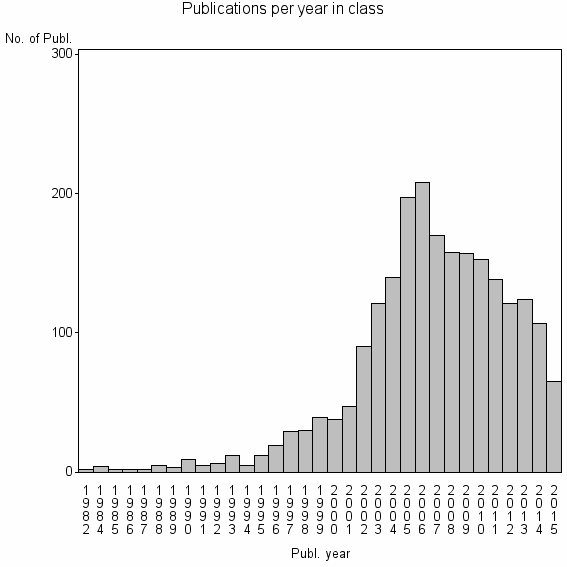
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 2302 | 2220 | 37.7 | 79% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 238 | 19563 | BIOINFORMATICS//BMC BIOINFORMATICS//GENOME RESEARCH |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | MOTIF DISCOVERY | Author keyword | 30 | 42% | 2% | 55 |
| 2 | CLOSEST STRING PROBLEM | Author keyword | 18 | 83% | 0% | 10 |
| 3 | MOTIF FINDING | Author keyword | 14 | 47% | 1% | 22 |
| 4 | TRANSCRIPTION FACTOR BINDING SITES | Author keyword | 14 | 31% | 2% | 37 |
| 5 | UKG | Address | 12 | 75% | 0% | 9 |
| 6 | BIOINFORMAT GENOMES EAUX BIGRE | Address | 10 | 52% | 1% | 14 |
| 7 | TRANSCRIPTION FACTOR BINDING SITE | Author keyword | 10 | 27% | 1% | 32 |
| 8 | POSITION WEIGHT MATRIX | Author keyword | 9 | 47% | 1% | 14 |
| 9 | CLOSEST SUBSTRING | Author keyword | 6 | 80% | 0% | 4 |
| 10 | BIOINFORMAT INTELLIGENT COMP | Address | 6 | 71% | 0% | 5 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | MOTIF DISCOVERY | 30 | 42% | 2% | 55 | Search MOTIF+DISCOVERY | Search MOTIF+DISCOVERY |
| 2 | CLOSEST STRING PROBLEM | 18 | 83% | 0% | 10 | Search CLOSEST+STRING+PROBLEM | Search CLOSEST+STRING+PROBLEM |
| 3 | MOTIF FINDING | 14 | 47% | 1% | 22 | Search MOTIF+FINDING | Search MOTIF+FINDING |
| 4 | TRANSCRIPTION FACTOR BINDING SITES | 14 | 31% | 2% | 37 | Search TRANSCRIPTION+FACTOR+BINDING+SITES | Search TRANSCRIPTION+FACTOR+BINDING+SITES |
| 5 | TRANSCRIPTION FACTOR BINDING SITE | 10 | 27% | 1% | 32 | Search TRANSCRIPTION+FACTOR+BINDING+SITE | Search TRANSCRIPTION+FACTOR+BINDING+SITE |
| 6 | POSITION WEIGHT MATRIX | 9 | 47% | 1% | 14 | Search POSITION+WEIGHT+MATRIX | Search POSITION+WEIGHT+MATRIX |
| 7 | CLOSEST SUBSTRING | 6 | 80% | 0% | 4 | Search CLOSEST+SUBSTRING | Search CLOSEST+SUBSTRING |
| 8 | PROMOTER PREDICTION | 5 | 33% | 1% | 12 | Search PROMOTER+PREDICTION | Search PROMOTER+PREDICTION |
| 9 | DE NOVO MOTIF DISCOVERY | 4 | 75% | 0% | 3 | Search DE+NOVO+MOTIF+DISCOVERY | Search DE+NOVO+MOTIF+DISCOVERY |
| 10 | IRREDUNDANT MOTIF | 4 | 75% | 0% | 3 | Search IRREDUNDANT+MOTIF | Search IRREDUNDANT+MOTIF |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | FACTOR BINDING SITES | 163 | 33% | 19% | 414 |
| 2 | COREGULATED GENES | 45 | 68% | 2% | 39 |
| 3 | CIS REGULATORY MODULES | 40 | 50% | 3% | 58 |
| 4 | UNALIGNED DNA FRAGMENTS | 38 | 93% | 1% | 14 |
| 5 | COMPUTATIONAL DETECTION | 34 | 93% | 1% | 13 |
| 6 | MOTIF DISCOVERY | 26 | 41% | 2% | 49 |
| 7 | STATISTICAL OVERREPRESENTATION | 23 | 86% | 1% | 12 |
| 8 | EXPECTATION MAXIMIZATION | 20 | 31% | 2% | 54 |
