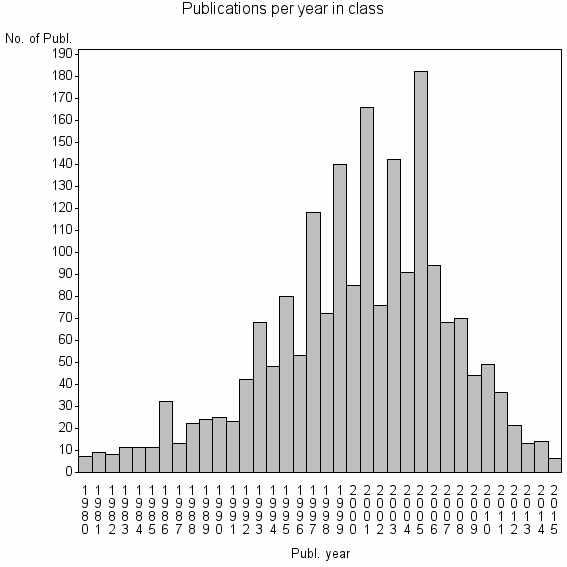
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 3139 | 1974 | 25.3 | 68% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 480 | 14980 | GENETIC EPIDEMIOLOGY//GENOMIC SELECTION//BMC GENETICS |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | GENETIC EPIDEMIOLOGY | Journal | 160 | 25% | 29% | 563 |
| 2 | LINKAGE ANALYSIS | Author keyword | 34 | 13% | 12% | 242 |
| 3 | MODEL FREE LINKAGE ANALYSIS | Author keyword | 31 | 92% | 1% | 12 |
| 4 | HASEMAN ELSTON | Author keyword | 22 | 81% | 1% | 13 |
| 5 | HASEMAN ELSTON REGRESSION | Author keyword | 21 | 90% | 0% | 9 |
| 6 | SIB PAIRS | Author keyword | 19 | 49% | 1% | 28 |
| 7 | BMC GENETICS | Journal | 17 | 10% | 8% | 153 |
| 8 | IDENTITY BY DESCENT | Author keyword | 16 | 32% | 2% | 41 |
| 9 | MOD SCORES | Author keyword | 15 | 88% | 0% | 7 |
| 10 | POSTERIOR PROBABILITY OF LINKAGE | Author keyword | 15 | 88% | 0% | 7 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | LINKAGE ANALYSIS | 34 | 13% | 12% | 242 | Search LINKAGE+ANALYSIS | Search LINKAGE+ANALYSIS |
| 2 | MODEL FREE LINKAGE ANALYSIS | 31 | 92% | 1% | 12 | Search MODEL+FREE+LINKAGE+ANALYSIS | Search MODEL+FREE+LINKAGE+ANALYSIS |
| 3 | HASEMAN ELSTON | 22 | 81% | 1% | 13 | Search HASEMAN+ELSTON | Search HASEMAN+ELSTON |
| 4 | HASEMAN ELSTON REGRESSION | 21 | 90% | 0% | 9 | Search HASEMAN+ELSTON+REGRESSION | Search HASEMAN+ELSTON+REGRESSION |
| 5 | SIB PAIRS | 19 | 49% | 1% | 28 | Search SIB+PAIRS | Search SIB+PAIRS |
| 6 | IDENTITY BY DESCENT | 16 | 32% | 2% | 41 | Search IDENTITY+BY+DESCENT | Search IDENTITY+BY+DESCENT |
| 7 | MOD SCORES | 15 | 88% | 0% | 7 | Search MOD+SCORES | Search MOD+SCORES |
| 8 | POSTERIOR PROBABILITY OF LINKAGE | 15 | 88% | 0% | 7 | Search POSTERIOR+PROBABILITY+OF+LINKAGE | Search POSTERIOR+PROBABILITY+OF+LINKAGE |
| 9 | LOD SCORES | 14 | 53% | 1% | 18 | Search LOD+SCORES | Search LOD+SCORES |
| 10 | INHERITANCE VECTOR | 13 | 80% | 0% | 8 | Search INHERITANCE+VECTOR | Search INHERITANCE+VECTOR |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | COMPLEX PEDIGREES | 45 | 56% | 3% | 55 |
| 2 | HASEMAN ELSTON METHOD | 36 | 79% | 1% | 23 |
| 3 | AFFECTED RELATIVE PAIRS | 35 | 38% | 4% | 74 |
| 4 | GENETICALLY COMPLEX TRAITS | 34 | 37% | 4% | 74 |
| 5 | ASSESSING GENETIC LINKAGE | 33 | 61% | 2% | 35 |
| 6 | LOD SCORE ANALYSIS | 33 | 75% | 1% | 24 |
| 7 | HASEMAN ELSTON | 27 | 92% | 1% | 11 |
| 8 | INFERRING MODE | 26 | 87% | 1% | 13 |
| 9 | GENERAL PEDIGREES | 26 | 39% | 3% | 53 |
