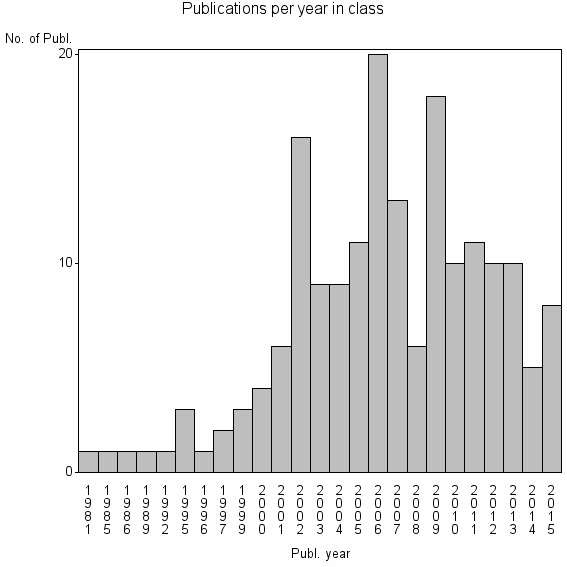
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 27819 | 180 | 42.2 | 90% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 238 | 19563 | BIOINFORMATICS//BMC BIOINFORMATICS//GENOME RESEARCH |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | PROCESSED PSEUDOGENES | Author keyword | 3 | 38% | 3% | 6 |
| 2 | PHYSICAL AND PSYCHOLOGICAL STRESSES | Author keyword | 2 | 67% | 1% | 2 |
| 3 | DUPLICATED PSEUDOGENES | Author keyword | 1 | 100% | 1% | 2 |
| 4 | NONFUNCTIONALIZATION | Author keyword | 1 | 33% | 1% | 2 |
| 5 | AEONS | Author keyword | 1 | 50% | 1% | 1 |
| 6 | CALMODULIN II | Author keyword | 1 | 50% | 1% | 1 |
| 7 | CAS MAX PLANK JR GRP | Address | 1 | 50% | 1% | 1 |
| 8 | CESCR | Author keyword | 1 | 50% | 1% | 1 |
| 9 | COMPARAT GENOMICS BIOINFORMAT | Address | 1 | 50% | 1% | 1 |
| 10 | EVOLUTIONARY PROC GRP | Address | 1 | 50% | 1% | 1 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | PROCESSED PSEUDOGENES | 3 | 38% | 3% | 6 | Search PROCESSED+PSEUDOGENES | Search PROCESSED+PSEUDOGENES |
| 2 | PHYSICAL AND PSYCHOLOGICAL STRESSES | 2 | 67% | 1% | 2 | Search PHYSICAL+AND+PSYCHOLOGICAL+STRESSES | Search PHYSICAL+AND+PSYCHOLOGICAL+STRESSES |
| 3 | DUPLICATED PSEUDOGENES | 1 | 100% | 1% | 2 | Search DUPLICATED+PSEUDOGENES | Search DUPLICATED+PSEUDOGENES |
| 4 | NONFUNCTIONALIZATION | 1 | 33% | 1% | 2 | Search NONFUNCTIONALIZATION | Search NONFUNCTIONALIZATION |
| 5 | AEONS | 1 | 50% | 1% | 1 | Search AEONS | Search AEONS |
| 6 | CALMODULIN II | 1 | 50% | 1% | 1 | Search CALMODULIN+II | Search CALMODULIN+II |
| 7 | CESCR | 1 | 50% | 1% | 1 | Search CESCR | Search CESCR |
| 8 | GENE DEATH | 1 | 50% | 1% | 1 | Search GENE+DEATH | Search GENE+DEATH |
| 9 | MGEA6 | 1 | 50% | 1% | 1 | Search MGEA6 | Search MGEA6 |
| 10 | POTOGENE | 1 | 50% | 1% | 1 | Search POTOGENE | Search POTOGENE |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PROCESSED PSEUDOGENES | 20 | 22% | 44% | 80 |
| 2 | HOMOLOGOUS CODING GENE | 10 | 54% | 7% | 13 |
| 3 | PROTEOME EVOLUTION | 8 | 75% | 3% | 6 |
| 4 | RIBOSOMAL PROTEIN PSEUDOGENES | 8 | 75% | 3% | 6 |
| 5 | TRANSCRIBED PSEUDOGENE | 8 | 100% | 3% | 5 |
| 6 | EXPRESSED PSEUDOGENE | 4 | 67% | 2% | 4 |
| 7 | ENCODE REGIONS | 4 | 41% | 4% | 7 |
| 8 | RETROPSEUDOGENES | 2 | 40% | 2% | 4 |
