV COMPUTAT MODELLING | 3 | 19% | 0.5% | 14 |
| 7 | BISON GRP | 3 | 60% | 0.1% | 3 |
| 8 | GUANGZHOU BIOINFORMAT | 3 | 42% | 0.2% | 5 |
| 9 | BIOCOMPLEX INFORMAT | 2 | 11% | 0.8% | 20 |
| 10 | AIHARA | 2 | 67% | 0.1% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000246439 | GENETIC REGULATORY NETWORKS GRNS//GENETIC REGULATORY NETWORKS//GENETIC REGULATORY NETWORK |
| 2 | 0.0000176912 | MASS ACTION KINETICS//COMPLEX BALANCING//CHEMICAL REACTION NETWORKS |
| 3 | 0.0000175145 | ITO STRATONOVICH DILEMMA//INFORMAT ENGN COMP SCI MATH DISIM//AGO ANTAGONIST MODELS |
| 4 | 0.0000161673 | CELL DECIS PROC//MATH ARTS SCI//SIGNALING SYST |
| 5 | 0.0000160133 | SYNTHETIC BIOLOGY//ACS SYNTHETIC BIOLOGY//CALIF QUANTITAT BIOL QB3 |
| 6 | 0.0000148205 | ERATO COMPLEX SYST BIOL//ISOLOGOUS DIVERSIFICATION//COHERENT SPIKING |
| 7 | 0.0000107393 | DIGITAL PCR//DROPLET DIGITAL PCR//SINGLE CELL TRANSCRIPTOMICS |
| 8 | 0.0000095241 | BOOLEAN NETWORKS//SEMI TENSOR PRODUCT//BOOLEAN NETWORK |
| 9 | 0.0000089865 | PROGRAM COMPUTAT GENOM//CIFN//PROGRAMA GENOM COMPUTAC |
| 10 | 0.0000089548 | CELLML//BIOINFORMAT TUEBINGEN ZBIT//SCI DATABASES VISUALIZAT GRP |