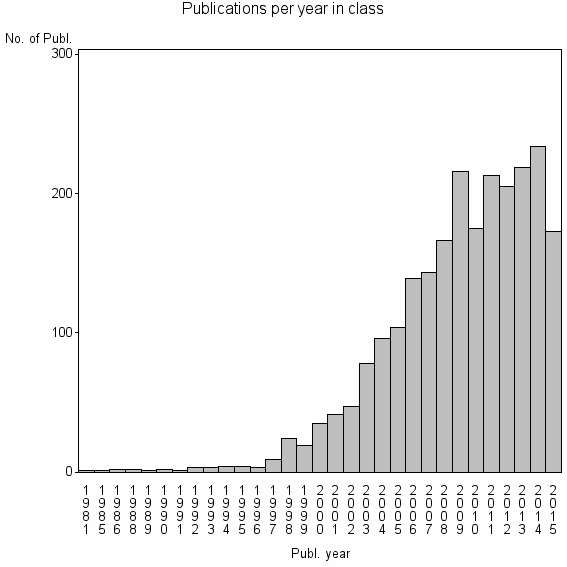
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 1908 | 2363 | 51.3 | 94% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 295 | 18105 | FANCONI ANEMIA//ATAXIA TELANGIECTASIA//WERNER SYNDROME |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | NIJMEGEN BREAKAGE SYNDROME | Author keyword | 76 | 67% | 3% | 69 |
| 2 | 53BP1 | Author keyword | 66 | 56% | 3% | 81 |
| 3 | NBS1 | Author keyword | 66 | 59% | 3% | 75 |
| 4 | GAMMA H2AX | Author keyword | 53 | 32% | 6% | 137 |
| 5 | MRE11 | Author keyword | 45 | 52% | 3% | 61 |
| 6 | H2AX | Author keyword | 43 | 41% | 4% | 83 |
| 7 | MDC1 | Author keyword | 30 | 68% | 1% | 26 |
| 8 | NBS1 GENE | Author keyword | 30 | 100% | 1% | 12 |
| 9 | RNF168 | Author keyword | 27 | 78% | 1% | 18 |
| 10 | RNF8 | Author keyword | 23 | 65% | 1% | 22 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | NIJMEGEN BREAKAGE SYNDROME | 76 | 67% | 3% | 69 | Search NIJMEGEN+BREAKAGE+SYNDROME | Search NIJMEGEN+BREAKAGE+SYNDROME |
| 2 | 53BP1 | 66 | 56% | 3% | 81 | Search 53BP1 | Search 53BP1 |
| 3 | NBS1 | 66 | 59% | 3% | 75 | Search NBS1 | Search NBS1 |
| 4 | GAMMA H2AX | 53 | 32% | 6% | 137 | Search GAMMA+H2AX | Search GAMMA+H2AX |
| 5 | MRE11 | 45 | 52% | 3% | 61 | Search MRE11 | Search MRE11 |
| 6 | H2AX | 43 | 41% | 4% | 83 | Search H2AX | Search H2AX |
| 7 | MDC1 | 30 | 68% | 1% | 26 | Search MDC1 | Search MDC1 |
| 8 | NBS1 GENE | 30 | 100% | 1% | 12 | Search NBS1+GENE | Search NBS1+GENE |
| 9 | RNF168 | 27 | 78% | 1% | 18 | Search RNF168 | Search RNF168 |
| 10 | RNF8 | 23 | 65% | 1% | 22 | Search RNF8 | Search RNF8 |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | DOUBLE STRAND BREAKS | 127 | 12% | 43% | 1021 |
| 2 | MRE11 COMPLEX | 126 | 49% | 8% | 189 |
| 3 | HISTONE H2AX | 117 | 30% | 14% | 333 |
| 4 | ATM ACTIVATION | 99 | 49% | 6% | 147 |
| 5 | HISTONE H2AX PHOSPHORYLATION | 98 | 41% | 8% | 184 |
| 6 | DAMAGE RESPONSE | 89 | 26% | 12% | 290 |
| 7 | END RESECTION | 86 | 49% | 5% | 126 |
| 8 | 53BP1 | 78 | 52% | 5% | 107 |
| 9 | MDC1 | 71 | 63% | 3% | 71 |
| 10 | NIJMEGEN BREAKAGE SYNDROME | 68 | 34% | 7% | 160 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Double-strand break repair: 53BP1 comes into focus | 2014 | 49 | 116 | 69% |
| The ATM protein kinase: regulating the cellular response to genotoxic stress, and more | 2013 | 204 | 197 | 38% |
| Double-Strand Break End Resection and Repair Pathway Choice | 2011 | 292 | 181 | 52% |
| Playing the End Game: DNA Double-Strand Break Repair Pathway Choice | 2012 | 224 | 108 | 40% |
| Dynamics of DNA damage response proteins at DNA breaks: a focus on protein modifications | 2011 | 277 | 354 | 48% |
| The MRE11 complex: starting from the ends | 2011 | 194 | 171 | 62% |
| More than just a focus: The chromatin response to DNA damage and its role in genome integrity maintenance | 2011 | 182 | 101 | 62% |
| Chromatin Remodeling at DNA Double-Strand Breaks | 2013 | 90 | 99 | 58% |
| OPINION gamma H2AX and cancer | 2008 | 516 | 159 | 50% |
| The DNA Damage Response: Making It Safe to Play with Knives | 2010 | 822 | 230 | 25% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | RADIAT BIOL DNA REPAIR | 17 | 79% | 0.5% | 11 |
| 2 | GENOTOX ST S | 10 | 28% | 1.3% | 31 |
| 3 | BRANDER CANC | 8 | 17% | 1.8% | 42 |
| 4 | GENOME DAMAGE STABIL | 7 | 11% | 2.5% | 59 |
| 5 | GENOME REPAIR DYNAM | 6 | 38% | 0.5% | 12 |
| 6 | GENOME STABIL | 6 | 19% | 1.1% | 26 |
| 7 | BIOREGULAT CELLULAR PONSE | 4 | 75% | 0.1% | 3 |
| 8 | CHROMOSOME STABIL DYNAM GRP | 4 | 75% | 0.1% | 3 |
| 9 | HRICHTUNG BIOPHYS | 3 | 31% | 0.4% | 9 |
| 10 | MOL RADIAT BIOL | 3 | 13% | 1.0% | 23 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000179338 | DNA PK//DNA DEPENDENT PROTEIN KINASE//KU |
| 2 | 0.0000165194 | CHK1//RAD9//CLASPIN |
| 3 | 0.0000149916 | ATAXIA TELANGIECTASIA//ATM GENE//ATAXIA TELEANGIECTASIA |
| 4 | 0.0000102088 | RAD51//HOMOLOGOUS RECOMBINATION//RAD52 |
| 5 | 0.0000094840 | OLAPARIB//SYNTHETIC LETHALITY//PARP INHIBITOR |
| 6 | 0.0000066988 | AOA2//ARSACS//APRATAXIN |
| 7 | 0.0000057236 | HISTONE CHAPERONE//ASF1//CAF 1 |
| 8 | 0.0000052194 | WR HEARST MICROBIOL//TELOMERE//TRF2 |
| 9 | 0.0000048132 | PREDICTIVE ASSAYS//INDIVIDUAL RADIOSENSITIVITY//NORMAL TISSUE RESPONSE |
| 10 | 0.0000046895 | THERMORADIOSENSITIZATION//THERMAL RADIOSENSITIZATION//LONG DURATION MILD HYPERTHERMIA |