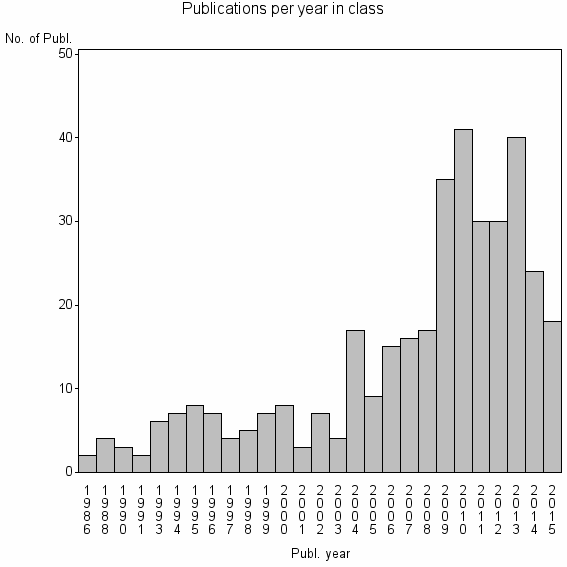
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 21192 | 369 | 48.4 | 89% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 356 | 16803 | PROTEOMICS//JOURNAL OF PROTEOME RESEARCH//PROTEIN IDENTIFICATION |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | SECRETOME | Author keyword | 16 | 17% | 23% | 86 |
| 2 | 90K MAC 2BP | Author keyword | 12 | 86% | 2% | 6 |
| 3 | 90K | Author keyword | 6 | 80% | 1% | 4 |
| 4 | GALECTIN 3 BINDING PROTEIN | Author keyword | 6 | 58% | 2% | 7 |
| 5 | 90K MAC 2 BINDING PROTEIN | Author keyword | 6 | 100% | 1% | 4 |
| 6 | MAC 2 BINDING PROTEIN | Author keyword | 5 | 55% | 2% | 6 |
| 7 | CANCER SECRETOME | Author keyword | 3 | 100% | 1% | 3 |
| 8 | LGALS3BP | Author keyword | 3 | 60% | 1% | 3 |
| 9 | CELL SECRETOME | Author keyword | 2 | 67% | 1% | 2 |
| 10 | COMPARATIVE PROTEOME PROFILING | Author keyword | 2 | 67% | 1% | 2 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | SECRETOME | 16 | 17% | 23% | 86 | Search SECRETOME | Search SECRETOME |
| 2 | 90K MAC 2BP | 12 | 86% | 2% | 6 | Search 90K+MAC+2BP | Search 90K+MAC+2BP |
| 3 | 90K | 6 | 80% | 1% | 4 | Search 90K | Search 90K |
| 4 | GALECTIN 3 BINDING PROTEIN | 6 | 58% | 2% | 7 | Search GALECTIN+3+BINDING+PROTEIN | Search GALECTIN+3+BINDING+PROTEIN |
| 5 | 90K MAC 2 BINDING PROTEIN | 6 | 100% | 1% | 4 | Search 90K+MAC+2+BINDING+PROTEIN | Search 90K+MAC+2+BINDING+PROTEIN |
| 6 | MAC 2 BINDING PROTEIN | 5 | 55% | 2% | 6 | Search MAC+2+BINDING+PROTEIN | Search MAC+2+BINDING+PROTEIN |
| 7 | CANCER SECRETOME | 3 | 100% | 1% | 3 | Search CANCER+SECRETOME | Search CANCER+SECRETOME |
| 8 | LGALS3BP | 3 | 60% | 1% | 3 | Search LGALS3BP | Search LGALS3BP |
| 9 | CELL SECRETOME | 2 | 67% | 1% | 2 | Search CELL+SECRETOME | Search CELL+SECRETOME |
| 10 | COMPARATIVE PROTEOME PROFILING | 2 | 67% | 1% | 2 | Search COMPARATIVE+PROTEOME+PROFILING | Search COMPARATIVE+PROTEOME+PROFILING |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MAC 2 BINDING PROTEIN | 43 | 62% | 12% | 45 |
| 2 | 90K | 12 | 58% | 4% | 14 |
| 3 | ANTIGEN 90K | 8 | 75% | 2% | 6 |
| 4 | 90K MAC 2 BP | 7 | 57% | 2% | 8 |
| 5 | CONDITIONED MEDIA | 6 | 24% | 6% | 22 |
| 6 | RING LIKE STRUCTURES | 5 | 54% | 2% | 7 |
| 7 | 90K MAC 2 BINDING PROTEIN | 4 | 67% | 1% | 4 |
| 8 | CANDIDATE SEROLOGICAL BIOMARKERS | 3 | 100% | 1% | 3 |
| 9 | MAC 2 BP | 3 | 100% | 1% | 3 |