| 9 | STRUCTURAL CENSUS | 1 | 38% | 2% | 3 |
| 10 | CONNEXIN43 PSEUDOGENE | 1 | 100% | 1% | 2 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Pseudogenes: Pseudo or Real Functional Elements? | 2013 | 10 | 67 | 63% |
| Pseudogenes: Pseudo-functional or key regulators in health and disease? | 2011 | 63 | 75 | 45% |
| The ambiguous boundary between genes and pseudogenes: the dead rise up, or do they? | 2007 | 55 | 45 | 56% |
| Pseudogenes as regulators of biological function | 2013 | 2 | 40 | 63% |
| Functional evidence of post-transcriptional regulation by pseudogenes | 2011 | 20 | 55 | 44% |
| Large-scale analysis of pseudogenes in the human genome | 2004 | 90 | 50 | 38% |
| Pseudogenes | 2012 | 2 | 19 | 63% |
| L1 elements, processed pseudogenes and retrogenes in mammalian genomes | 2006 | 22 | 61 | 28% |
| Pseudogene in cancer: real functions and promising signature | 2015 | 0 | 53 | 40% |
| Pseudogenes: Newly Discovered Players in Human Cancer | 2012 | 22 | 164 | 14% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CAS MAX PLANK JR GRP | 1 | 50% | 0.6% | 1 |
| 2 | COMPARAT GENOMICS BIOINFORMAT | 1 | 50% | 0.6% | 1 |
| 3 | EVOLUTIONARY PROC GRP | 1 | 50% | 0.6% | 1 |
| 4 | ONCOGENOM UNIT | 1 | 50% | 0.6% | 1 |
| 5 | PROGRAM COMP BIOL BIOINFORMAT | 1 | 50% | 0.6% | 1 |
| 6 | RNA BIOL CLIN PLICAT | 1 | 29% | 1.1% | 2 |
| 7 | BIOL SYSTEXPT ANIM | 0 | 33% | 0.6% | 1 |
| 8 | IST TOSCANO TUMORI CRL ITT | 0 | 33% | 0.6% | 1 |
| 9 | MOL GENET BIOINFORMAT | 0 | 18% | 1.1% | 2 |
| 10 | DONNELLY CCBR | 0 | 14% | 1.1% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000215159 | SEX BIASED GENE EXPRESSION//SEXUAL ANTAGONISM//INTRALOCUS SEXUAL CONFLICT |
| 2 | 0.0000183623 | WHOLE GENOME DUPLICATION//GENET MOL CELLULAIRE ECT LEVU INTERE//GENOME DUPLICATION |
| 3 | 0.0000158490 | NUMT//NUMTS//NUMTDNA |
| 4 | 0.0000157459 | WRAP53//OVERLAPPING GENES//CUSTOM BIOTECHNOL SERV GRP |
| 5 | 0.0000145722 | BIOL COMPUTAT VISUALIZAT//SINE//NON LTR RETROTRANSPOSON |
| 6 | 0.0000115272 | GENE PREDICTION//GENE FINDING//GENE RECOGNITION |
| 7 | 0.0000100073 | LONG NONCODING RNAS//LNCRNA//LONG NON CODING RNA |
| 8 | 0.0000094262 | INTRON GAIN//INTRON LOSS//INTRON EVOLUTION |
| 9 | 0.0000077339 | ISOCHORES//EVOLUZ MOL//ISOCHORE |
| 10 | 0.0000068353 | INSECT NERVOUS SYSTEM//MOL NEUROBIOL CELLULAR PHYSIOL//PLEUROBRANCHAEA |