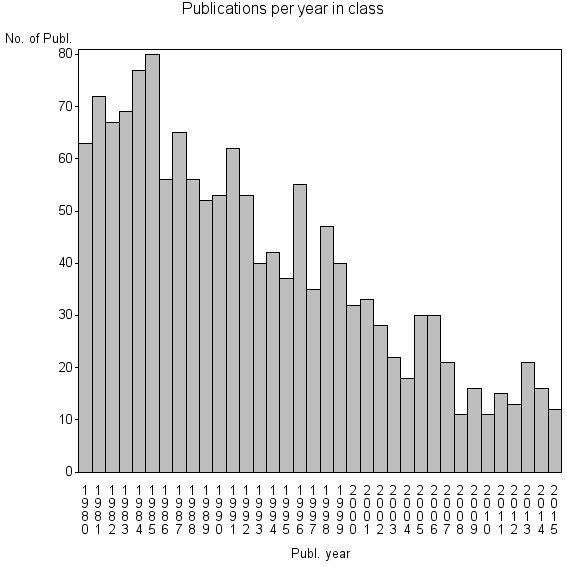
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 6207 | 1450 | 39.0 | 60% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 302 | 17871 | TRANSLESION SYNTHESIS//DNA POLYMERASE//RECA PROTEIN |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | SOS MUTAGENESIS | Author keyword | 44 | 77% | 2% | 30 |
| 2 | UMUDC | Author keyword | 34 | 70% | 2% | 28 |
| 3 | LEXA REPRESSOR | Author keyword | 32 | 85% | 1% | 17 |
| 4 | MUCAB | Author keyword | 26 | 87% | 1% | 13 |
| 5 | UMUD | Author keyword | 19 | 80% | 1% | 12 |
| 6 | SECT DNA REPLICAT REPAIR MUTAGENESIS | Address | 16 | 41% | 2% | 30 |
| 7 | LEXA | Author keyword | 15 | 47% | 2% | 23 |
| 8 | SOS BOX | Author keyword | 13 | 71% | 1% | 10 |
| 9 | SOS RESPONSE | Author keyword | 12 | 20% | 4% | 52 |
| 10 | MOL MICROBIOL BACTERIAL GENET GRP | Address | 11 | 100% | 0% | 6 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | SOS MUTAGENESIS | 44 | 77% | 2% | 30 | Search SOS+MUTAGENESIS | Search SOS+MUTAGENESIS |
| 2 | UMUDC | 34 | 70% | 2% | 28 | Search UMUDC | Search UMUDC |
| 3 | LEXA REPRESSOR | 32 | 85% | 1% | 17 | Search LEXA+REPRESSOR | Search LEXA+REPRESSOR |
| 4 | MUCAB | 26 | 87% | 1% | 13 | Search MUCAB | Search MUCAB |
| 5 | UMUD | 19 | 80% | 1% | 12 | Search UMUD | Search UMUD |
| 6 | LEXA | 15 | 47% | 2% | 23 | Search LEXA | Search LEXA |
| 7 | SOS BOX | 13 | 71% | 1% | 10 | Search SOS+BOX | Search SOS+BOX |
| 8 | SOS RESPONSE | 12 | 20% | 4% | 52 | Search SOS+RESPONSE | Search SOS+RESPONSE |
| 9 | UMUC | 10 | 52% | 1% | 13 | Search UMUC | Search UMUC |
| 10 | SOS SYSTEM | 9 | 59% | 1% | 10 | Search SOS+SYSTEM | Search SOS+SYSTEM |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | SOS MUTAGENESIS | 141 | 73% | 7% | 108 |
| 2 | UV MUTAGENESIS | 68 | 57% | 6% | 81 |
| 3 | UMUD | 42 | 72% | 2% | 33 |
| 4 | INTACT UMUD | 41 | 100% | 1% | 15 |
| 5 | AUTODIGESTION | 37 | 81% | 2% | 22 |
| 6 | ULTRAVIOLET LIGHT MUTAGENESIS | 33 | 100% | 1% | 13 |
| 7 | RECA PROTEIN | 32 | 19% | 10% | 150 |
| 8 | POL V | 23 | 62% | 2% | 24 |
| 9 | GROE MUTANTS | 22 | 81% | 1% | 13 |
| 10 | INDUCIBLE MUTAGENESIS | 21 | 78% | 1% | 14 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| The bacterial LexA transcriptional repressor | 2009 | 91 | 120 | 60% |
