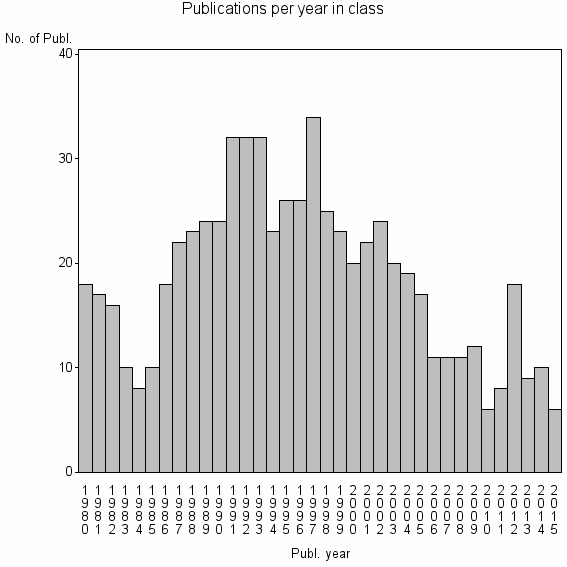
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 15008 | 667 | 32.6 | 68% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 501 | 14604 | RNA POLYMERASE//SITE SPECIFIC RECOMBINATION//H NS |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | T7 RNA POLYMERASE | Author keyword | 31 | 34% | 11% | 73 |
| 2 | T7 LYSOZYME | Author keyword | 19 | 76% | 2% | 13 |
| 3 | PHAGE RNA POLYMERASE | Author keyword | 9 | 83% | 1% | 5 |
| 4 | MORSE MOL GENET | Address | 8 | 39% | 3% | 17 |
| 5 | ABORTIVE TRANSCRIPTION | Author keyword | 4 | 67% | 1% | 4 |
| 6 | BACTERIOPHAGE T7 | Author keyword | 4 | 26% | 2% | 13 |
| 7 | DNTP UTILIZATION | Author keyword | 3 | 100% | 0% | 3 |
| 8 | K11 RNA POLYMERASE | Author keyword | 3 | 100% | 0% | 3 |
| 9 | BACTERIOPHAGE T7 RNA POLYMERASE | Author keyword | 3 | 60% | 0% | 3 |
| 10 | SUMMER UNDER EXPERIENCE PROGRAM | Address | 2 | 67% | 0% | 2 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | T7 RNA POLYMERASE | 31 | 34% | 11% | 73 | Search T7+RNA+POLYMERASE | Search T7+RNA+POLYMERASE |
| 2 | T7 LYSOZYME | 19 | 76% | 2% | 13 | Search T7+LYSOZYME | Search T7+LYSOZYME |
| 3 | PHAGE RNA POLYMERASE | 9 | 83% | 1% | 5 | Search PHAGE+RNA+POLYMERASE | Search PHAGE+RNA+POLYMERASE |
| 4 | ABORTIVE TRANSCRIPTION | 4 | 67% | 1% | 4 | Search ABORTIVE+TRANSCRIPTION | Search ABORTIVE+TRANSCRIPTION |
| 5 | BACTERIOPHAGE T7 | 4 | 26% | 2% | 13 | Search BACTERIOPHAGE+T7 | Search BACTERIOPHAGE+T7 |
| 6 | DNTP UTILIZATION | 3 | 100% | 0% | 3 | Search DNTP+UTILIZATION | Search DNTP+UTILIZATION |
| 7 | K11 RNA POLYMERASE | 3 | 100% | 0% | 3 | Search K11+RNA+POLYMERASE | Search K11+RNA+POLYMERASE |
| 8 | BACTERIOPHAGE T7 RNA POLYMERASE | 3 | 60% | 0% | 3 | Search BACTERIOPHAGE+T7+RNA+POLYMERASE | Search BACTERIOPHAGE+T7+RNA+POLYMERASE |
| 9 | TRNA REPORTER GENE | 2 | 67% | 0% | 2 | Search TRNA+REPORTER+GENE | Search TRNA+REPORTER+GENE |
| 10 | RNA DNA HYBRID | 2 | 43% | 0% | 3 | Search RNA++DNA+HYBRID | Search RNA++DNA+HYBRID |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BACTERIOPHAGE T7 LATE PROMOTERS | 35 | 86% | 3% | 18 |
| 2 | T7 RNA POLYMERASE | 24 | 19% | 17% | 115 |
| 3 | BACTERIOPHAGE T7 | 24 | 21% | 15% | 103 |
| 4 | SMALL SYNTHETIC PROMOTERS | 14 | 100% | 1% | 7 |
| 5 | INITIAL BUBBLE COLLAPSE | 12 | 86% | 1% | 6 |
| 6 | BACTERIOPHAGE T3 | 10 | 52% | 2% | 14 |
| 7 | RIBONUCLEIC ACID POLYMERASE | 9 | 27% | 4% | 30 |
| 8 | TRANSCRIBING ACTIVITIES | 9 | 83% | 1% | 5 |
