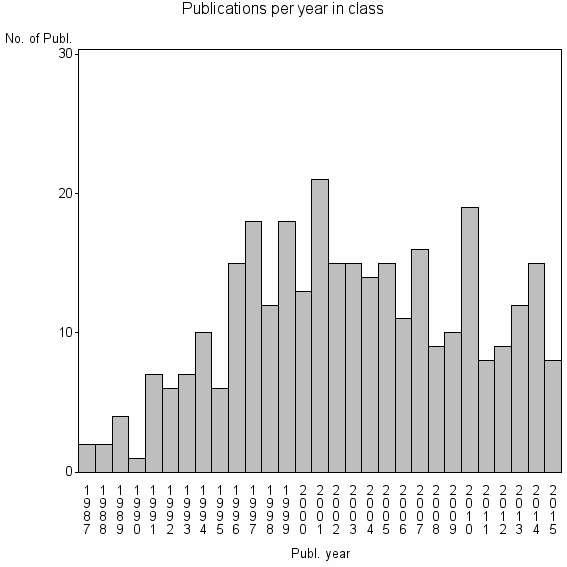
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 22650 | 318 | 42.7 | 90% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | CUTL1 | Author keyword | 18 | 83% | 3% | 10 |
| 2 | SMAR1 | Author keyword | 13 | 69% | 3% | 11 |
| 3 | CDP CUX | Author keyword | 9 | 83% | 2% | 5 |
| 4 | CCAAT DISPLACEMENT PROTEIN | Author keyword | 8 | 62% | 3% | 8 |
| 5 | CUX1 | Author keyword | 6 | 48% | 3% | 10 |
| 6 | CDP CUT | Author keyword | 6 | 80% | 1% | 4 |
| 7 | ST7 | Author keyword | 6 | 71% | 2% | 5 |
| 8 | CUX 1 | Author keyword | 4 | 67% | 1% | 4 |
| 9 | CUX | Author keyword | 3 | 45% | 2% | 5 |
| 10 | HBP1 | Author keyword | 3 | 33% | 2% | 7 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | CUTL1 | 18 | 83% | 3% | 10 | Search CUTL1 | Search CUTL1 |
| 2 | SMAR1 | 13 | 69% | 3% | 11 | Search SMAR1 | Search SMAR1 |
| 3 | CDP CUX | 9 | 83% | 2% | 5 | Search CDP+CUX | Search CDP+CUX |
| 4 | CCAAT DISPLACEMENT PROTEIN | 8 | 62% | 3% | 8 | Search CCAAT+DISPLACEMENT+PROTEIN | Search CCAAT+DISPLACEMENT+PROTEIN |
| 5 | CUX1 | 6 | 48% | 3% | 10 | Search CUX1 | Search CUX1 |
| 6 | CDP CUT | 6 | 80% | 1% | 4 | Search CDP+CUT | Search CDP+CUT |
| 7 | ST7 | 6 | 71% | 2% | 5 | Search ST7 | Search ST7 |
| 8 | CUX 1 | 4 | 67% | 1% | 4 | Search CUX+1 | Search CUX+1 |
| 9 | CUX | 3 | 45% | 2% | 5 | Search CUX | Search CUX |
| 10 | HBP1 | 3 | 33% | 2% | 7 | Search HBP1 | Search HBP1 |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CCAAT DISPLACEMENT PROTEIN | 52 | 47% | 26% | 82 |
| 2 | CUT HOMEODOMAIN PROTEIN | 46 | 76% | 10% | 32 |
| 3 | SENSORY ORGAN IDENTITY | 24 | 50% | 11% | 34 |
| 4 | CELL TYPE SPECIFICATION | 18 | 51% | 8% | 25 |
| 5 | DROSOPHILA WING MARGIN | 17 | 100% | 3% | 8 |
| 6 | DROSOPHILA CUT | 15 | 68% | 4% | 13 |
| 7 | FACTOR HINF D | 14 | 40% | 8% | 27 |
| 8 | CDP CUX TRANSCRIPTION FACTOR | 11 | 67% | 3% | 10 |
| 9 | CUT REPEATS | 9 | 55% | 3% | 11 |
| 10 | CUX CDP HOMEOPROTEIN | 8 | 100% | 2% | 5 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Role of the multifunctional CDP/Cut/Cux homeodomain transcription factor in regulating differentiation, cell growth and development | 2001 | 155 | 113 | 48% |
| CUX1 transcription factors: From biochemical activities and cell-based assays to mouse models and human diseases | 2012 | 15 | 92 | 48% |