| 10 | LOCATION SCORES | 25 | 77% | 1% | 17 |
Journals |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | GENETIC EPIDEMIOLOGY | 160 | 25% | 29% | 563 |
| 2 | BMC GENETICS | 17 | 10% | 8% | 153 |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Methods for Handling Multiple Testing | 2008 | 80 | 38 | 32% |
| Variance component methods for detecting complex trait loci | 2001 | 145 | 40 | 70% |
| GENETIC DISSECTION OF COMPLEX TRAITS | 1994 | 2295 | 191 | 33% |
| Key concepts in genetic epidemiology | 2005 | 102 | 54 | 24% |
| False positives and false negatives in genome scans | 2001 | 36 | 11 | 91% |
| Contemporary Model-Free Methods for Linkage Analysis | 2008 | 7 | 66 | 76% |
| A Kernel of Truth: Statistical Advances in Polygenic Variance Component Models for Complex Human Pedigrees | 2013 | 6 | 55 | 25% |
| Finding genes underlying human disease | 2009 | 10 | 43 | 40% |
| Genomewide scans of complex human diseases: True linkage is hard to find | 2001 | 318 | 120 | 11% |
| Localization and identification of human quantitative trait loci: King Harvest has surely come | 2004 | 58 | 39 | 44% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PROGRAM PUBL HLTH GENET | 6 | 38% | 0.6% | 12 |
| 2 | RAMMELKAMP EDUC | 3 | 12% | 1.3% | 26 |
| 3 | BATTELLE MATH MED | 3 | 18% | 0.8% | 16 |
| 4 | GENET MOL PSYCHIAT | 1 | 100% | 0.1% | 2 |
| 5 | RECH BIOTECHNOL MARINES | 1 | 100% | 0.1% | 2 |
| 6 | SPECIAL POPULAT HLTH | 1 | 50% | 0.1% | 2 |
| 7 | SPIELBERG PEDIAT BURNS ALLEN | 1 | 100% | 0.1% | 2 |
| 8 | GENOMETR SECT | 1 | 15% | 0.4% | 8 |
| 9 | UNIT 24 | 1 | 21% | 0.3% | 5 |
| 10 | METHODOL HLTH | 1 | 40% | 0.1% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000211677 | RELATIONSHIP ESTIMATION//PEDIGREE RECONSTRUCTION//RELATIONSHIP INFERENCE |
| 2 | 0.0000206128 | GENETIC EPIDEMIOLOGY//EFFICIENCY ROBUSTNESS//FAMILY BASED ASSOCIATION TESTS |
| 3 | 0.0000164255 | CALPAIN 10//MED CELLULAR MOL PHYSIOL BIOSTAT//MOL GENET COMPLEX MONOGEN DISORDERS |
| 4 | 0.0000145042 | SPOUSAL CONCORDANCE//GENETIC AND ENVIRONMENTAL COMPONENTS//SPOUSE CONCORDANCE |
| 5 | 0.0000144697 | ZFAT//CENT ADV MOL MED//ZBTB38 GENE |
| 6 | 0.0000134893 | INTERVAL MAPPING//SYSTEMS MAPPING//SELECTIVE GENOTYPING |
| 7 | 0.0000128494 | G72//DAOA//MANIC DEPRESSIVE ILLNESS |
| 8 | 0.0000091594 | HAPPY MAPPING//MULTILOCUS ORDERING//EXOGAMIC POPULATIONS |
| 9 | 0.0000087852 | BETA3 ADRENERGIC RECEPTOR GENE//TRP64ARG POLYMORPHISM//BETA3 ADRENERGIC RECEPTOR |
| 10 | 0.0000081017 | HAPLOTYPE INFERENCE//PURE PARSIMONY//HAPLOTYPE ASSEMBLY |