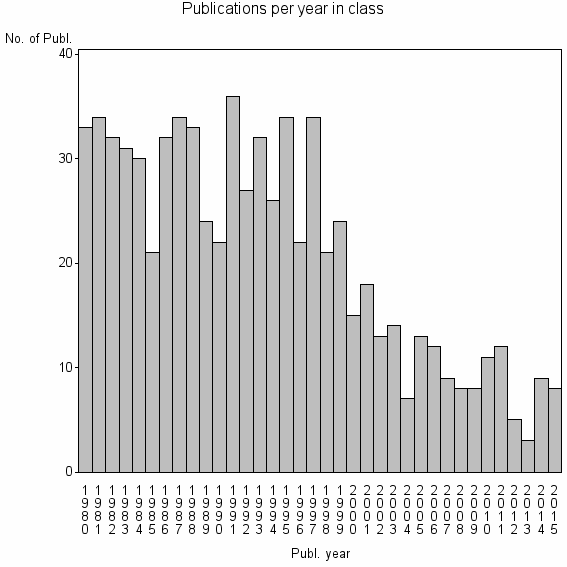
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 13832 | 747 | 37.8 | 56% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 1343 | 7871 | BACTERIOPHAGE//PHAGE THERAPY//DNA PACKAGING |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | M13 COAT PROTEIN | Author keyword | 14 | 100% | 1% | 7 |
| 2 | GENE 3 PROTEIN | Author keyword | 11 | 78% | 1% | 7 |
| 3 | FILAMENTOUS BACTERIOPHAGE | Author keyword | 9 | 36% | 3% | 21 |
| 4 | GENE 5 PROTEIN | Author keyword | 9 | 67% | 1% | 8 |
| 5 | SELECTIVELY INFECTIVE PHAGE | Author keyword | 8 | 100% | 1% | 5 |
| 6 | PHAGE ASSEMBLY | Author keyword | 5 | 54% | 1% | 7 |
| 7 | PHAGE FD | Author keyword | 4 | 67% | 1% | 4 |
| 8 | FF GENE 5 PROTEIN | Author keyword | 4 | 75% | 0% | 3 |
| 9 | FIBRE DIFFRACTION | Author keyword | 4 | 40% | 1% | 8 |
| 10 | GENE V PROTEIN | Author keyword | 4 | 56% | 1% | 5 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | M13 COAT PROTEIN | 14 | 100% | 1% | 7 | Search M13+COAT+PROTEIN | Search M13+COAT+PROTEIN |
| 2 | GENE 3 PROTEIN | 11 | 78% | 1% | 7 | Search GENE+3+PROTEIN | Search GENE+3+PROTEIN |
| 3 | FILAMENTOUS BACTERIOPHAGE | 9 | 36% | 3% | 21 | Search FILAMENTOUS+BACTERIOPHAGE | Search FILAMENTOUS+BACTERIOPHAGE |
| 4 | GENE 5 PROTEIN | 9 | 67% | 1% | 8 | Search GENE+5+PROTEIN | Search GENE+5+PROTEIN |
| 5 | SELECTIVELY INFECTIVE PHAGE | 8 | 100% | 1% | 5 | Search SELECTIVELY+INFECTIVE+PHAGE | Search SELECTIVELY+INFECTIVE+PHAGE |
| 6 | PHAGE ASSEMBLY | 5 | 54% | 1% | 7 | Search PHAGE+ASSEMBLY | Search PHAGE+ASSEMBLY |
| 7 | PHAGE FD | 4 | 67% | 1% | 4 | Search PHAGE+FD | Search PHAGE+FD |
| 8 | FF GENE 5 PROTEIN | 4 | 75% | 0% | 3 | Search FF+GENE+5+PROTEIN | Search FF+GENE+5+PROTEIN |
| 9 | FIBRE DIFFRACTION | 4 | 40% | 1% | 8 | Search FIBRE+DIFFRACTION | Search FIBRE+DIFFRACTION |
| 10 | GENE V PROTEIN | 4 | 56% | 1% | 5 | Search GENE+V+PROTEIN | Search GENE+V+PROTEIN |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | FF FD | 51 | 91% | 3% | 21 |
| 2 | 5 PROTEIN | 45 | 94% | 2% | 16 |
| 3 | IKE | 42 | 78% | 4% | 28 |
| 4 | FD | 39 | 38% | 11% | 83 |
| 5 | FILAMENTOUS BACTERIOPHAGE M13 | 27 | 92% | 1% | 11 |
| 6 | BACTERIOPHAGE M13 | 26 | 53% | 5% | 35 |
| 7 | BACTERIOPHAGE FD | 26 | 53% | 5% | 34 |
| 8 | FD ADSORPTION PROTEIN | 21 | 75% | 2% | 15 |
| 9 | INOVIRUS | 21 | 90% | 1% | 9 |
