Class information for: |
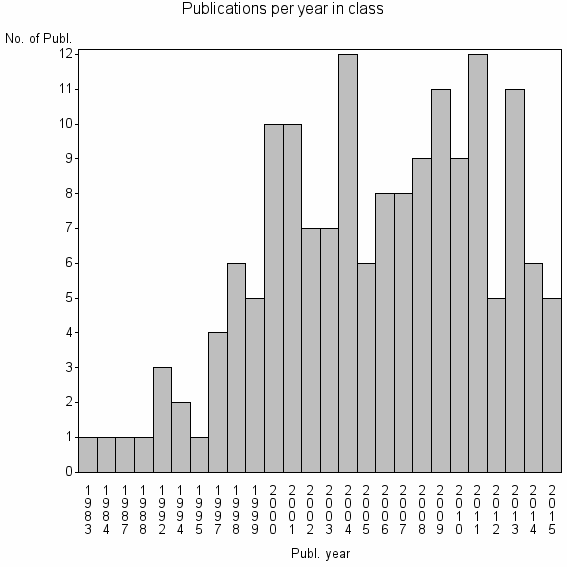
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 28864 | 161 | 58.8 | 79% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | GENOME THEORY | Author keyword | 37 | 100% | 9% | 14 |
| 2 | GENOME CHAOS | Author keyword | 21 | 90% | 6% | 9 |
| 3 | REPLICATION PATTERN | Author keyword | 7 | 67% | 4% | 6 |
| 4 | MED KLIN MANNHEIM 3 | Address | 5 | 47% | 5% | 8 |
| 5 | NON CLONAL CHROMOSOME ABERRATION NCCA | Author keyword | 3 | 100% | 2% | 3 |
| 6 | NON CLONAL CHROMOSOME ABERRATIONS NCCAS | Author keyword | 3 | 100% | 2% | 3 |
| 7 | SYSTEM INHERITANCE | Author keyword | 3 | 100% | 2% | 3 |
| 8 | CANCER EVOLUTIONARY MECHANISM | Author keyword | 1 | 100% | 1% | 2 |
| 9 | CLONAL CHROMOSOME ABERRATION CCA | Author keyword | 1 | 100% | 1% | 2 |
| 10 | EVOLUTIONARY MECHANISM OF CANCER | Author keyword | 1 | 100% | 1% | 2 |
Web of Science journal categories |
Author Key Words |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | 21Q22 LOCI | 21 | 85% | 7% | 11 |
| 2 | CLONAL CHROMOSOME ABERRATIONS | 16 | 51% | 14% | 22 |
| 3 | EPIGENETIC HETEROGENEITY | 8 | 70% | 4% | 7 |
| 4 | EVOLUTIONARY MECHANISM | 7 | 53% | 6% | 9 |
| 5 | ALLELE SPECIFIC REPLICATION | 7 | 38% | 9% | 14 |
| 6 | RANDOM ANEUPLOIDY | 6 | 100% | 2% | 4 |
| 7 | ASYNCHRONOUS REPLICATION | 5 | 21% | 12% | 20 |
| 8 | CENTRIC CONCEPT | 3 | 100% | 2% | 3 |
| 9 | ALLELIC REPLICATION | 3 | 50% | 2% | 4 |
| 10 | SEGMENTAL ANEUPLOIDY | 2 | 36% | 3% | 5 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Chromosomal instability (CIN): what it is and why it is crucial to cancer evolution | 2013 | 12 | 102 | 31% |
| Evolutionary Mechanisms and Diversity in Cancer | 2011 | 20 | 87 | 25% |
| Genetic and Epigenetic Heterogeneity in Cancer: A Genome-Centric Perspective | 2009 | 49 | 80 | 15% |
| Why imatinib remains an exception of cancer research | 2013 | 9 | 65 | 20% |
| Cancer-causing karyotypes: chromosomal equilibria between destabilizing aneuploidy and stabilizing selection for oncogenic function | 2009 | 24 | 83 | 19% |
| Comparison of mitotic cell death by chromosome fragmentation to premature chromosome condensation | 2010 | 19 | 61 | 20% |
| Ovarian cancer evolution through stochastic genome alterations: defining the genomic role in ovarian cancer | 2014 | 3 | 113 | 16% |
| Aneuploidy, the primary cause of the multilateral genomic instability of neoplastic and preneoplastic cells | 2004 | 83 | 144 | 8% |
| Evolutionary Mechanism Unifies the Hallmarks of Cancer | 2015 | 0 | 69 | 32% |
| Chromosomal instability and transcriptome dynamics in cancer | 2013 | 6 | 67 | 15% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MED KLIN MANNHEIM 3 | 5 | 47% | 5.0% | 8 |
| 2 | FOSTER | 1 | 100% | 1.2% | 2 |
| 3 | MOL MED GENETKARMANOS CANC | 1 | 100% | 1.2% | 2 |
| 4 | EHS MOL CELLULAR TOXICOL HUMAN PLICAT | 1 | 40% | 1.2% | 2 |
| 5 | BIOSYNTH NUCLE ACIDS | 1 | 29% | 1.2% | 2 |
| 6 | CNRS UMR8126 | 0 | 33% | 0.6% | 1 |
| 7 | DANA FARBER CANC DERMATOL | 0 | 33% | 0.6% | 1 |
| 8 | E 361 | 0 | 25% | 0.6% | 1 |
| 9 | MED HEMATOL A | 0 | 25% | 0.6% | 1 |
| 10 | CELLULAR MOL ANAL | 0 | 13% | 0.6% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000173157 | EVOLUT CANC//PETOS PARADOX//TRANSLAT CANC THER EUT |
| 2 | 0.0000147702 | PROGRESSIVE STATE SELECTION//SPONTANEOUS TRANSFORMATION//GROWTH CONSTRAINT |
| 3 | 0.0000136653 | INTER SIMPLE SEQUENCE REPEAT PCR//BIOPHYS CYTOMETRY//HOSP UNIV SALAMANCA IBSAL |
| 4 | 0.0000134610 | SPECTRAL SIMILARITY MAPPING//FAST FISH//CHROMOSOME SPREADING |
| 5 | 0.0000132820 | REVERSINE//MITOTIC CATASTROPHE//WP 631 |
| 6 | 0.0000114840 | KINETOCHORE//SPINDLE ASSEMBLY CHECKPOINT//MAD2 |
| 7 | 0.0000112658 | ROBERT ROSEN//M R SYSTEMS//RELATIONAL BIOLOGY |
| 8 | 0.0000102155 | BINUCLEATED LYMPHOCYTES//POLYCLONAL LYMPHOCYTOSIS//PPBL |
| 9 | 0.0000092548 | SPAG5//OLIGOMEGANEPHRONIA//ASTRIN |
| 10 | 0.0000088816 | JUMPING TRANSLOCATION//INTERSTITIAL TELOMERIC SEQUENCES//TELOMERE ASSOCIATION |