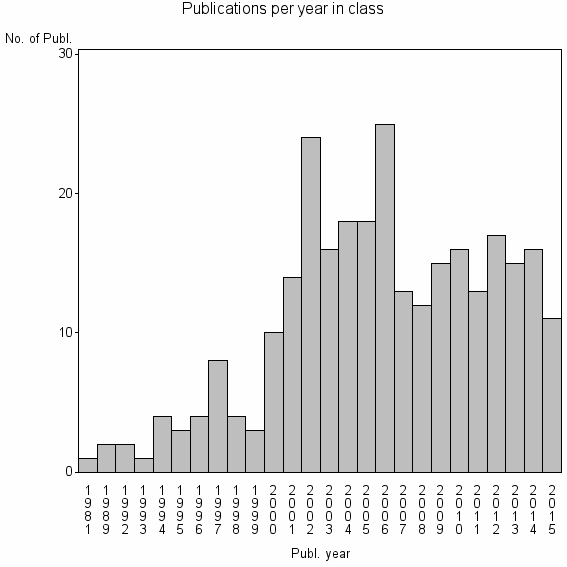
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 23636 | 285 | 39.5 | 84% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | LY573636 | Author keyword | 15 | 88% | 2% | 7 |
| 2 | TASISULAM | Author keyword | 8 | 75% | 2% | 6 |
| 3 | NCI 60 | Author keyword | 5 | 40% | 4% | 10 |
| 4 | CLUSTERED IMAGE MAP | Author keyword | 3 | 100% | 1% | 3 |
| 5 | INFORMAT TECHNOL BRANCH | Address | 2 | 19% | 4% | 11 |
| 6 | COURS ARGONNE 229 | Address | 2 | 67% | 1% | 2 |
| 7 | GLOBAL DISCOVERY DEV STAT | Address | 2 | 67% | 1% | 2 |
| 8 | INFORMAT TECHNOL BRANCHDEV THER EUT PROGRAM | Address | 2 | 67% | 1% | 2 |
| 9 | GENOM BIOINFORMAT GRP | Address | 2 | 20% | 3% | 8 |
| 10 | ADV BIOINFORMAT SYST MED | Address | 1 | 50% | 1% | 2 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | LY573636 | 15 | 88% | 2% | 7 | Search LY573636 | Search LY573636 |
| 2 | TASISULAM | 8 | 75% | 2% | 6 | Search TASISULAM | Search TASISULAM |
| 3 | NCI 60 | 5 | 40% | 4% | 10 | Search NCI+60 | Search NCI+60 |
| 4 | CLUSTERED IMAGE MAP | 3 | 100% | 1% | 3 | Search CLUSTERED+IMAGE+MAP | Search CLUSTERED+IMAGE+MAP |
| 5 | ANTICANCER DRUG METABOLISM | 1 | 50% | 1% | 2 | Search ANTICANCER+DRUG+METABOLISM | Search ANTICANCER+DRUG+METABOLISM |
| 6 | CANCER CELL LINE MODELING | 1 | 100% | 1% | 2 | Search CANCER+CELL+LINE+MODELING | Search CANCER+CELL+LINE+MODELING |
| 7 | KARYOTYPIC COMPLEXITY | 1 | 100% | 1% | 2 | Search KARYOTYPIC+COMPLEXITY | Search KARYOTYPIC+COMPLEXITY |
| 8 | NCI 60 CELL LINES | 1 | 50% | 1% | 2 | Search NCI+60+CELL+LINES | Search NCI+60+CELL+LINES |
| 9 | GENE TRANSCRIPTS | 1 | 21% | 2% | 5 | Search GENE+TRANSCRIPTS | Search GENE+TRANSCRIPTS |
| 10 | CANCER CHEMORESISTANCE | 1 | 27% | 1% | 3 | Search CANCER+CHEMORESISTANCE | Search CANCER+CHEMORESISTANCE |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ELLIPTICINE ANALOGS | 22 | 81% | 5% | 13 |
| 2 | MOLECULAR PHARMACOLOGY | 5 | 12% | 15% | 43 |
| 3 | ANTICANCER DRUG SCREEN | 5 | 15% | 11% | 30 |
| 4 | DRUG DISCOVERY DATABASES | 5 | 55% | 2% | 6 |
| 5 | TUMOR SCREENING DATABASE | 3 | 100% | 1% | 3 |
| 6 | SODIUM LY573636 SODIUM | 2 | 67% | 1% | 2 |
| 7 | INSTITUTES ANTICANCER SCREEN | 2 | 50% | 1% | 3 |
| 8 | DIFFERENTIAL CYTOTOXICITY DATA | 2 | 22% | 3% | 8 |
| 9 | LY573636 SODIUM | 1 | 100% | 1% | 2 |
| 10 | MULTICASE EXPERT SYSTEM | 1 | 33% | 1% | 2 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Using drug response data to identify molecular effectors, and molecular "omic" data to identify candidate drugs in cancer | 2015 | 1 | 61 | 34% |
