Class information for: |
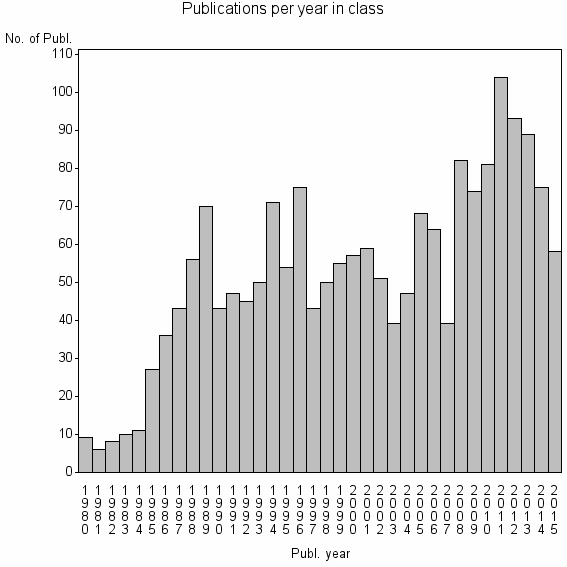
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 3512 | 1889 | 47.5 | 81% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | BICOID | Author keyword | 58 | 68% | 3% | 50 |
| 2 | GAP GENES | Author keyword | 53 | 85% | 1% | 28 |
| 3 | HUNCHBACK | Author keyword | 45 | 63% | 2% | 45 |
| 4 | PAIR RULE GENES | Author keyword | 30 | 79% | 1% | 19 |
| 5 | EVEN SKIPPED | Author keyword | 30 | 49% | 2% | 44 |
| 6 | GAP GENE | Author keyword | 22 | 68% | 1% | 19 |
| 7 | PAIR RULE GENE | Author keyword | 21 | 71% | 1% | 17 |
| 8 | TERMINAL SYSTEM | Author keyword | 21 | 90% | 0% | 9 |
| 9 | FUSHI TARAZU | Author keyword | 20 | 62% | 1% | 21 |
| 10 | DROSOPHILA SEGMENTATION | Author keyword | 17 | 79% | 1% | 11 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | BICOID | 58 | 68% | 3% | 50 | Search BICOID | Search BICOID |
| 2 | GAP GENES | 53 | 85% | 1% | 28 | Search GAP+GENES | Search GAP+GENES |
| 3 | HUNCHBACK | 45 | 63% | 2% | 45 | Search HUNCHBACK | Search HUNCHBACK |
| 4 | PAIR RULE GENES | 30 | 79% | 1% | 19 | Search PAIR+RULE+GENES | Search PAIR+RULE+GENES |
| 5 | EVEN SKIPPED | 30 | 49% | 2% | 44 | Search EVEN+SKIPPED | Search EVEN+SKIPPED |
| 6 | GAP GENE | 22 | 68% | 1% | 19 | Search GAP+GENE | Search GAP+GENE |
| 7 | PAIR RULE GENE | 21 | 71% | 1% | 17 | Search PAIR+RULE+GENE | Search PAIR+RULE+GENE |
| 8 | TERMINAL SYSTEM | 21 | 90% | 0% | 9 | Search TERMINAL+SYSTEM | Search TERMINAL+SYSTEM |
| 9 | FUSHI TARAZU | 20 | 62% | 1% | 21 | Search FUSHI+TARAZU | Search FUSHI+TARAZU |
| 10 | DROSOPHILA SEGMENTATION | 17 | 79% | 1% | 11 | Search DROSOPHILA+SEGMENTATION | Search DROSOPHILA+SEGMENTATION |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | HUNCHBACK | 124 | 59% | 7% | 139 |
| 2 | EARLY DROSOPHILA EMBRYO | 78 | 57% | 5% | 93 |
| 3 | SEGMENTATION GENE | 64 | 43% | 6% | 113 |
| 4 | POSITIONAL INFORMATION | 63 | 33% | 8% | 157 |
| 5 | PAIR RULE GENES | 62 | 80% | 2% | 39 |
| 6 | BICOID PROTEIN | 60 | 49% | 5% | 90 |
| 7 | ANTERIOR PATTERN | 60 | 69% | 3% | 51 |
| 8 | GAP GENES | 54 | 65% | 3% | 51 |
| 9 | BICOID MORPHOGEN GRADIENT | 54 | 75% | 2% | 39 |
| 10 | FUSHI TARAZU | 54 | 33% | 7% | 135 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Coordination of Patterning and Growth by the Morphogen DPP | 2014 | 17 | 71 | 32% |
| The gap gene network | 2011 | 43 | 337 | 83% |
| Developmental roles of 21 Drosophila transcription factors are determined by quantitative differences in binding to an overlapping set of thousands of genomic regions | 2009 | 146 | 93 | 43% |
| Modelling the Bicoid gradient | 2010 | 44 | 47 | 83% |
| The Bicoid Morphogen System | 2010 | 40 | 48 | 88% |
| The interpretation of morphogen gradients | 2006 | 235 | 89 | 39% |
| Morphogen Gradients: From Generation to Interpretation | 2011 | 88 | 199 | 37% |
| How Cells Know Where They Are | 2013 | 19 | 46 | 46% |
| Developmental Pattern Formation: Insights from Physics and Biology | 2012 | 23 | 42 | 60% |
| Arthropod segmentation: Beyond the Drosophila paradigm | 2005 | 112 | 85 | 73% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | EMBL CRG UNIT SYST BIOL | 15 | 63% | 0.8% | 15 |
| 2 | ABT MOL ENTWICKLUNGSBIOL | 9 | 27% | 1.5% | 29 |
| 3 | CHICAGO SYST BIOL | 6 | 80% | 0.2% | 4 |
| 4 | DEV EVOLUT | 6 | 28% | 1.0% | 18 |
| 5 | GENET GENOM DEV | 5 | 26% | 0.8% | 16 |
| 6 | EXCELLENCE WIRELESS INFORMAT TECHNOL | 4 | 67% | 0.2% | 4 |
| 7 | BIOTECHNOL BIO OURCES AGR | 4 | 75% | 0.2% | 3 |
| 8 | UNIT SYST BIOL | 4 | 75% | 0.2% | 3 |
| 9 | ABT GENET TIERE | 4 | 36% | 0.4% | 8 |
| 10 | CEWIT | 3 | 60% | 0.2% | 3 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000200797 | LEG DEVELOPMENT//VESTIGIAL//HOMOTHORAX |
| 2 | 0.0000196970 | TWISTED GASTRULATION//TOLLOID//CHORDIN |
| 3 | 0.0000169618 | DROSOPHILA MATERNAL EFFECT MUTATION//DROSOPHILA MELANOGASTER EMBRYO//IMAGINAL PRIMORDIA |
| 4 | 0.0000131842 | HOX//HOX GENES//PROBOSCIPEDIA |
| 5 | 0.0000105208 | RHOMBOID//SPITZ//RHOMBOID PROTEASE |
| 6 | 0.0000104785 | ACHAETE SCUTE//GENET 240//ENHANCER OF SPLIT |
| 7 | 0.0000100432 | TINMAN//DEV SYST BIOL//STICKS AND STONES |
| 8 | 0.0000096207 | OSKAR//RNA LOCALIZATION//VASA |
| 9 | 0.0000095219 | ONYCHOPHORA//PYCNOGONIDA//PERIPATOPSIDAE |
| 10 | 0.0000083721 | DREF//INSECT BIOMED//DNA REPLICATION RELATED GENE |