| 10 | ADSORPTION PROTEIN | 18 | 59% | 3% | 20 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Viruses: incredible nanomachines. New advances with filamentous phages | 2010 | 32 | 62 | 60% |
| Structure and assembly of filamentous bacteriophages | 2014 | 14 | 322 | 51% |
| Raman Tensors and their application in structural studies of biological systems | 2009 | 12 | 47 | 85% |
| Filamentous Bacteriophage: Biology, Phage Display and Nanotechnology Applications | 2011 | 56 | 218 | 25% |
| FILAMENTOUS PHAGE ASSEMBLY | 1991 | 128 | 32 | 66% |
| Protein-lipid interactions of bacteriophage M13 major coat protein | 2003 | 28 | 77 | 82% |
| Raman spectroscopy of protein and nucleic acid assemblies | 1999 | 142 | 73 | 29% |
| CD of single-stranded, double-stranded, and G-quartet nucleic acids in complexes with a single-stranded DNA-binding protein | 2002 | 11 | 22 | 91% |
| DNA PACKING IN FILAMENTOUS BACTERIOPHAGES | 1988 | 90 | 67 | 76% |
| From 'I' to 'L' and back again: the odyssey of membrane-bound M13 protein | 2009 | 7 | 36 | 53% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BAYREUTHER ZENTRUM MOL BIOWISSEN | 2 | 20% | 1.3% | 10 |
| 2 | BIOCHEM BAYREUTHER | 1 | 50% | 0.3% | 2 |
| 3 | PHYS BIOPHYS MITTELDEUT | 1 | 50% | 0.3% | 2 |
| 4 | ZENTRUM MOL BIOWISSEN | 1 | 33% | 0.3% | 2 |
| 5 | PROT KRISTALLOG | 1 | 21% | 0.4% | 3 |
| 6 | BIOPHYS CHEM NIJMEGEN SON | 1 | 50% | 0.1% | 1 |
| 7 | ZENTRUM STRUKTUR DYNAM PROT MZP | 1 | 50% | 0.1% | 1 |
| 8 | BAYREUTHER ZENTRUM MOL BIOWISSEN AFTEN | 0 | 33% | 0.1% | 1 |
| 9 | MRC PROT STRUCT FUNCT GRP | 0 | 33% | 0.1% | 1 |
| 10 | MITTELDEUT ZENTRUM STRUKTUR DYNAM PROT MZP | 0 | 13% | 0.3% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000195447 | BACTERIOPHAGE ALPHA 3//MICROVIRUS//BACTERIOPHAGE PHI X174 |
| 2 | 0.0000120189 | PHAGE DISPLAY//LANDSCAPE PHAGE//IN VIVO PHAGE DISPLAY |
| 3 | 0.0000109975 | ULTRAVIOLET RESONANCE RAMAN//UV RESONANCE RAMAN SPECTROSCOPY//DEEP UV RESONANCE RAMAN SPECTROSCOPY |
| 4 | 0.0000105255 | REPLICATION PROTEIN A//SINGLE STRANDED DNA BINDING PROTEIN//REPLICATION PROTEIN A RPA |
| 5 | 0.0000074162 | MEGAPRIMER//OVERLAP EXTENSION//MULTIPLE SITE DIRECTED MUTAGENESIS |
| 6 | 0.0000060542 | SPEKT ABT//NON LINEAR EPR//ABT SPEKTROSKOPIE |
| 7 | 0.0000057310 | TYPE II SECRETION//TYPE IV PILI//GENERAL SECRETORY PATHWAY |
| 8 | 0.0000056275 | COLICIN//COLICIN E1//TOLA |
| 9 | 0.0000053132 | ROLLING CIRCLE REPLICATION//PLASMID REPLICATION CONTROL//CRYPTIC PLASMID |
| 10 | 0.0000049034 | WARWICK ANALYT SCI//GELRED//DNA TOPOISOMERS |