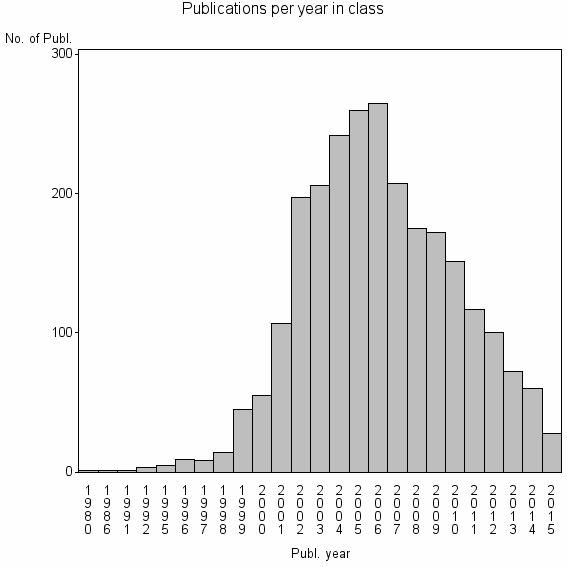
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 1587 | 2501 | 30.1 | 75% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 1650 | 6377 | BICLUSTERING//PROGRAM TRANSLAT IMMUNOVIROL BIODEF//MAYO VACCINE GRP |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | MICROARRAY IMAGE PROCESSING | Author keyword | 9 | 83% | 0% | 5 |
| 2 | GENE FILTERING | Author keyword | 8 | 75% | 0% | 6 |
| 3 | MAQC | Author keyword | 8 | 100% | 0% | 5 |
| 4 | PLATFORM COMPARISON | Author keyword | 7 | 64% | 0% | 7 |
| 5 | DYE BIAS | Author keyword | 6 | 80% | 0% | 4 |
| 6 | BATCH EFFECT REMOVAL | Author keyword | 6 | 100% | 0% | 4 |
| 7 | MICROARRAY EXPERIMENTS | Author keyword | 5 | 47% | 0% | 8 |
| 8 | MICROARRAY IMAGE | Author keyword | 5 | 63% | 0% | 5 |
| 9 | MAQC II | Author keyword | 4 | 67% | 0% | 4 |
| 10 | CDNA MICROARRAY IMAGES | Author keyword | 4 | 75% | 0% | 3 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | MICROARRAY IMAGE PROCESSING | 9 | 83% | 0% | 5 | Search MICROARRAY+IMAGE+PROCESSING | Search MICROARRAY+IMAGE+PROCESSING |
| 2 | GENE FILTERING | 8 | 75% | 0% | 6 | Search GENE+FILTERING | Search GENE+FILTERING |
| 3 | MAQC | 8 | 100% | 0% | 5 | Search MAQC | Search MAQC |
| 4 | PLATFORM COMPARISON | 7 | 64% | 0% | 7 | Search PLATFORM+COMPARISON | Search PLATFORM+COMPARISON |
| 5 | DYE BIAS | 6 | 80% | 0% | 4 | Search DYE+BIAS | Search DYE+BIAS |
| 6 | BATCH EFFECT REMOVAL | 6 | 100% | 0% | 4 | Search BATCH+EFFECT+REMOVAL | Search BATCH+EFFECT+REMOVAL |
| 7 | MICROARRAY EXPERIMENTS | 5 | 47% | 0% | 8 | Search MICROARRAY+EXPERIMENTS | Search MICROARRAY+EXPERIMENTS |
| 8 | MICROARRAY IMAGE | 5 | 63% | 0% | 5 | Search MICROARRAY+IMAGE | Search MICROARRAY+IMAGE |
| 9 | MAQC II | 4 | 67% | 0% | 4 | Search MAQC+II | Search MAQC+II |
| 10 | CDNA MICROARRAY IMAGES | 4 | 75% | 0% | 3 | Search CDNA+MICROARRAY+IMAGES | Search CDNA+MICROARRAY+IMAGES |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | AFFYMETRIX | 40 | 69% | 1% | 34 |
| 2 | OLIGONUCLEOTIDE ARRAYS | 35 | 15% | 9% | 220 |
| 3 | GENE EXPRESSION MEASUREMENTS | 28 | 44% | 2% | 48 |
| 4 | PLATFORM CONSISTENCY | 23 | 100% | 0% | 10 |
| 5 | CDNA MICROARRAYS | 22 | 17% | 5% | 117 |
| 6 | ART NO E15 | 20 | 62% | 1% | 21 |
| 7 | CONTROL MAQC PROJECT | 18 | 37% | 2% | 40 |
| 8 | SUMMARIES | 18 | 31% | 2% | 48 |
| 9 | DNA MICROARRAYS | 17 | 10% | 6% | 151 |
| 10 | DNA MICROARRAY | 17 | 11% | 6% | 139 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| TM4 microarray software suite | 2006 | 861 | 44 | 50% |