| 9 | INVITRO TRANSCRIPTION PROPERTIES | 8 | 100% | 1% | 5 |
| 10 | SP6 | 7 | 53% | 1% | 10 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| USE OF T7 RNA-POLYMERASE TO DIRECT EXPRESSION OF CLONED GENES | 1990 | 6055 | 16 | 56% |
| T7 RNA polymerase | 2003 | 24 | 110 | 65% |
| Mechanism of transcription initiation by the yeast mitochondrial RNA polymerase | 2012 | 8 | 95 | 34% |
| Insights into transcription: structure and function of single-subunit DNA-dependent RNA polymerases | 2000 | 72 | 34 | 65% |
| Structure and function in promoter escape by T7 RNA polymerase | 2005 | 7 | 57 | 81% |
| Evaluation of fluorescence spectroscopy methods for mapping melted regions of DNA along the transcription pathway | 2003 | 14 | 45 | 51% |
| Recent studies of T7 RNA polymerase mechanism | 1998 | 15 | 36 | 83% |
| Structural-functional analysis of bacteriophage T7 RNA polymerase | 2002 | 12 | 36 | 86% |
| THE PHAGE RNA-POLYMERASES ARE RELATED TO DNA-POLYMERASES AND REVERSE TRANSCRIPTASES | 1993 | 53 | 42 | 62% |
| Visualizing polynucleotide polymerase machines at work | 2006 | 54 | 43 | 23% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MORSE MOL GENET | 8 | 39% | 2.5% | 17 |
| 2 | SUMMER UNDER EXPERIENCE PROGRAM | 2 | 67% | 0.3% | 2 |
| 3 | SANDOZ CO | 1 | 50% | 0.1% | 1 |
| 4 | HIGH THROUGHPUT TORY | 1 | 29% | 0.3% | 2 |
| 5 | STRATFORD | 0 | 33% | 0.1% | 1 |
| 6 | UMR 8183 | 0 | 33% | 0.1% | 1 |
| 7 | BK21 VET SCI | 0 | 25% | 0.1% | 1 |
| 8 | ENGN MED SCI | 0 | 13% | 0.3% | 2 |
| 9 | OR BECKER | 0 | 20% | 0.1% | 1 |
| 10 | RIKEN TSUKUBA LIFE SCI | 0 | 20% | 0.1% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000164781 | BACTERIOPHAGE T5//HOST PHAGE INTERACTION//LYTIC CONVERSION |
| 2 | 0.0000153309 | RNA POLYMERASE//ESCHERICHIA COLI RNA POLYMERASE//SIGMA70 |
| 3 | 0.0000141103 | NUSG//TRANSCRIPTION TERMINATION//RHO FACTOR |
| 4 | 0.0000134492 | REX EXCLUSION//BACTERIOPHAGE LAMBDA LAMBDA//CI REXA REXB OPERON |
| 5 | 0.0000094283 | DNA PACKAGING//BACTERIOPHAGE P22//PORTAL PROTEIN |
| 6 | 0.0000090286 | DNA POLYMERASE//DNA POLYMERASE BETA//DNA POLYMERASE LAMBDA |
| 7 | 0.0000084660 | MITOCHONDRIAL NUCLEOIDS//TFAM//MITOCHONDRIAL NUCLEOID |
| 8 | 0.0000078999 | OBSTET GYNECOLSEALY STRUCT BIOL//PRIMASE//HELICASE |
| 9 | 0.0000068288 | DISACCHARIDE NUCLEOSIDES//DISACCHARIDE NUCLEOSIDE//2 O MODIFIED NUCLEOSIDES |
| 10 | 0.0000056917 | SD SEQUENCE//RIBOSOMAL PROTEIN S1//SHINE DALGARNO SEQUENCE |