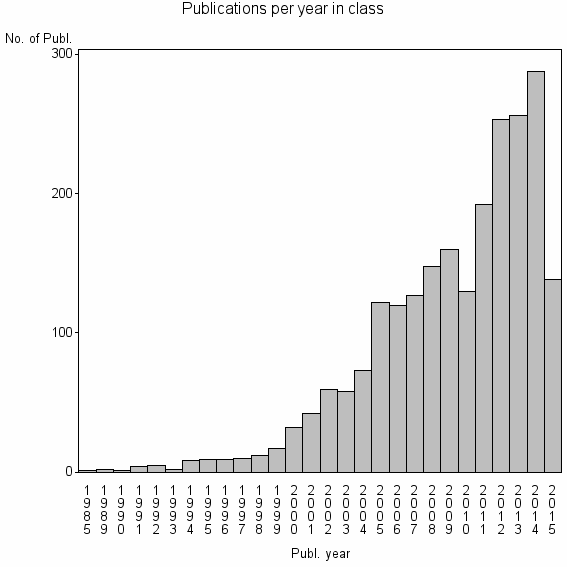
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 2131 | 2278 | 40.2 | 90% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 823 | 11253 | NON SMALL CELL LUNG CANCER//XRCC1//GEFITINIB |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | XRCC1 | Author keyword | 436 | 75% | 14% | 317 |
| 2 | XRCC3 | Author keyword | 210 | 83% | 5% | 118 |
| 3 | XRCC1 GENE | Author keyword | 130 | 97% | 2% | 36 |
| 4 | XPD | Author keyword | 89 | 54% | 5% | 115 |
| 5 | ERCC2 | Author keyword | 65 | 76% | 2% | 45 |
| 6 | X RAY REPAIR CROSS COMPLEMENTING GROUP 1 | Author keyword | 45 | 94% | 1% | 16 |
| 7 | HOGG1 | Author keyword | 42 | 51% | 3% | 58 |
| 8 | XRCC2 | Author keyword | 42 | 75% | 1% | 30 |
| 9 | ARG399GLN | Author keyword | 33 | 100% | 1% | 13 |
| 10 | X RAY REPAIR CROSS COMPLEMENTING GROUP 3 | Author keyword | 27 | 92% | 0% | 11 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | XRCC1 | 436 | 75% | 14% | 317 | Search XRCC1 | Search XRCC1 |
| 2 | XRCC3 | 210 | 83% | 5% | 118 | Search XRCC3 | Search XRCC3 |
| 3 | XRCC1 GENE | 130 | 97% | 2% | 36 | Search XRCC1+GENE | Search XRCC1+GENE |
| 4 | XPD | 89 | 54% | 5% | 115 | Search XPD | Search XPD |
| 5 | ERCC2 | 65 | 76% | 2% | 45 | Search ERCC2 | Search ERCC2 |
| 6 | X RAY REPAIR CROSS COMPLEMENTING GROUP 1 | 45 | 94% | 1% | 16 | Search X+RAY+REPAIR+CROSS+COMPLEMENTING+GROUP+1 | Search X+RAY+REPAIR+CROSS+COMPLEMENTING+GROUP+1 |
| 7 | HOGG1 | 42 | 51% | 3% | 58 | Search HOGG1 | Search HOGG1 |
| 8 | XRCC2 | 42 | 75% | 1% | 30 | Search XRCC2 | Search XRCC2 |
| 9 | ARG399GLN | 33 | 100% | 1% | 13 | Search ARG399GLN | Search ARG399GLN |
| 10 | X RAY REPAIR CROSS COMPLEMENTING GROUP 3 | 27 | 92% | 0% | 11 | Search X+RAY+REPAIR+CROSS+COMPLEMENTING+GROUP+3 | Search X+RAY+REPAIR+CROSS+COMPLEMENTING+GROUP+3 |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | DNA REPAIR GENES | 334 | 58% | 17% | 388 |
| 2 | XPD POLYMORPHISMS | 309 | 90% | 6% | 137 |
| 3 | XPD | 280 | 78% | 8% | 186 |
| 4 | ARG399GLN | 186 | 100% | 2% | 48 |
| 5 | XRCC1 | 180 | 54% | 10% | 230 |
| 6 | XRCC1 POLYMORPHISMS | 132 | 85% | 3% | 71 |
| 7 | ACID SUBSTITUTION VARIANTS | 125 | 76% | 4% | 88 |
| 8 | ARG399GLN POLYMORPHISM | 81 | 96% | 1% | 25 |
| 9 | CHROMOSOME 19Q132 3 | 73 | 96% | 1% | 23 |
| 10 | SER326CYS POLYMORPHISM | 71 | 71% | 3% | 57 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references |
% act. ref. to same field |
|---|---|---|---|---|
| X-ray repair cross-complementing group 1 (XRCC1) Arg399Gln polymorphism significantly associated with prostate cancer | 2015 | 1 | 40 | 75% |
