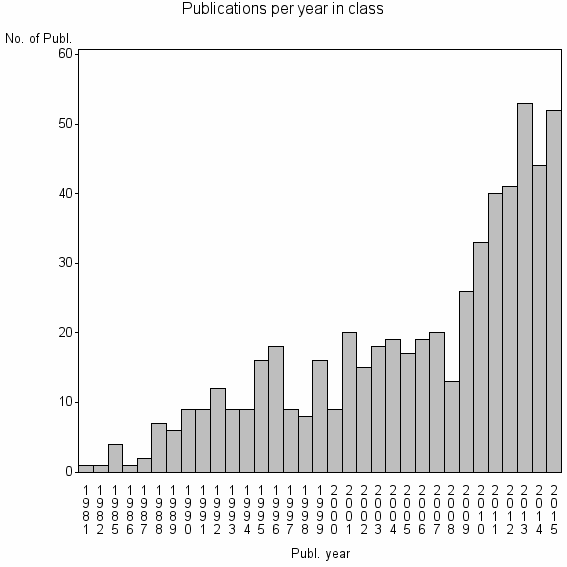
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 16697 | 576 | 37.0 | 89% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 1568 | 6715 | NEXT GENERATION SEQUENCING//RNA SEQ//COPY NUMBER VARIATION |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | SPLICED LEADER | Author keyword | 21 | 46% | 6% | 33 |
| 2 | TRANS SPLICING | Author keyword | 20 | 28% | 11% | 61 |
| 3 | CIRCULAR RNA | Author keyword | 8 | 45% | 2% | 13 |
| 4 | RNA TRANS SPLICING | Author keyword | 4 | 67% | 1% | 4 |
| 5 | SPLICE LEADER | Author keyword | 4 | 75% | 1% | 3 |
| 6 | CDK10 | Author keyword | 4 | 56% | 1% | 5 |
| 7 | CIRCRNA | Author keyword | 3 | 57% | 1% | 4 |
| 8 | CHIMERIC RNAS | Author keyword | 3 | 100% | 1% | 3 |
| 9 | MOL DERMATOL EB HOUSE AUSTRIA | Address | 3 | 60% | 1% | 3 |
| 10 | CHIMERIC MRNA | Author keyword | 2 | 67% | 0% | 2 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | SPLICED LEADER | 21 | 46% | 6% | 33 | Search SPLICED+LEADER | Search SPLICED+LEADER |
| 2 | TRANS SPLICING | 20 | 28% | 11% | 61 | Search TRANS+SPLICING | Search TRANS+SPLICING |
| 3 | CIRCULAR RNA | 8 | 45% | 2% | 13 | Search CIRCULAR+RNA | Search CIRCULAR+RNA |
| 4 | RNA TRANS SPLICING | 4 | 67% | 1% | 4 | Search RNA+TRANS+SPLICING | Search RNA+TRANS+SPLICING |
| 5 | SPLICE LEADER | 4 | 75% | 1% | 3 | Search SPLICE+LEADER | Search SPLICE+LEADER |
| 6 | CDK10 | 4 | 56% | 1% | 5 | Search CDK10 | Search CDK10 |
| 7 | CIRCRNA | 3 | 57% | 1% | 4 | Search CIRCRNA | Search CIRCRNA |
| 8 | CHIMERIC RNAS | 3 | 100% | 1% | 3 | Search CHIMERIC+RNAS | Search CHIMERIC+RNAS |
| 9 | CHIMERIC MRNA | 2 | 67% | 0% | 2 | Search CHIMERIC+MRNA | Search CHIMERIC+MRNA |
| 10 | MRNA REPAIR | 2 | 67% | 0% | 2 | Search MRNA+REPAIR | Search MRNA+REPAIR |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | EXON REPETITION | 21 | 85% | 2% | 11 |
| 2 | EXON CIRCULARIZATION | 18 | 83% | 2% | 10 |
| 3 | CAENORHABDITIS ELEGANS OPERONS | 8 | 70% | 1% | 7 |
| 4 | SPLICED LEADER ADDITION | 8 | 75% | 1% | 6 |
| 5 | TRANSSPLICING INVITRO | 8 | 75% | 1% | 6 |
| 6 | TRIMETHYLGUANOSINE CAP | 7 | 43% | 2% | 12 |
| 7 | RECURRENT FUSION | 5 | 54% | 1% | 7 |
| 8 | HUMAN PARASITE | 5 | 41% | 2% | 9 |