| 10 | SERUM 90K ANTIGEN | 3 | 100% | 1% | 3 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Secretome proteomics for discovery of cancer biomarkers | 2010 | 81 | 99 | 49% |
| Conditioned media from cell lines: A complementary model to clinical specimens for the discovery of disease-specific biomarkers | 2011 | 45 | 69 | 41% |
| The cancer secretome, current status and opportunities in the lung, breast and colorectal cancer context | 2013 | 12 | 106 | 33% |
| Mapping of the secretome of primary isolates of mammalian cells, stem cells and derived cell lines | 2011 | 64 | 75 | 17% |
| The cancer cell secretome: A good source for discovering biomarkers? | 2010 | 69 | 75 | 23% |
| Sieving through the cancer secretome | 2013 | 4 | 99 | 46% |
| Advances in the proteomic investigation of the cell secretome | 2012 | 17 | 70 | 24% |
| Mass Spectrometry-Based Proteomics in Molecular Diagnostics: Discovery of Cancer Biomarkers Using Tissue Culture | 2013 | 15 | 101 | 13% |
| Approaches to the study of the cell secretome | 2007 | 72 | 34 | 18% |
| Methodologies to decipher the cell secretome | 2013 | 6 | 75 | 21% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | SECT BIOMED SCI | 1 | 25% | 1.4% | 5 |
| 2 | TUMOR PROFILING UNIT | 1 | 33% | 0.8% | 3 |
| 3 | CHANG GUNG PROTEOM CORE | 1 | 33% | 0.5% | 2 |
| 4 | JOSEPH WOLF LEBOVIC | 1 | 33% | 0.5% | 2 |
| 5 | ANALYT BIOTECHNOL SECT | 1 | 50% | 0.3% | 1 |
| 6 | AXON GROWTH REGENERAT GRP | 1 | 50% | 0.3% | 1 |
| 7 | BIOL SEQUENCE ANAL BIO | 1 | 50% | 0.3% | 1 |
| 8 | CATTEDRA ONCOL CLIN | 1 | 50% | 0.3% | 1 |
| 9 | DIPARTIMENTO MED SCI SALUTE V TIBERIO | 1 | 50% | 0.3% | 1 |
| 10 | DOERENKAMP ZBINDEN IN VITRO TOXICOL BIOMED | 1 | 50% | 0.3% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000152346 | GRIM 19//OLFACTOMEDIN 4//GW112 |
| 2 | 0.0000142097 | MAZUMDAR SHAW TRANSLAT//2 DIMENSIONAL LIQUID CHROMATOGRAPHY//BIOLOGICAL VARIATION ANALYSIS |
| 3 | 0.0000107400 | SELDI//SELDI TOF MS//LOW ABUNDANCE PROTEOME |
| 4 | 0.0000105102 | ALCAM//CD166//CD6 |
| 5 | 0.0000101590 | VASC PHYSIOPATHOL//MEDIA LAYER//CORONARY ATHEROSCLEROTIC HEART DISEASE |
| 6 | 0.0000091923 | CALUMENIN//BIOCHEM FUNCT PROTE//RETICULOCALBIN |
| 7 | 0.0000085557 | DMBT1//SALIVARY AGGLUTININ//GP 340 |
| 8 | 0.0000076924 | ENDOSIALIN//CD248//TEM1 |
| 9 | 0.0000075719 | LEUCINE RICH ALPHA 2 GLYCOPROTEIN 1//GLOBAL OLARS PROGRAM//LRG1 |
| 10 | 0.0000069254 | PEPTIDE IDENTIFICATION//JOURNAL OF PROTEOME RESEARCH//QUANTITATIVE PROTEOMICS |