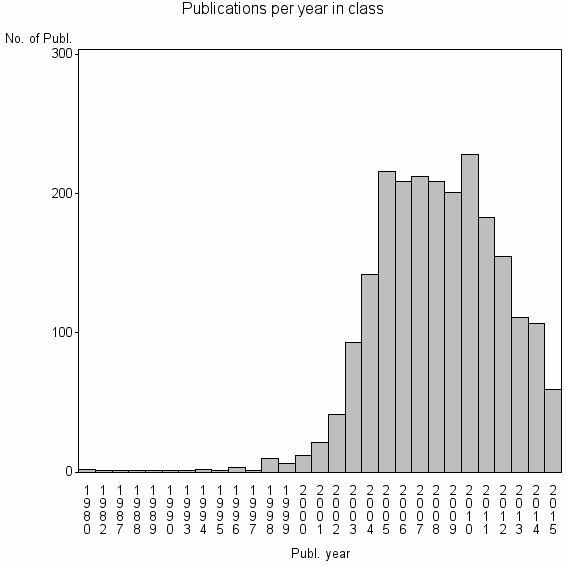
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 2270 | 2230 | 40.9 | 84% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 356 | 16803 | PROTEOMICS//JOURNAL OF PROTEOME RESEARCH//PROTEIN IDENTIFICATION |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | SELDI | Author keyword | 82 | 54% | 5% | 105 |
| 2 | SELDI TOF MS | Author keyword | 57 | 42% | 5% | 105 |
| 3 | LOW ABUNDANCE PROTEOME | Author keyword | 35 | 73% | 1% | 27 |
| 4 | SURFACE ENHANCED LASER DESORPTION IONIZATION | Author keyword | 29 | 61% | 1% | 31 |
| 5 | SELDI TOF | Author keyword | 25 | 45% | 2% | 42 |
| 6 | SERUM PROTEOME | Author keyword | 22 | 66% | 1% | 21 |
| 7 | SURFACE ENHANCED LASER DESORPTION IONIZATION TIME OF FLIGHT MASS SPECTROMETRY | Author keyword | 22 | 60% | 1% | 24 |
| 8 | HEXAPEPTIDE LIGANDS | Author keyword | 22 | 81% | 1% | 13 |
| 9 | COMBINATORIAL PEPTIDE LIGAND LIBRARIES | Author keyword | 21 | 71% | 1% | 17 |
| 10 | CLIN PROTEOM PROGRAM | Address | 17 | 59% | 1% | 19 |
Web of Science journal categories |
Author Key Words |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PROTEOMIC PATTERNS | 128 | 51% | 8% | 179 |
| 2 | SELDI TOF MS | 109 | 56% | 6% | 134 |
| 3 | ENHANCED LASER DESORPTION | 108 | 79% | 3% | 70 |
| 4 | HOST RESPONSE PROTEINS | 61 | 95% | 1% | 20 |
| 5 | BIOMARKER DISCOVERY | 54 | 20% | 11% | 246 |
| 6 | HUMAN PLASMA PROTEOME | 52 | 35% | 5% | 121 |
| 7 | SELDI TOF | 48 | 61% | 2% | 51 |
| 8 | CLINICAL PROTEOMICS | 47 | 41% | 4% | 87 |
| 9 | LASER DESORPTION IONIZATION TIME | 35 | 28% | 5% | 106 |
| 10 | PLASMA PROTEOME | 31 | 28% | 4% | 93 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Combinatorial Peptide Ligand Libraries as a "Trojan Horse" in Deep Discovery Proteomics | 2015 | 1 | 68 | 54% |
| Liquid-phase-based separation systems for depletion, prefractionation and enrichment of proteins in biological fluids and matrices for in-depth proteomics analysis - An update covering the period 2008-2011 | 2012 | 40 | 93 | 43% |
| Liquid phase based separation systems for depletion, prefractionation, and enrichment of proteins in biological fluids and matrices for in-depth proteomics analysis-An update covering the period 2011-2014 | 2015 | 2 | 60 | 27% |
| Mining the plasma proteome for cancer biomarkers | 2008 | 346 | 49 | 22% |
| High-abundance polypeptides of the human plasma proteome comprising the top 4 logs of polypeptide abundance | 2008 | 96 | 27 | 63% |
| Clinical proteomics: searching for better tumour markers with SELDI-TOF mass spectrometry | 2006 | 125 | 59 | 76% |
| Biological and methodical challenges of blood-based proteomics in the field of neurological research | 2013 | 17 | 164 | 46% |
| Opportunities and limitations of SELDI-TOF-MS in biomedical research: practical advices | 2007 | 90 | 78 | 64% |
| Challenges for Biomarker Discovery in Body Fluids Using SELDI-TOF-MS | 2010 | 34 | 123 | 64% |
| Proteomic Approaches to the Discovery of Cancer Biomarkers for Early Detection and Personalized Medicine | 2013 | 23 | 38 | 32% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CLIN PROTEOM PROGRAM | 17 | 59% | 0.9% | 19 |
| 2 | CUCA | 10 | 46% | 0.7% | 16 |
| 3 | CHEM MAT CHEM ENGN GIULIO NATTA | 10 | 16% | 2.5% | 56 |
| 4 | OVARIAN CANC EARLY DETECT PROGRAM | 8 | 70% | 0.3% | 7 |
| 5 | BIOMED PROTEOM PROGRAM | 8 | 60% | 0.4% | 9 |
| 6 | CLIN PROTE REFERENCE | 8 | 100% | 0.2% | 5 |
| 7 | FDA NCI CLIN PROTEOM PROGRAM | 7 | 67% | 0.3% | 6 |
| 8 | MICROBIOL MOL CELL BIOL | 6 | 15% | 1.7% | 39 |
| 9 | VIRGINIA PROSTATE | 6 | 33% | 0.7% | 16 |
| 10 | BIOMED PROTEOM | 6 | 50% | 0.4% | 9 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000218217 | URINARY PROTEOMICS//URINARY PROTEOME//URINE PROTEOME |
| 2 | 0.0000167791 | AMNIOTIC FLUID SUPERNATANT//CERVICAL VAGINAL FLUID//UNITE MED MATERNO FOETALE |
| 3 | 0.0000155385 | SISCAPA//PSAQ//BIOMARKER VERIFICATION |
| 4 | 0.0000143895 | CALUMENIN//BIOCHEM FUNCT PROTE//RETICULOCALBIN |
| 5 | 0.0000122480 | CIBER BBN SALUD CARLOS 3//RIER RED INFLAMAC ENFERMEDADES REUMAT//PROTEORED ISCIII PROTE GRP |
| 6 | 0.0000110136 | HUMAN BRONCHOALVEOLAR LAVAGE FLUID PROTEINS//CALCYPHOSINE//FUNCT PROTE SECT |
| 7 | 0.0000107400 | SECRETOME//90K MAC 2BP//90K |
| 8 | 0.0000104003 | MAZUMDAR SHAW TRANSLAT//2 DIMENSIONAL LIQUID CHROMATOGRAPHY//BIOLOGICAL VARIATION ANALYSIS |
| 9 | 0.0000097893 | LEUCINE RICH ALPHA 2 GLYCOPROTEIN 1//GLOBAL OLARS PROGRAM//LRG1 |
| 10 | 0.0000094398 | VASC PHYSIOPATHOL//MEDIA LAYER//CORONARY ATHEROSCLEROTIC HEART DISEASE |