| 9 | PISSLRE | 4 | 44% | 1% | 7 |
| 10 | ELEGANS GENE | 3 | 100% | 1% | 3 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Detecting and characterizing circular RNAs | 2014 | 39 | 48 | 54% |
| SL trans-splicing: easy come or easy go? | 2005 | 93 | 62 | 52% |
| Circular RNAs: diversity of form and function | 2014 | 15 | 67 | 43% |
| Alternative trans-splicing: a novel mode of pre-mRNA processing | 2006 | 65 | 32 | 53% |
| Nematode spliced leaders - Ubiquity, evolution and utility | 1996 | 91 | 43 | 51% |
| Trans-splicing | 2011 | 28 | 137 | 58% |
| Spliceosome-mediated RNA trans-splicing | 2005 | 24 | 33 | 64% |
| TRANSSPLICING AND POLYCISTRONIC TRANSCRIPTION IN CAENORHABDITIS-ELEGANS | 1995 | 159 | 27 | 48% |
| Implications of chimaeric non-co-linear transcripts | 2009 | 75 | 45 | 31% |
| Chimeric RNAs as potential biomarkers for tumor diagnosis | 2012 | 4 | 44 | 57% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MOL DERMATOL EB HOUSE AUSTRIA | 3 | 60% | 0.5% | 3 |
| 2 | INRIA BAMBOO | 1 | 50% | 0.3% | 2 |
| 3 | 115 | 1 | 50% | 0.2% | 1 |
| 4 | CNRSUMR 1599 | 1 | 50% | 0.2% | 1 |
| 5 | COMPETENCE PERSONALIZED MED | 1 | 50% | 0.2% | 1 |
| 6 | DPT BIOL MOL CELULAR | 1 | 50% | 0.2% | 1 |
| 7 | GRUP RECERCA INFORMAT BIOMED | 1 | 50% | 0.2% | 1 |
| 8 | INSERMSC13 | 1 | 50% | 0.2% | 1 |
| 9 | INT MAX PLANCK MOL CELLULAR LIFE SCI | 1 | 50% | 0.2% | 1 |
| 10 | JR GRP TRANSCRIPTOME BIOINFORMAT | 1 | 50% | 0.2% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000124032 | BIDIRECTIONAL PROMOTER//NEIGHBORING GENES//GENOME NETWORK PROJECT CORE GRPGENOM SCI |
| 2 | 0.0000109858 | RNA SEQ//DE NOVO ASSEMBLY//TRANSCRIPTOME ASSEMBLY |
| 3 | 0.0000081463 | TRYPANOSOMA BRUCEI//VARIANT SURFACE GLYCOPROTEIN//TRYPANOSOME |
| 4 | 0.0000076623 | SPLICEOSOME//PRE MRNA SPLICING//PRP24 |
| 5 | 0.0000070173 | PCA3//TMPRSS2 ERG//PROSTATE CANCER ANTIGEN 3 |
| 6 | 0.0000064763 | SR PROTEINS//SR PROTEIN//ALTERNATIVE SPLICING |
| 7 | 0.0000060486 | LONG NONCODING RNAS//LNCRNA//LONG NON CODING RNA |
| 8 | 0.0000056346 | ZNF703//C8ORF4//8P12 |
| 9 | 0.0000055963 | GENOME ASSEMBLY//VARIANT CALLING//TARGET ENRICHMENT |
| 10 | 0.0000051583 | EVOLUT CANC//PETOS PARADOX//TRANSLAT CANC THER EUT |