Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
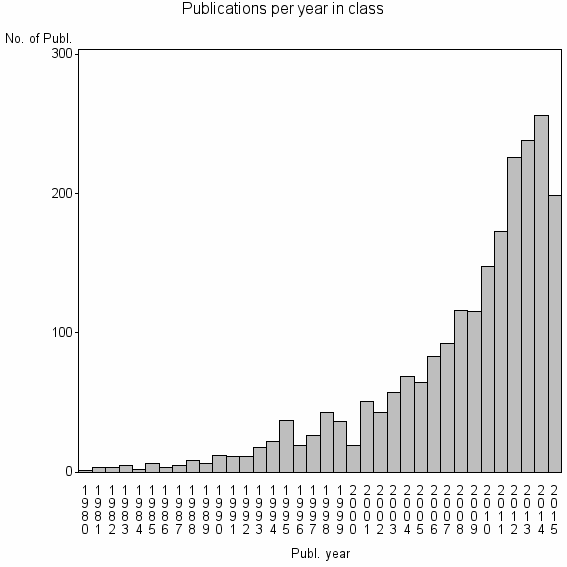
| 2284 | 2226 | 51.7 | 93% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | EZH2 | Author keyword | 248 | 62% | 11% | 255 |
| 2 | POLYCOMB | Author keyword | 160 | 52% | 10% | 218 |
| 3 | POLYCOMB GROUP | Author keyword | 119 | 72% | 4% | 92 |
| 4 | BMI 1 | Author keyword | 94 | 62% | 4% | 96 |
| 5 | BMI1 | Author keyword | 85 | 61% | 4% | 91 |
| 6 | POLYCOMB GROUP GENES | Author keyword | 53 | 85% | 1% | 28 |
| 7 | ENHANCER OF ZESTE HOMOLOG 2 | Author keyword | 52 | 82% | 1% | 31 |
| 8 | POLYHOMEOTIC | Author keyword | 41 | 90% | 1% | 18 |
| 9 | RING1B | Author keyword | 38 | 89% | 1% | 17 |
| 10 | PLEIOHOMEOTIC | Author keyword | 27 | 92% | 0% | 11 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | EZH2 | 248 | 62% | 11% | 255 | Search EZH2 | Search EZH2 |
| 2 | POLYCOMB | 160 | 52% | 10% | 218 | Search POLYCOMB | Search POLYCOMB |
| 3 | POLYCOMB GROUP | 119 | 72% | 4% | 92 | Search POLYCOMB+GROUP | Search POLYCOMB+GROUP |
| 4 | BMI 1 | 94 | 62% | 4% | 96 | Search BMI+1 | Search BMI+1 |
| 5 | BMI1 | 85 | 61% | 4% | 91 | Search BMI1 | Search BMI1 |
| 6 | POLYCOMB GROUP GENES | 53 | 85% | 1% | 28 | Search POLYCOMB+GROUP+GENES | Search POLYCOMB+GROUP+GENES |
| 7 | ENHANCER OF ZESTE HOMOLOG 2 | 52 | 82% | 1% | 31 | Search ENHANCER+OF+ZESTE+HOMOLOG+2 | Search ENHANCER+OF+ZESTE+HOMOLOG+2 |
| 8 | POLYHOMEOTIC | 41 | 90% | 1% | 18 | Search POLYHOMEOTIC | Search POLYHOMEOTIC |
| 9 | RING1B | 38 | 89% | 1% | 17 | Search RING1B | Search RING1B |
| 10 | PLEIOHOMEOTIC | 27 | 92% | 0% | 11 | Search PLEIOHOMEOTIC | Search PLEIOHOMEOTIC |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | HISTONE METHYLTRANSFERASE ACTIVITY | 239 | 62% | 11% | 247 |
| 2 | GROUP PROTEIN EZH2 | 158 | 57% | 8% | 186 |
| 3 | CHROMOSOME BINDING SITES | 146 | 87% | 3% | 72 |
| 4 | GROUP PROTEINS | 112 | 53% | 7% | 146 |
| 5 | POSTERIOR SEX COMBS | 105 | 89% | 2% | 48 |
| 6 | POLYCOMB GROUP PROTEINS | 97 | 42% | 8% | 181 |
| 7 | DEVELOPMENTAL REGULATORS | 97 | 36% | 10% | 220 |
| 8 | POLYCOMB | 86 | 31% | 11% | 234 |
| 9 | ZESTE HOMOLOG 2 | 79 | 71% | 3% | 64 |
