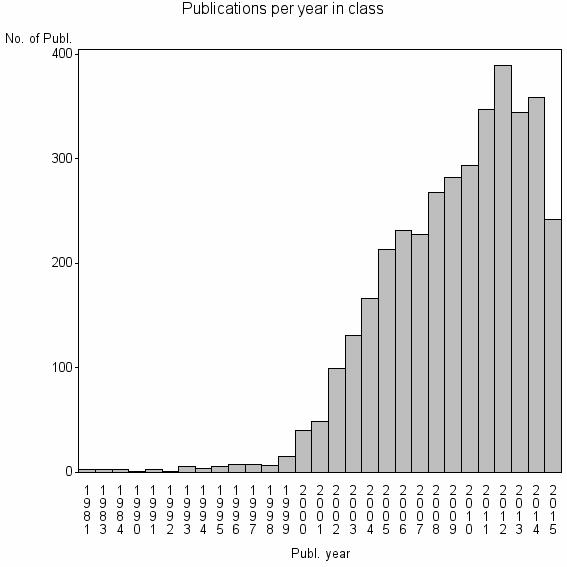
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 246 | 3739 | 40.3 | 87% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 356 | 16803 | PROTEOMICS//JOURNAL OF PROTEOME RESEARCH//PROTEIN IDENTIFICATION |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | PEPTIDE IDENTIFICATION | Author keyword | 97 | 63% | 3% | 97 |
| 2 | JOURNAL OF PROTEOME RESEARCH | Journal | 87 | 12% | 18% | 666 |
| 3 | QUANTITATIVE PROTEOMICS | Author keyword | 74 | 31% | 5% | 204 |
| 4 | PROTEIN INFERENCE | Author keyword | 57 | 95% | 1% | 19 |
| 5 | PROTEOMICS STANDARDS INITIATIVE | Author keyword | 57 | 95% | 1% | 19 |
| 6 | PROTEIN IDENTIFICATION | Author keyword | 51 | 29% | 4% | 152 |
| 7 | MOLECULAR & CELLULAR PROTEOMICS | Journal | 41 | 12% | 9% | 326 |
| 8 | MOL SYST BIOL | Address | 31 | 18% | 4% | 154 |
| 9 | PROTEOGENOMICS | Author keyword | 26 | 50% | 1% | 38 |
| 10 | SHOTGUN PROTEOMICS | Author keyword | 26 | 28% | 2% | 81 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | PEPTIDE IDENTIFICATION | 97 | 63% | 3% | 97 | Search PEPTIDE+IDENTIFICATION | Search PEPTIDE+IDENTIFICATION |
| 2 | QUANTITATIVE PROTEOMICS | 74 | 31% | 5% | 204 | Search QUANTITATIVE+PROTEOMICS | Search QUANTITATIVE+PROTEOMICS |
| 3 | PROTEIN INFERENCE | 57 | 95% | 1% | 19 | Search PROTEIN+INFERENCE | Search PROTEIN+INFERENCE |
| 4 | PROTEOMICS STANDARDS INITIATIVE | 57 | 95% | 1% | 19 | Search PROTEOMICS+STANDARDS+INITIATIVE | Search PROTEOMICS+STANDARDS+INITIATIVE |
| 5 | PROTEIN IDENTIFICATION | 51 | 29% | 4% | 152 | Search PROTEIN+IDENTIFICATION | Search PROTEIN+IDENTIFICATION |
| 6 | PROTEOGENOMICS | 26 | 50% | 1% | 38 | Search PROTEOGENOMICS | Search PROTEOGENOMICS |
| 7 | SHOTGUN PROTEOMICS | 26 | 28% | 2% | 81 | Search SHOTGUN+PROTEOMICS | Search SHOTGUN+PROTEOMICS |
| 8 | HUMAN PROTEOME PROJECT | 25 | 63% | 1% | 25 | Search HUMAN+PROTEOME+PROJECT | Search HUMAN+PROTEOME+PROJECT |
| 9 | CHROMOSOME CENTRIC HUMAN PROTEOME PROJECT | 24 | 82% | 0% | 14 | Search CHROMOSOME+CENTRIC+HUMAN+PROTEOME+PROJECT | Search CHROMOSOME+CENTRIC+HUMAN+PROTEOME+PROJECT |
| 10 | DECOY DATABASE | 23 | 86% | 0% | 12 | Search DECOY+DATABASE | Search DECOY+DATABASE |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PROTEIN IDENTIFICATION | 291 | 39% | 16% | 590 |
| 2 | SHOTGUN PROTEOMICS | 200 | 35% | 12% | 461 |
| 3 | PEPTIDE IDENTIFICATION | 193 | 55% | 6% | 242 |
| 4 | QUANTITATIVE PROTEOMICS | 120 | 26% | 11% | 393 |
| 5 | SEQUENCE DATABASES | 105 | 33% | 7% | 260 |
| 6 | DATABASE SEARCH | 96 | 45% | 4% | 163 |
| 7 | COMPLEX PROTEIN MIXTURES | 89 | 55% | 3% | 110 |
| 8 | SPECTROMETRY DATA | 86 | 60% | 3% | 95 |
| 9 | CODED AFFINITY TAGS | 86 | 34% | 6% | 209 |