| Inducible SOS Response System of DNA Repair and Mutagenesis in Escherichia coli | 2008 | 68 | 32 | 75% |
| Aeons of distress: an evolutionary perspective on the bacterial SOS response | 2007 | 103 | 135 | 53% |
| Mutagenesis and more: umuDC and the Escherichia coli SOS response | 1998 | 101 | 101 | 69% |
| A new model for SOS-induced mutagenesis: how RecA protein activates DNA polymerase V | 2010 | 33 | 75 | 49% |
| The SOS response: Recent insights into umuDC-dependent mutagenesis and DNA damage tolerance | 2000 | 163 | 85 | 58% |
| MUTAGENESIS AND INDUCIBLE RESPONSES TO DEOXYRIBONUCLEIC-ACID DAMAGE IN ESCHERICHIA-COLI | 1984 | 1711 | 229 | 48% |
| Y-family DNA polymerases in Escherichia coli | 2007 | 68 | 73 | 45% |
| MUTAGENESIS INDUCED BY BACTERIAL UMUDC PROTEINS AND THEIR PLASMID HOMOLOGS | 1992 | 68 | 43 | 88% |
| SOS, the formidable strategy of bacteria against aggressions | 2014 | 8 | 187 | 21% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | SECT DNA REPLICAT REPAIR MUTAGENESIS | 16 | 41% | 2.1% | 30 |
| 2 | MOL MICROBIOL BACTERIAL GENET GRP | 11 | 100% | 0.4% | 6 |
| 3 | BIOMED PLICAT GRP | 5 | 43% | 0.6% | 9 |
| 4 | GENOM INTEGR | 3 | 23% | 0.8% | 11 |
| 5 | HEDCO MOL BIOL S | 3 | 24% | 0.7% | 10 |
| 6 | SERV MICROBIOL INIBIC | 1 | 31% | 0.3% | 4 |
| 7 | ST PETERSBURG KONSTANTINOV NUCL PHYS | 1 | 21% | 0.2% | 3 |
| 8 | BIOCHEM BIOPHYS RAKOWIECKA 36 | 1 | 50% | 0.1% | 1 |
| 9 | BIOL 68 633 | 1 | 50% | 0.1% | 1 |
| 10 | BIOMED PL GRP | 1 | 50% | 0.1% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000255217 | RECA PROTEIN//RECA//DNA STRAND EXCHANGE |
| 2 | 0.0000202502 | UVRB//UVRABC//UVRC |
| 3 | 0.0000165552 | CLAMP LOADER//REPLICATION FACTOR C//DNA REPLICAT |
| 4 | 0.0000161791 | ADAPTIVE MUTATION//STATIONARY PHASE MUTATION//STATIONARY PHASE MUTAGENESIS |
| 5 | 0.0000142897 | HOLLIDAY JUNCTION//RECBCD ENZYME//RECBCD |
| 6 | 0.0000126767 | UMU TEST//SOS CHROMOTEST//COMPARISON ON MUTAGENICITY |
| 7 | 0.0000110575 | REX EXCLUSION//BACTERIOPHAGE LAMBDA LAMBDA//CI REXA REXB OPERON |
| 8 | 0.0000108044 | TRANSLESION SYNTHESIS//DNA POLYMERASE ETA//DNA DAMAGE TOLERANCE |
| 9 | 0.0000098098 | T OLIGOS//T OLIGO//MER PHENOTYPE |
| 10 | 0.0000087473 | S METHYL METHANETHIOSULFONATE//BIO ANTIMUTAGEN//DIAGNOST TOXICIDADE GENET |