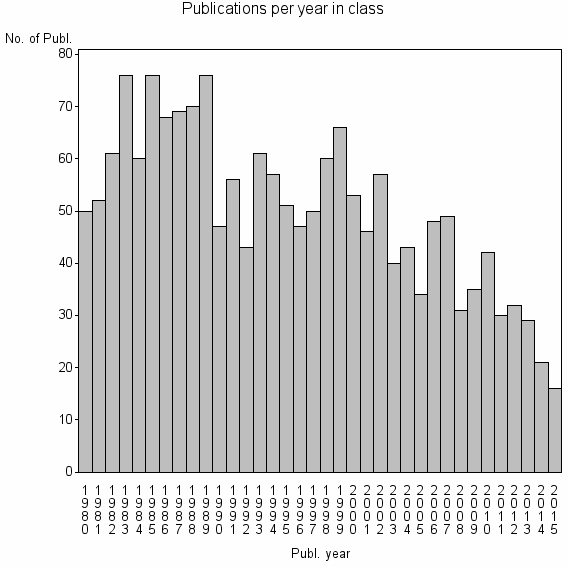
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 3938 | 1802 | 39.4 | 68% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | NIFA PROTEIN | Author keyword | 41 | 100% | 1% | 15 |
| 2 | SIGMA54 | Author keyword | 37 | 67% | 2% | 33 |
| 3 | NIFA | Author keyword | 31 | 59% | 2% | 35 |
| 4 | GLNB | Author keyword | 29 | 75% | 1% | 21 |
| 5 | GLNK | Author keyword | 29 | 75% | 1% | 21 |
| 6 | SIGMA 54 | Author keyword | 23 | 56% | 2% | 28 |
| 7 | NITROGENASE REGULATION | Author keyword | 21 | 90% | 0% | 9 |
| 8 | NIFL | Author keyword | 18 | 83% | 1% | 10 |
| 9 | NTRC | Author keyword | 18 | 59% | 1% | 20 |
| 10 | NIF REGULATION | Author keyword | 17 | 100% | 0% | 8 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | NIFA PROTEIN | 41 | 100% | 1% | 15 | Search NIFA+PROTEIN | Search NIFA+PROTEIN |
| 2 | SIGMA54 | 37 | 67% | 2% | 33 | Search SIGMA54 | Search SIGMA54 |
| 3 | NIFA | 31 | 59% | 2% | 35 | Search NIFA | Search NIFA |
| 4 | GLNB | 29 | 75% | 1% | 21 | Search GLNB | Search GLNB |
| 5 | GLNK | 29 | 75% | 1% | 21 | Search GLNK | Search GLNK |
| 6 | SIGMA 54 | 23 | 56% | 2% | 28 | Search SIGMA+54 | Search SIGMA+54 |
| 7 | NITROGENASE REGULATION | 21 | 90% | 0% | 9 | Search NITROGENASE+REGULATION | Search NITROGENASE+REGULATION |
| 8 | NIFL | 18 | 83% | 1% | 10 | Search NIFL | Search NIFL |
| 9 | NTRC | 18 | 59% | 1% | 20 | Search NTRC | Search NTRC |
| 10 | NIF REGULATION | 17 | 100% | 0% | 8 | Search NIF+REGULATION | Search NIF+REGULATION |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | SIGNAL TRANSDUCTION PROTEINS | 88 | 44% | 8% | 150 |
| 2 | FACTOR SIGMA 54 | 73 | 96% | 1% | 23 |
| 3 | GLNALG OPERON | 64 | 80% | 2% | 40 |
| 4 | REVERSIBLE ADP RIBOSYLATION | 59 | 78% | 2% | 39 |
| 5 | NTRC | 59 | 59% | 4% | 66 |
| 6 | GLNA | 49 | 69% | 2% | 42 |
| 7 | P II PROTEIN | 49 | 60% | 3% | 53 |
| 8 | P II | 45 | 34% | 6% | 108 |
| 9 | UPSTREAM ACTIVATOR SEQUENCES | 42 | 83% | 1% | 24 |
| 10 | SIGMA 54 | 42 | 78% | 2% | 28 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| PII signal transduction proteins: nitrogen regulation and beyond | 2013 | 24 | 283 | 77% |
| P-II signal transduction proteins, pivotal players in microbial nitrogen control | 2001 | 263 | 226 | 70% |
| Post-translational modification of P-II signal transduction proteins | 2015 | 1 | 38 | 87% |
| Nitrogen regulation in bacteria and archaea | 2007 | 145 | 100 | 68% |
| The Role of Bacterial Enhancer Binding Proteins as Specialized Activators of sigma(54)-Dependent Transcription | 2012 | 37 | 233 | 57% |
| PII signal transduction proteins: sensors of alpha-ketoglutarate that regulate nitrogen metabolism | 2005 | 141 | 42 | 83% |
| P-II signal transducers: novel functional and structural insights | 2008 | 85 | 58 | 67% |
| Genetic regulation of biological nitrogen fixation | 2004 | 237 | 74 | 58% |
| NITROGEN CONTROL IN BACTERIA | 1995 | 403 | 245 | 65% |
| PII signal transduction proteins | 2000 | 184 | 45 | 80% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | STUDY NITROGEN FIXAT | 14 | 55% | 0.9% | 17 |
| 2 | NACL CIENCIA TECNOL FIXACAO BIOL NITROGENI | 7 | 64% | 0.4% | 7 |
| 3 | INTEGRAT BIOSCI DEGREE PROGRAM CHEM BIOL | 3 | 100% | 0.2% | 3 |
| 4 | SCI TECHNOL BIOL NITROGEN FIXAT | 3 | 100% | 0.2% | 3 |
| 5 | RECONOCIMIENTO MOL BIOESTRUCTURA | 2 | 44% | 0.2% | 4 |
| 6 | AD TAT MICROORGANISMS | 2 | 67% | 0.1% | 2 |
| 7 | STRUCT BIOL MOL BIOSCI | 2 | 67% | 0.1% | 2 |
| 8 | URA 1300UNITE PHYSIOL CELLULAIRE | 2 | 67% | 0.1% | 2 |
| 9 | GRP SYMBIOSIS | 2 | 43% | 0.2% | 3 |
| 10 | INTERFAK MIKROBIOL INFEKT MED | 1 | 19% | 0.3% | 6 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000230664 | TRANSCRIPTION FACTOR TNRA//GLNR//BECKMAN COULTER |
| 2 | 0.0000217451 | ALGAL RESPIRATION//METHYLAMMONIUM UPTAKE//AMMONIUM TRANSPORT |
| 3 | 0.0000136146 | MOFE PROTEIN//FEMO COFACTOR//NITROGENASE |
| 4 | 0.0000118144 | HEME SENSOR//FIXL//COOA |
| 5 | 0.0000097495 | BEIJERINCKIA DERXII//GRP ENVIRONM MICROBIOL//VITAMIN PRODUCTION |
| 6 | 0.0000096912 | XYLS//TOL PLASMID//XYLR |
| 7 | 0.0000089820 | PPSR//CRTJ//FNRL |
| 8 | 0.0000080859 | NTCA//BIOQUIM VEGETAL FOTOSINTESIS//HETEROCYST |
| 9 | 0.0000066848 | CYCLOSOPHORAOSES//LBMPS//NOD GENES |
| 10 | 0.0000063526 | AZOSPIRILLUM//AZOSPIRILLUM BRASILENSE//BIOCHEM PHYSIOL PLANTS MICROORGANISMS |