| 10 | MYC TRANSGENIC MICE | 78 | 50% | 5% | 111 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| The Polycomb complex PRC2 and its mark in life | 2011 | 598 | 96 | 65% |
| Occupying Chromatin: Polycomb Mechanisms for Getting to Genomic Targets, Stopping Transcriptional Traffic, and Staying Put | 2013 | 144 | 151 | 75% |
| Mechanisms of Polycomb gene silencing: knowns and unknowns | 2009 | 559 | 167 | 67% |
| Transcriptional regulation by Polycomb group proteins | 2013 | 71 | 110 | 75% |
| Genome regulation by polycomb and trithorax proteins | 2007 | 658 | 76 | 55% |
| Polycomb silencing mechanisms and the management of genomic programmes | 2007 | 469 | 143 | 65% |
| The role of EZH2 in tumour progression | 2012 | 101 | 32 | 75% |
| What are memories made of ? How Polycomb and Trithorax proteins mediate epigenetic memory | 2014 | 24 | 171 | 62% |
| A new world of Polycombs: unexpected partnerships and emerging functions | 2013 | 41 | 140 | 80% |
| Roles of the EZH2 histone methyltransferase in cancer epigenetics | 2008 | 313 | 125 | 68% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CANC EPIGENET DISCOVERY PERFORMANCE UNIT | 17 | 79% | 0.5% | 11 |
| 2 | EQUIPE CHROMATINE DEV | 6 | 80% | 0.2% | 4 |
| 3 | DEV EPIGENET GRP | 4 | 41% | 0.3% | 7 |
| 4 | DEV BIOL MED | 4 | 46% | 0.3% | 6 |
| 5 | SURG ONCOL DEV THER EUT | 3 | 57% | 0.2% | 4 |
| 6 | COMPREHENS CANC 4217 | 3 | 50% | 0.2% | 4 |
| 7 | BIOMED SCI BIOCHEM GENET | 2 | 67% | 0.1% | 2 |
| 8 | CENT PROTE ILUBIQUITIN PROTEOLYSIS GRP | 2 | 67% | 0.1% | 2 |
| 9 | HEMATOL LEUKEMIA CELL BANK QUEBEC | 2 | 67% | 0.1% | 2 |
| 10 | ICFM | 2 | 67% | 0.1% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000129553 | CTCF//CONTROL GENET PROC//ENHANCER BLOCKING |
| 2 | 0.0000127952 | HISTONE DEMETHYLASE//HISTONE DEMETHYLATION//LSD1 |
| 3 | 0.0000080651 | YY1//YIN YANG 1//GON4L |
| 4 | 0.0000079118 | HP1//POSITION EFFECT VARIEGATION//HETEROCHROMATIN |
| 5 | 0.0000069192 | BAP1//BRCA1 ASSOCIATED PROTEIN 1//SETD2 |
| 6 | 0.0000053404 | HOX//HOX GENES//PROBOSCIPEDIA |
| 7 | 0.0000052791 | ARID3B//JUMONJI//AT RICH INTERACTION DOMAIN |
| 8 | 0.0000050781 | CHIP SEQ//PEAK CALLING//DNASE SEQ |
| 9 | 0.0000046484 | SWI SNF//BRG1//ISWI |
| 10 | 0.0000046287 | MLL//MLL GENE//11Q23 |