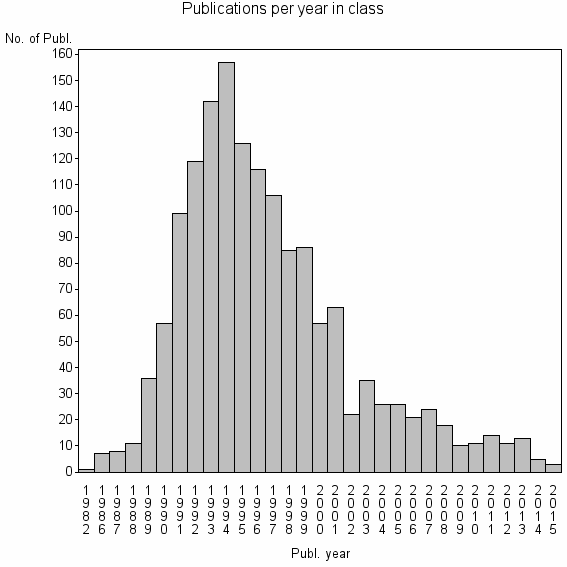
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 5742 | 1515 | 28.6 | 80% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | BIBAC | Author keyword | 21 | 90% | 1% | 9 |
| 2 | TAR CLONING | Author keyword | 7 | 64% | 0% | 7 |
| 3 | BIBAC LIBRARY | Author keyword | 6 | 100% | 0% | 4 |
| 4 | YEAST ARTIFICIAL CHROMOSOME | Author keyword | 5 | 25% | 1% | 17 |
| 5 | CLONE ORDERING | Author keyword | 4 | 75% | 0% | 3 |
| 6 | TRANSFORMATION COMPETENT ARTIFICIAL CHROMOSOME | Author keyword | 4 | 75% | 0% | 3 |
| 7 | YEAST ARTIFICIAL CHROMOSOME LIBRARY | Author keyword | 4 | 75% | 0% | 3 |
| 8 | BAC LIBRARY | Author keyword | 4 | 14% | 2% | 24 |
| 9 | BACTERIAL ARTIFICIAL CHROMOSOME BAC LIBRARY | Author keyword | 4 | 46% | 0% | 6 |
| 10 | GENETIC ANALYSIS-BIOMOLECULAR ENGINEERING | Journal | 3 | 11% | 2% | 29 |
Web of Science journal categories |
Author Key Words |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | YEAST ARTIFICIAL CHROMOSOMES | 56 | 37% | 8% | 121 |
| 2 | ARTIFICIAL CHROMOSOME LIBRARY | 33 | 29% | 6% | 97 |
| 3 | HUMAN XQ24 XQ28 | 23 | 76% | 1% | 16 |
| 4 | LARGE SEGMENTS | 22 | 68% | 1% | 19 |
| 5 | FACTOR BASED VECTOR | 21 | 47% | 2% | 34 |
| 6 | GENOME EQUIVALENTS | 21 | 78% | 1% | 14 |
| 7 | LARGE INSERTS | 20 | 62% | 1% | 21 |
| 8 | CONTIGS | 19 | 55% | 2% | 24 |
| 9 | PHYSICAL MAP | 16 | 11% | 9% | 135 |
| 10 | TRANSCRIBED SEQUENCES | 15 | 41% | 2% | 29 |
Journals |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | GENETIC ANALYSIS-BIOMOLECULAR ENGINEERING | 3 | 11% | 2% | 29 |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Advances in BAC-Based Physical Mapping and Map Integration Strategies in Plants | 2012 | 8 | 87 | 49% |
| Mapping and sequencing complex genomes: Let's get physical! | 2004 | 48 | 49 | 65% |
| Size matters: Use of YACs, BACs and PACs in transgenic animals | 2001 | 159 | 164 | 24% |
| YACS, BACS, PACS AND MACS - ARTIFICIAL CHROMOSOMES AS RESEARCH TOOLS | 1994 | 75 | 40 | 55% |
| BAC as tools for genome sequencing | 2001 | 35 | 41 | 63% |
| Zebrafish YAC, BAG, and PAC genomic libraries | 1999 | 33 | 101 | 51% |
| Strategies for the systematic sequencing of complex genomes | 2001 | 64 | 77 | 34% |
| Bacterial Artificial Chromosome Libraries of Pulse Crops: Characteristics and Applications | 2012 | 3 | 60 | 35% |
| Artificial chromosome-based transgenes in the study of genome function | 2006 | 19 | 130 | 19% |
| Cosmid-based exon trapping | 1999 | 1 | 7 | 86% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ARIZONA GENOM COMPUTAT | 3 | 32% | 0.5% | 7 |
| 2 | ARIZONA GENOM | 1 | 10% | 0.9% | 13 |
| 3 | FO TRY FRUIT | 1 | 50% | 0.1% | 2 |
| 4 | INSERM U184 11 RUE HUMANN | 1 | 100% | 0.1% | 2 |
| 5 | LGME CNRS | 1 | 100% | 0.1% | 2 |
| 6 | MED GENET MED MOLEC MICROBIOL | 1 | 100% | 0.1% | 2 |
| 7 | MULTIMEGABASE SEQUENCING | 1 | 50% | 0.1% | 2 |
| 8 | ABT ZUCHTUNGSFOR ERTRAGSPHYSIOL | 1 | 50% | 0.1% | 1 |
| 9 | ARIZONA GEN | 1 | 50% | 0.1% | 1 |
| 10 | BAC EST OURCE | 1 | 50% | 0.1% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000214042 | HAPPY MAPPING//MULTILOCUS ORDERING//EXOGAMIC POPULATIONS |
| 2 | 0.0000131068 | RICE GENOME PROGRAM//CROP DESIGN//GENOME IL |
| 3 | 0.0000107188 | RECOMBINEERING//LAMBDA RED//LAMBDA RED RECOMBINATION |
| 4 | 0.0000101573 | SPECTRAL SIMILARITY MAPPING//FAST FISH//CHROMOSOME SPREADING |
| 5 | 0.0000100820 | MKIAA//SMART CDNA LIBRARY//CDNA SEQUENCING |
| 6 | 0.0000094709 | GENOME WALKING//FALSE POSITIVE EXCLUSION//UNKNOWN FLANKING SEQUENCES |
| 7 | 0.0000092516 | HN1//CANC 6401 E THOMAS RD//HORMONAL PROFILING |
| 8 | 0.0000087733 | RBM5//HUMAN CHROMOSOME 3//LUCA 15 |
| 9 | 0.0000076230 | MOL IMMUNOL RUMENTAT//GENOMIC MATCHING TECHNIQUE//HLA A ALLELES |
| 10 | 0.0000073528 | DEV BIOL PROGRAM OBSTET GYNECOL//HUMAN GENOME ORGANIZATION//KLEISLI |