| 9 | FINDING MOTIFS | 20 | 67% | 1% | 18 |
| 10 | NONCODING SEQUENCES | 16 | 22% | 3% | 63 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Determining the specificity of protein-DNA interactions | 2010 | 92 | 55 | 60% |
| Applied bioinformatics for the identification of regulatory elements | 2004 | 533 | 80 | 53% |
| Absence of a simple code: how transcription factors read the genome | 2014 | 17 | 234 | 26% |
| Eukaryotic transcription factor binding sites - modeling and integrative search methods | 2008 | 54 | 67 | 78% |
| MAPPER: a search engine for the computational identification of putative transcription factor binding sites in multiple genomes | 2005 | 143 | 78 | 63% |
| A survey of motif discovery methods in an integrated framework | 2006 | 55 | 83 | 89% |
| Generic eukaryotic core promoter prediction using structural features of DNA | 2008 | 90 | 97 | 39% |
| Computational identification of transcriptional regulatory elements in DNA sequence | 2006 | 61 | 154 | 71% |
| COMPUTATIONAL STRATEGIES FOR THE GENOME-WIDE IDENTIFICATION OF CIS-REGULATORY ELEMENTS AND TRANSCRIPTIONAL TARGETS | 2012 | 8 | 124 | 61% |
| A survey of motif finding Web tools for detecting binding site motifs in ChIP-Seq data | 2014 | 2 | 70 | 59% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | UKG | 12 | 75% | 0.4% | 9 |
| 2 | BIOINFORMAT GENOMES EAUX BIGRE | 10 | 52% | 0.6% | 14 |
| 3 | BIOINFORMAT INTELLIGENT COMP | 6 | 71% | 0.2% | 5 |
| 4 | THEORET GENET | 6 | 47% | 0.4% | 9 |
| 5 | HARVARD MIT HLTH SCI TECHNOL HST | 4 | 42% | 0.4% | 8 |
| 6 | ARCO GRP | 4 | 50% | 0.3% | 6 |
| 7 | GRP ALGORISM GENET | 3 | 100% | 0.1% | 3 |
| 8 | KNOWLEDGE EXTRACT | 3 | 40% | 0.3% | 6 |
| 9 | GENOM FUNCT ANAL SECT | 3 | 60% | 0.1% | 3 |
| 10 | LBNC | 2 | 67% | 0.1% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000198284 | PROGRAM COMPUTAT GENOM//CIFN//PROGRAMA GENOM COMPUTAC |
| 2 | 0.0000171496 | CHIP SEQ//PEAK CALLING//DNASE SEQ |
| 3 | 0.0000148554 | IROQUOIS//CONSERVED NONCODING ELEMENTS//ENERGY ENVIRONM BIOL COMP |
| 4 | 0.0000117816 | INTERACTION PROPENSITY//RNA BINDING RESIDUES//COMPUTAT INTELLIGENCE LEARNING DISCOVERY |
| 5 | 0.0000095897 | NETWORK INFERENCE//GENE REGULATORY NETWORKS//GENE NETWORK INFERENCE |
| 6 | 0.0000091487 | GENE PREDICTION//GENE FINDING//GENE RECOGNITION |
| 7 | 0.0000080249 | CIS MOTIF//ARABIDOPSIS T87 CELL LINE//GENE ATLAS |
| 8 | 0.0000069672 | MULTIPLE SEQUENCE ALIGNMENT//T COFFEE//STATISTICAL ALIGNMENT |
| 9 | 0.0000063057 | NUCLEOSOME POSITIONING//NUCLEOSOME//NUCLEOSOME MAPPING |
| 10 | 0.0000056257 | EMBL OUTSTN//PROT INFORMAT OURCE//HLTH SCI TECHNOL ICIR |