| Transcriptomic analysis of the NCI-60 cancer cell lines | 2003 | 29 | 43 | 42% |
| Integrating the NCI-60 Data with "Omics" for Drug Discovery | 2012 | 1 | 31 | 45% |
| Connecting chemosensitivity, gene expression and disease | 2008 | 4 | 41 | 32% |
| Analysis of TP53 Mutation Status in Human Cancer Cell Lines: A Reassessment | 2014 | 5 | 38 | 8% |
| Drug sensitivity and resistance genes in cancer chemotherapy: a chemogenomics approach | 2003 | 25 | 38 | 26% |
| Mining the National Cancer Institute's tumor-screening database: Identification of compounds with similar cellular activities | 2002 | 81 | 87 | 13% |
| Molecular targets in cancer drug discovery: Cell-based profiling | 2000 | 14 | 26 | 42% |
| Gene expression microarray technologies in the development of new therapeutic agents | 2004 | 53 | 133 | 11% |
| Neural network techniques for informatics of cancer drug discovery | 2000 | 3 | 24 | 42% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | INFORMAT TECHNOL BRANCH | 2 | 19% | 3.9% | 11 |
| 2 | COURS ARGONNE 229 | 2 | 67% | 0.7% | 2 |
| 3 | GLOBAL DISCOVERY DEV STAT | 2 | 67% | 0.7% | 2 |
| 4 | INFORMAT TECHNOL BRANCHDEV THER EUT PROGRAM | 2 | 67% | 0.7% | 2 |
| 5 | GENOM BIOINFORMAT GRP | 2 | 20% | 2.8% | 8 |
| 6 | ADV BIOINFORMAT SYST MED | 1 | 50% | 0.7% | 2 |
| 7 | FREDERICK CANC DEV SAIC | 1 | 100% | 0.7% | 2 |
| 8 | CANC METAB DRUG DISCOVERY | 1 | 40% | 0.7% | 2 |
| 9 | PROGRAM PHARMACOGENOM | 1 | 14% | 1.8% | 5 |
| 10 | CHEMIOMETRIA CHEMIOINFORMAT | 1 | 50% | 0.4% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000200063 | CHEM TECHNOL DRUGS//2 MERCAPTOBENZENESULFONAMIDE//2 MERCAPTOBENZENESULFONAMIDES |
| 2 | 0.0000104149 | FP3//HOLLOW FIBRE ASSAY//AFFILIATED ZHUJI HOSP |
| 3 | 0.0000097122 | POLYPHARMACOLOGY//NETWORK PHARMACOLOGY//DRUG REPOSITIONING |
| 4 | 0.0000085390 | BIOIMAGING PROBE DEV//SING ORE BIOIMAGING CONSORTIUM//MEDCHEM PROGRAM LIFE SCI |
| 5 | 0.0000082069 | SINOP ARTS SCI//TG MS SYSTEM//ELE OCHIM MAT MOL COMPLEXES |
| 6 | 0.0000077048 | MALCOLM H FILSON S//INTEGRAL APPROXIMATIONS ACCORDING TO MULLIKEN AND RUDENBERG//NEGLECT OF DIATOMIC DIFFERENTIAL OVERLAP NDDO |
| 7 | 0.0000074790 | KIAA1199//MIR 188 5P//TRANSCRIPTOMIC PROFILES |
| 8 | 0.0000069227 | FIBROBLAST LINE//FIBROBLAST CELL LINE//CELLBANK AUSTRALIA |
| 9 | 0.0000068880 | IN VITRO CARCINOGENESIS CELLULAR CHEMOTHER//VITRO CARCINOGENESIS CELLULAR CHEMOTHER Y//TARGET BASED DRUG |
| 10 | 0.0000068628 | PHILIP MORRIS INT RD//COMPUTAT PL GENOM PROGRAM//SECT COMPUTAT BIOMED |