| Association Between the XRCC6 Polymorphisms and Cancer Risks A Systematic Review and Meta-analysis | 2015 | 1 | 42 | 55% |
| Associations of Lys939Gln and Ala499Val polymorphisms of the XPC gene with cancer susceptibility: A meta-analysis | 2013 | 22 | 84 | 85% |
| Genetic polymorphisms in the base excision repair pathway and cancer risk: A HuGE review | 2005 | 329 | 93 | 83% |
| The association between the ERCC1/2 polymorphisms and the clinical outcomes of the platinum-based chemotherapy in non-small cell lung cancer (NSCLC): a systematic review and meta-analysis | 2014 | 5 | 61 | 82% |
| Polymorphisms in DNA repair genes and associations with cancer risk | 2002 | 789 | 84 | 56% |
| Polymorphism in the DNA repair enzyme XRCC1: Utility of current database and implications for human health risk assessment | 2011 | 36 | 90 | 83% |
| The effect of XPD/ERCC2 Lys751Gln polymorphism on acute leukemia risk: A systematic review and meta-analysis | 2014 | 3 | 40 | 93% |
| XRCC3 and XPD/ERCC2 single nucleotide polymorphisms and the risk of cancer: A HuGE review | 2006 | 128 | 23 | 78% |
| X-Ray Repair Cross-Complementing Group 1 (XRCC1) Genetic Polymorphisms and Gastric Cancer Risk: A HuGE Review and Meta-Analysis | 2011 | 37 | 34 | 62% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | TERRY FOX CANC | 16 | 34% | 1.7% | 38 |
| 2 | CLIN GYNAECOL SURG | 15 | 88% | 0.3% | 7 |
| 3 | CATHEDRAL MOTHERS CHILDS HLTH | 12 | 86% | 0.3% | 6 |
| 4 | ZAKLAD PATOMORFOL KLIN | 6 | 50% | 0.4% | 8 |
| 5 | MENOPAUSAL DIS | 5 | 60% | 0.3% | 6 |
| 6 | MOL EPIDEMIOL MED | 4 | 67% | 0.2% | 4 |
| 7 | MOL RADIOBIOL GRP | 4 | 75% | 0.1% | 3 |
| 8 | EASTERN MED GRP SHENYANG | 3 | 100% | 0.1% | 3 |
| 9 | MOL GENET EPIDEMIOL SECT | 3 | 45% | 0.2% | 5 |
| 10 | UNIT 1365 | 3 | 22% | 0.4% | 10 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000228063 | ERCC1//RRM1//MED OVARIAN CANC SECT |
| 2 | 0.0000139110 | G2 ASSAY//MUTAGEN SENSITIVITY//CHROMOSOMAL RADIOSENSITIVITY |
| 3 | 0.0000074171 | APE1//AP ENDONUCLEASE//APE1 REF 1 |
| 4 | 0.0000069351 | XERODERMA PIGMENTOSUM//COCKAYNE SYNDROME//NUCLEOTIDE EXCISION REPAIR |
| 5 | 0.0000068681 | DNA GLYCOSYLASE//8 OXOGUANINE//BASE EXCISION REPAIR |
| 6 | 0.0000066360 | P53 CODON 72//P53 POLYMORPHISM//CODON 72 POLYMORPHISM |
| 7 | 0.0000064927 | GSTM1//GSTT1//GSTP1 |
| 8 | 0.0000051323 | PREDICTIVE ASSAYS//INDIVIDUAL RADIOSENSITIVITY//NORMAL TISSUE RESPONSE |
| 9 | 0.0000050068 | CHEK2//PALB2//SERV CIRUGIA GEN ESPECIALIDADES |
| 10 | 0.0000047523 | FAMILIAL GLIOMA//RTEL1//EVANSTON KELLOGG CANC CARE |