| 10 | TANDEM MASS SPECTRA | 83 | 43% | 4% | 149 |
Journals |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | JOURNAL OF PROTEOME RESEARCH | 87 | 12% | 18% | 666 |
| 2 | MOLECULAR & CELLULAR PROTEOMICS | 41 | 12% | 9% | 326 |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Insights into the regulation of protein abundance from proteomic and transcriptomic analyses | 2012 | 410 | 49 | 37% |
| Selected reaction monitoring-based proteomics: workflows, potential, pitfalls and future directions | 2012 | 273 | 101 | 52% |
| Quantitative mass spectrometry in proteomics: a critical review | 2007 | 652 | 113 | 75% |
| Quantitative mass spectrometry in proteomics: critical review update from 2007 to the present | 2012 | 135 | 212 | 60% |
| Mass spectrometry-based proteomics turns quantitative | 2005 | 855 | 113 | 63% |
| A guided tour of the Trans-Proteomic Pipeline | 2010 | 162 | 42 | 95% |
| Review - Mass spectrometry and protein analysis | 2006 | 881 | 30 | 43% |
| Less label, more free: Approaches in label-free quantitative mass spectrometry | 2011 | 190 | 146 | 64% |
| The Coming Age of Complete, Accurate, and Ubiquitous Proteomes | 2013 | 66 | 34 | 74% |
| Selected reaction monitoring for quantitative proteomics: a tutorial | 2008 | 425 | 43 | 56% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MOL SYST BIOL | 31 | 18% | 4.1% | 154 |
| 2 | PROTE SIGNAL TRANSDUCT | 25 | 29% | 2.0% | 73 |
| 3 | LUXEMBOURG CLIN PROTE LCP | 18 | 83% | 0.3% | 10 |
| 4 | PROTEOME INFORMAT GRP | 14 | 46% | 0.6% | 23 |
| 5 | JIM AYERS PRECANC DETECT DIAG | 13 | 51% | 0.5% | 18 |
| 6 | EMBL OUTSTN | 11 | 21% | 1.3% | 48 |
| 7 | MED PROT | 10 | 13% | 1.9% | 70 |
| 8 | BEIJING PROTEOME | 9 | 17% | 1.3% | 48 |
| 9 | CELL BIOL SR11 | 8 | 75% | 0.2% | 6 |
| 10 | PROTEOM SIGNAL TRANSDUCT | 7 | 32% | 0.5% | 19 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000218404 | SISCAPA//PSAQ//BIOMARKER VERIFICATION |
| 2 | 0.0000152136 | SAMPLE PREFRACTIONATION//PREFRACTIONATION//ALKALINE PROTEINS |
| 3 | 0.0000139074 | PHOSPHOPROTEOMICS//PHOSPHOPEPTIDE ENRICHMENT//PHOSPHOPEPTIDES |
| 4 | 0.0000105597 | PEPTIDE FRAGMENTATION//COMPUTAT PROTE GRP//PROLINE EFFECT |
| 5 | 0.0000103530 | TANDEM AFFINITY PURIFICATION//TAP TANDEM AFFINITY PURIFICATION//TANDEM AFFINITY PURIFICATION TAP |
| 6 | 0.0000103158 | GPLLI PROGRAM//IMMOBILIZED TRYPSIN//PROTEIN DIGESTION |
| 7 | 0.0000101515 | ION ION REACTIONS//ELECTRON CAPTURE DISSOCIATION//ELECTRON TRANSFER DISSOCIATION |
| 8 | 0.0000096255 | PROTEIN SIP//SOIL METAPROTEOMICS//CHEM DIRECTORATE |
| 9 | 0.0000094830 | TWO DIMENSIONAL POLYACRYLAMIDE GEL ELECTROPHORESIS//MYOCARDIAL PROTEINS//PROTEOME GENE PROD M PING |
| 10 | 0.0000087966 | 2 DE MAP//DATA INDEPENDENT SCANNING//LEUKAEMIA FUND PROTEOM IL |