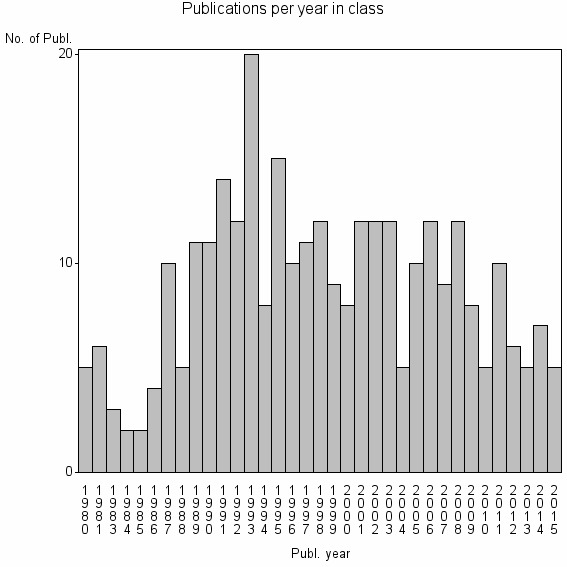
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 22920 | 308 | 27.7 | 71% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 501 | 14604 | RNA POLYMERASE//SITE SPECIFIC RECOMBINATION//H NS |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | RETARDATION ANALYSIS | Author keyword | 6 | 80% | 1% | 4 |
| 2 | NONSPECIFIC PROTEIN DNA COMPLEXES | Author keyword | 4 | 75% | 1% | 3 |
| 3 | GEL CAGE | Author keyword | 3 | 100% | 1% | 3 |
| 4 | PEDIAT DPMSC | Address | 2 | 67% | 1% | 2 |
| 5 | MOBILITY SHIFT ASSAY | Author keyword | 2 | 21% | 3% | 8 |
| 6 | DNA OLIGOMER SEPARATION | Author keyword | 1 | 100% | 1% | 2 |
| 7 | DNA AFFINITY CHROMATOGRAPHY | Author keyword | 1 | 40% | 1% | 2 |
| 8 | CY5 DYE | Author keyword | 1 | 50% | 0% | 1 |
| 9 | ECORI RESTRICTION ENDONUCLEASE | Author keyword | 1 | 50% | 0% | 1 |
| 10 | ENDOGENOUS SIGNALS | Author keyword | 1 | 50% | 0% | 1 |
Web of Science journal categories |
Author Key Words |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | DNA AFFINITY CHROMATOGRAPHY | 13 | 80% | 3% | 8 |
| 2 | MOBILITY SHIFT ASSAY | 5 | 24% | 6% | 19 |
| 3 | SEPHAROSE COLUMNS | 2 | 67% | 1% | 2 |
| 4 | SPHI | 2 | 67% | 1% | 2 |
| 5 | CATALYTIC CHROMATOGRAPHY | 1 | 100% | 1% | 2 |
| 6 | CDNA SILICA | 1 | 100% | 1% | 2 |
| 7 | OLIGODT30 LATEX | 1 | 100% | 1% | 2 |
| 8 | EMSA | 1 | 40% | 1% | 2 |
| 9 | LACTOSE PROMOTER | 1 | 27% | 1% | 3 |
| 10 | CLEAVABLE BIOTINYLATED NUCLEOTIDE | 1 | 50% | 0% | 1 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Methods for proteomic analysis of transcription factors | 2009 | 22 | 48 | 46% |
| Electrophoretic mobility shift assay (EMSA) for detecting protein-nucleic acid interactions | 2007 | 193 | 93 | 27% |
| Theory and practice of gel electrophoresis of interacting macromolecules | 1996 | 36 | 32 | 31% |
| Standard In Vitro Assays for Protein-Nucleic Acid Interactions - Gel Shift Assays for RNA and DNA Binding | 2014 | 0 | 5 | 80% |
| DNA affinity chromatography of transcription factors | 2001 | 32 | 129 | 26% |
| ELECTROPHORETIC MOBILITY SHIFT ASSAY | 1995 | 59 | 14 | 14% |
| MEASUREMENT AND ANALYSIS OF EQUILIBRIUM BINDING TITRATIONS: A BEGINNER'S GUIDE | 2011 | 1 | 3 | 33% |
| Mass spectrometry-based identification of proteins interacting with nucleic acids | 2013 | 3 | 106 | 16% |
| GEL RETARDATION | 1991 | 306 | 11 | 45% |
| Modem tools for identification of nucleic acid-binding proteins | 2008 | 11 | 59 | 15% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PEDIAT DPMSC | 2 | 67% | 0.6% | 2 |
| 2 | MCKUSICK NATHANS INS GENET MED | 1 | 50% | 0.3% | 1 |
| 3 | MSU FUNCT GENOM CORE IL | 1 | 50% | 0.3% | 1 |
| 4 | RCMI PROTE CORE | 1 | 50% | 0.3% | 1 |
| 5 | EQUIPE SCI SEPERAT BIOPHARMACEUT 2SB | 0 | 33% | 0.3% | 1 |
| 6 | GOFORSYS UNIT SYST BIOL | 0 | 33% | 0.3% | 1 |
| 7 | KYOTO NANOTECHNOL CLUSTER | 0 | 33% | 0.3% | 1 |
| 8 | NUCLEAR SIGNALLING | 0 | 33% | 0.3% | 1 |
| 9 | BIOCHEM CELL BIOL URBC | 0 | 17% | 0.6% | 2 |
| 10 | PROT BIOMARKERS CO | 0 | 25% | 0.3% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000134355 | USF1//UPSTREAM TRANSCRIPTION FACTOR 1//USF |
| 2 | 0.0000114046 | TRP REPRESSOR//LAC REPRESSOR//LACTOSE REPRESSOR |
| 3 | 0.0000083503 | CYCLIC AMP RECEPTOR PROTEIN//CATABOLITE ACTIVATOR PROTEIN CAP//CAMP RECEPTOR PROTEIN |
| 4 | 0.0000070383 | AD LACZ//CHICKEN EMBRYO FIBROBLAST CULTURE//SURFACE PROTEIN BIOTINYLATION |
| 5 | 0.0000069852 | GRHL2//GRAINYHEAD//GRAINY HEAD |
| 6 | 0.0000065654 | NUCLEAR FACTOR 1//NUCLEAR FACTOR I//DEV GENOM GRP |
| 7 | 0.0000058906 | SEQUENCING BY HYBRIDIZATION//DNA SEQUENCING WITH ERRORS//GAPPED PROBES |
| 8 | 0.0000054351 | WEAK AFFINITY//HPLAC//WEAK AFFINITY CHROMATOGRAPHY WAC |
| 9 | 0.0000053429 | NARROW ESCAPE//DEN DANS DM2S SERMA LTSD//NANOMED MECH BIOL |
| 10 | 0.0000053136 | BIOMIMETIC DYE//TRIAZINE DYE//BEAD CELLULOSE |