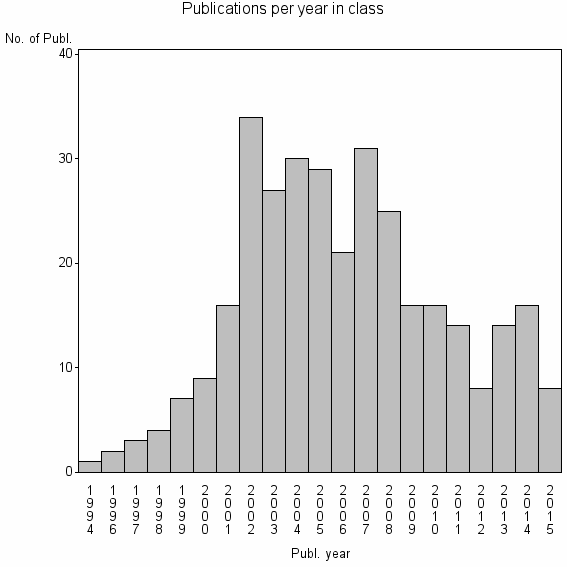
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 22261 | 331 | 38.1 | 89% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | CALPAIN 10 | Author keyword | 37 | 64% | 11% | 36 |
| 2 | MED CELLULAR MOL PHYSIOL BIOSTAT | Address | 21 | 90% | 3% | 9 |
| 3 | MOL GENET COMPLEX MONOGEN DISORDERS | Address | 10 | 63% | 3% | 10 |
| 4 | PSMD9 | Author keyword | 8 | 75% | 2% | 6 |
| 5 | CAPN10 | Author keyword | 5 | 47% | 2% | 8 |
| 6 | LIPGENE STUDY | Author keyword | 2 | 50% | 1% | 3 |
| 7 | 12Q24 | Author keyword | 1 | 50% | 1% | 2 |
| 8 | CAPN10 POLYMORPHISMS | Author keyword | 1 | 100% | 1% | 2 |
| 9 | NUTRIGEN GRP | Address | 1 | 11% | 3% | 10 |
| 10 | GENOMEWIDE SCAN | Author keyword | 1 | 30% | 1% | 3 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | CALPAIN 10 | 37 | 64% | 11% | 36 | Search CALPAIN+10 | Search CALPAIN+10 |
| 2 | PSMD9 | 8 | 75% | 2% | 6 | Search PSMD9 | Search PSMD9 |
| 3 | CAPN10 | 5 | 47% | 2% | 8 | Search CAPN10 | Search CAPN10 |
| 4 | LIPGENE STUDY | 2 | 50% | 1% | 3 | Search LIPGENE+STUDY | Search LIPGENE+STUDY |
| 5 | 12Q24 | 1 | 50% | 1% | 2 | Search 12Q24 | Search 12Q24 |
| 6 | CAPN10 POLYMORPHISMS | 1 | 100% | 1% | 2 | Search CAPN10+POLYMORPHISMS | Search CAPN10+POLYMORPHISMS |
| 7 | GENOMEWIDE SCAN | 1 | 30% | 1% | 3 | Search GENOMEWIDE+SCAN | Search GENOMEWIDE+SCAN |
| 8 | CALPAIN 10 GENE | 1 | 40% | 1% | 2 | Search CALPAIN+10+GENE | Search CALPAIN+10+GENE |
| 9 | LIPGENE | 1 | 23% | 1% | 3 | Search LIPGENE | Search LIPGENE |
| 10 | CALPAIN10 | 1 | 50% | 0% | 1 | Search CALPAIN10 | Search CALPAIN10 |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CAPN10 GENE | 24 | 91% | 3% | 10 |
| 2 | INDEPENDENT REPLICATION | 21 | 40% | 12% | 41 |
| 3 | PSMD9 | 14 | 100% | 2% | 7 |
| 4 | CAPN10 | 13 | 69% | 3% | 11 |
| 5 | CHROMOSOME 1Q21 Q24 | 12 | 46% | 6% | 19 |
| 6 | PROVIDES INDEPENDENT REPLICATION | 12 | 86% | 2% | 6 |
| 7 | CALPAIN 10 GENE | 11 | 49% | 5% | 17 |
| 8 | 12Q24 LOCUS | 11 | 100% | 2% | 6 |
| 9 | NORTHERN EUROPEAN CAUCASIANS | 11 | 78% | 2% | 7 |
| 10 | MELLITUS GENETICS FUSION | 10 | 52% | 4% | 13 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Role of Calpain-10 in the Development of Diabetes Mellitus and Its Complications | 2014 | 1 | 105 | 51% |