| Microarray data analysis: from disarray to consolidation and consensus | 2006 | 657 | 76 | 45% |
| Comprehensive literature review and statistical considerations for microarray meta-analysis | 2012 | 48 | 118 | 40% |
| Reuse of public genome-wide gene expression data | 2013 | 43 | 91 | 30% |
| Microarray data normalization and transformation | 2002 | 904 | 9 | 67% |
| Reproducible and reliable microarray results through quality control: good laboratory proficiency and appropriate data analysis practices are essential | 2008 | 59 | 29 | 93% |
| Fundamentals of experimental design for cDNA microarrays | 2002 | 589 | 16 | 81% |
| Reliability and reproducibility issues in DNA microarray measurements | 2006 | 286 | 57 | 42% |
| Key Issues in Conducting a Meta-Analysis of Gene Expression Microarray Datasets | 2008 | 110 | 91 | 35% |
| Exploring the new world of the genome with DNA microarrays | 1999 | 1424 | 20 | 50% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MAX MCGEE JUVENILE DIABET | 3 | 21% | 0.5% | 12 |
| 2 | Z TECH | 2 | 38% | 0.2% | 5 |
| 3 | BIOMOL ANAL IMAGING | 2 | 67% | 0.1% | 2 |
| 4 | TRANSCRIPT GENOM CORE | 2 | 67% | 0.1% | 2 |
| 5 | IAN POTTER FDN CANC GENOM PREDICT MED | 2 | 50% | 0.1% | 3 |
| 6 | CANC GENOM CORE | 2 | 23% | 0.3% | 7 |
| 7 | INFORMAT NANTES ATLANTIQUE LINA | 1 | 100% | 0.1% | 2 |
| 8 | LOTTERYW STATE MICROARRAY IL | 1 | 100% | 0.1% | 2 |
| 9 | SHANGHAI INFORMAT SCI | 1 | 100% | 0.1% | 2 |
| 10 | WALLACE TUMOR 153 | 1 | 50% | 0.1% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000179702 | DIFFERENTIAL GENE EXPRESSION DETECTION//CANCER OUTLIER PROFILE ANALYSIS//BREAST CANC DIAG TREATMENT HENAN |
| 2 | 0.0000175366 | BICLUSTERING//GENE EXPRESSION DATA//GENE EXPRESSION DATA ANALYSIS |
| 3 | 0.0000138546 | FALSE DISCOVERY RATE//MULTIPLE TESTING//FAMILYWISE ERROR RATE |
| 4 | 0.0000133474 | GENE SET ANALYSIS//ENRICHMENT ANALYSIS//FUNCT GENOM NODE |
| 5 | 0.0000123076 | NONSPECIFIC HYBRIDIZATIONS//NEAREST NEIGHBOR PROPERTIES//NUCLEIC ACID OLIGOMERS |
| 6 | 0.0000109666 | RTF//ESTADD//ESTOUT |
| 7 | 0.0000108654 | SEQUENCING BY HYBRIDIZATION//DNA SEQUENCING WITH ERRORS//GAPPED PROBES |
| 8 | 0.0000107136 | TOXICOGENOMICS//TOXICOGENOM INFORMAT PROJECT//HAZCHEM |
| 9 | 0.0000097039 | RNA AMPLIFICATION//LASER CAPTURE MICRODISSECTION//LASER MICRODISSECTION |
| 10 | 0.0000095173 | GENE SELECTION//FEATURE SELECTION//CANCER CLASSIFICATION |