| Cutl1: A potential target for cancer therapy | 2013 | 1 | 35 | 89% |
| CUX1, a haploinsufficient tumour suppressor gene overexpressed in advanced cancers | 2014 | 1 | 79 | 61% |
| Regulation of cell proliferation and differentiation in the kidney | 2009 | 5 | 102 | 52% |
| Contribution of CDP/Cux, a transcription factor, to cell cycle progression | 2007 | 2 | 55 | 56% |
| -7/7q-syndrome in myeloid-lineage hematopoietic malignancies: attempts to understand this complex disease entity | 2015 | 1 | 208 | 14% |
| Identification of genetic loci that interact with cut during Drosophila wing-margin development | 2005 | 11 | 104 | 27% |
| The HBP1 transcriptional repressor and the p38 MAP kinase: unlikely partners in G1 regulation and tumor suppression | 2004 | 42 | 84 | 14% |
| Cux/CDP homeoprotein is a component of NF-mu NR and represses the immunoglobulin heavy chain intronic enhancer by antagonizing the bright transcription activator | 1999 | 56 | 95 | 17% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PROGRAM CELL MOL NUTR | 2 | 67% | 0.6% | 2 |
| 2 | SECT PEDIAT HEMATOL ONCOL BIOCHEM MOL BIOL | 1 | 50% | 0.6% | 2 |
| 3 | CARDIOLTIANJIN ION MOL FUNCT CARDIO | 1 | 50% | 0.3% | 1 |
| 4 | HEDLEY ATKINS CANC UK | 1 | 50% | 0.3% | 1 |
| 5 | INSERMU EQUIPE CHROMATINE EXP S GENES 309 | 1 | 50% | 0.3% | 1 |
| 6 | POST MED EDUC IPGMER | 1 | 50% | 0.3% | 1 |
| 7 | BALTIMORE VA HOSP | 0 | 33% | 0.3% | 1 |
| 8 | BIOL MOL CELLULAIRE DIFFERENTIAT | 0 | 33% | 0.3% | 1 |
| 9 | CANC CYTOGENET SECT | 0 | 33% | 0.3% | 1 |
| 10 | CATHCART | 0 | 33% | 0.3% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000132386 | JUVENILE MYELOMONOCYTIC LEUKEMIA//JMML//JUVENILE MYELOMONOCYTIC LEUKAEMIA |
| 2 | 0.0000089316 | HISTONE GENE EXPRESSION//HISTONE MRNA//SLBP |
| 3 | 0.0000078423 | SATB1//2Q DELETION//SATB2 |
| 4 | 0.0000074163 | DREF//INSECT BIOMED//DNA REPLICATION RELATED GENE |
| 5 | 0.0000072145 | ACHAETE SCUTE//GENET 240//ENHANCER OF SPLIT |
| 6 | 0.0000062042 | CLIN BREAST CARE PROJECT//16Q243//CHROMOSOMAL SCANNING |
| 7 | 0.0000053309 | HUMAN CYTOGENET TOXICOL GENET//C KI RAS ONCOGENE//JOAO BARROS BARRETO UNIV HOSP |
| 8 | 0.0000053193 | SPLENIC MARGINAL ZONE LYMPHOMA//NODAL MARGINAL ZONE LYMPHOMA//MONOCYTOID B CELL LYMPHOMA |
| 9 | 0.0000051422 | HMGA1//HMGA2//HMGIY |
| 10 | 0.0000049280 | CTCF//CONTROL GENET PROC//ENHANCER BLOCKING |