| Agreement among type 2 diabetes linkage studies but a poor correlation with results from genome-wide association studies | 2009 | 14 | 125 | 46% |
| A strategy to search for common obesity and type 2 diabetes genes | 2007 | 23 | 75 | 39% |
| Proteasome Modulator 9 and Depression in Type 2 Diabetes | 2012 | 3 | 42 | 50% |
| Catpain-10: from genome search to function | 2005 | 32 | 85 | 36% |
| Calpain-10 (NIDDM1) as a susceptibility gene for common type 2 diabetes | 2006 | 5 | 40 | 65% |
| The search for type 2 diabetes susceptibitity genes using whole-genome scans: an epidemiologist's perspective | 2002 | 23 | 27 | 56% |
| Challenges in identifying genetic variation affecting susceptibility to type 2 diabetes: Examples from studies of the calpain-10 gene | 2001 | 20 | 12 | 58% |
| Metabolic syndrome: Evidences for a personalized nutrition | 2012 | 8 | 83 | 13% |
| Genes and type 2 diabetes mellitus | 2005 | 17 | 87 | 30% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MED CELLULAR MOL PHYSIOL BIOSTAT | 21 | 90% | 2.7% | 9 |
| 2 | MOL GENET COMPLEX MONOGEN DISORDERS | 10 | 63% | 3.0% | 10 |
| 3 | NUTRIGEN GRP | 1 | 11% | 3.0% | 10 |
| 4 | UCD PUBL HLTH POPULAT SCI | 1 | 15% | 1.8% | 6 |
| 5 | LIPID ATHEROSCLEROSIS UNIT | 1 | 12% | 2.1% | 7 |
| 6 | UNIDAD REPROD GENET HUMANA | 1 | 33% | 0.6% | 2 |
| 7 | BIOL ANTHROPOL GRP | 1 | 50% | 0.3% | 1 |
| 8 | BUFFIELD MED | 1 | 50% | 0.3% | 1 |
| 9 | CARDIOVASC METAB ENDOCRINE CLIN | 1 | 50% | 0.3% | 1 |
| 10 | CHINESE HUMAN GENOME BEIJING | 1 | 50% | 0.3% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000262563 | TCF7L2//CDKAL1//SLC30A8 |
| 2 | 0.0000238099 | MODY//MATURITY ONSET DIABETES OF THE YOUNG//MATURITY ONSET DIABETES OF THE YOUNG MODY |
| 3 | 0.0000164255 | GENETIC EPIDEMIOLOGY//LINKAGE ANALYSIS//MODEL FREE LINKAGE ANALYSIS |
| 4 | 0.0000124933 | BETA3 ADRENERGIC RECEPTOR GENE//TRP64ARG POLYMORPHISM//BETA3 ADRENERGIC RECEPTOR |
| 5 | 0.0000115973 | USF1//UPSTREAM TRANSCRIPTION FACTOR 1//USF |
| 6 | 0.0000111383 | ENPP1//PLASMA CELL MEMBRANE GLYCOPROTEIN 1//PC 1 |
| 7 | 0.0000102642 | CALPAIN//CALPASTATIN//CALPAIN INHIBITOR |
| 8 | 0.0000091947 | GLUCOSE EFFECTIVENESS//INTRAVENOUS GLUCOSE TOLERANCE TEST//MINIMAL MODEL |
| 9 | 0.0000083960 | PRO12ALA POLYMORPHISM//PRO12ALA//PPAR GAMMA 2 |
| 10 | 0.0000062255 | NSY MOUSE//NEW ZEALAND OBESE MOUSE//MOUSE GENOM OURCE |