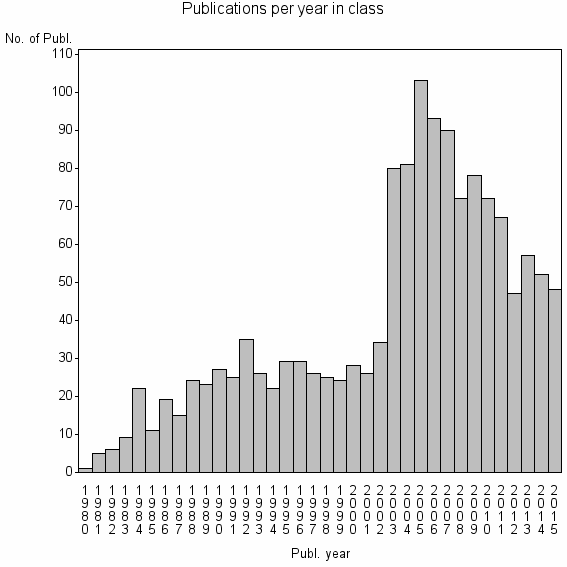
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 6346 | 1431 | 30.2 | 67% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 238 | 19563 | BIOINFORMATICS//BMC BIOINFORMATICS//GENOME RESEARCH |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | MULTIPLE SEQUENCE ALIGNMENT | Author keyword | 33 | 27% | 7% | 106 |
| 2 | T COFFEE | Author keyword | 21 | 90% | 1% | 9 |
| 3 | STATISTICAL ALIGNMENT | Author keyword | 18 | 83% | 1% | 10 |
| 4 | SPACED SEEDS | Author keyword | 15 | 73% | 1% | 11 |
| 5 | SEQUENCE ALIGNMENT | Author keyword | 10 | 11% | 6% | 81 |
| 6 | INSERTION DELETION MODEL | Author keyword | 6 | 80% | 0% | 4 |
| 7 | INTERDISCIPLINARY STUDIES STRUCT FORMAT | Address | 6 | 71% | 0% | 5 |
| 8 | WHOLE GENOME ALIGNMENTS | Author keyword | 6 | 100% | 0% | 4 |
| 9 | ALIGNMENT UNCERTAINTY | Author keyword | 4 | 75% | 0% | 3 |
| 10 | DEP LENGUAJES CIENCIAS COMPUTAC | Address | 4 | 75% | 0% | 3 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | MULTIPLE SEQUENCE ALIGNMENT | 33 | 27% | 7% | 106 | Search MULTIPLE+SEQUENCE+ALIGNMENT | Search MULTIPLE+SEQUENCE+ALIGNMENT |
| 2 | T COFFEE | 21 | 90% | 1% | 9 | Search T+COFFEE | Search T+COFFEE |
| 3 | STATISTICAL ALIGNMENT | 18 | 83% | 1% | 10 | Search STATISTICAL+ALIGNMENT | Search STATISTICAL+ALIGNMENT |
| 4 | SPACED SEEDS | 15 | 73% | 1% | 11 | Search SPACED+SEEDS | Search SPACED+SEEDS |
| 5 | SEQUENCE ALIGNMENT | 10 | 11% | 6% | 81 | Search SEQUENCE+ALIGNMENT | Search SEQUENCE+ALIGNMENT |
| 6 | INSERTION DELETION MODEL | 6 | 80% | 0% | 4 | Search INSERTION+DELETION+MODEL | Search INSERTION+DELETION+MODEL |
| 7 | WHOLE GENOME ALIGNMENTS | 6 | 100% | 0% | 4 | Search WHOLE+GENOME+ALIGNMENTS | Search WHOLE+GENOME+ALIGNMENTS |
| 8 | ALIGNMENT UNCERTAINTY | 4 | 75% | 0% | 3 | Search ALIGNMENT+UNCERTAINTY | Search ALIGNMENT+UNCERTAINTY |
| 9 | PAIR HIDDEN MARKOV MODELS | 4 | 75% | 0% | 3 | Search PAIR+HIDDEN+MARKOV+MODELS | Search PAIR+HIDDEN+MARKOV+MODELS |
| 10 | SUBSTITUTION SCORE MATRICES | 4 | 75% | 0% | 3 | Search SUBSTITUTION+SCORE+MATRICES | Search SUBSTITUTION+SCORE+MATRICES |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MAXIMUM LIKELIHOOD ALIGNMENT | 28 | 69% | 2% | 24 |
| 2 | DIALIGN | 25 | 65% | 2% | 24 |
| 3 | SPACED SEEDS | 21 | 61% | 2% | 23 |
| 4 | STATISTICAL ALIGNMENT | 18 | 89% | 1% | 8 |
| 5 | JOINT BAYESIAN ESTIMATION | 17 | 75% | 1% | 12 |
| 6 | HOMOLOGY SEARCH | 15 | 42% | 2% | 28 |
| 7 | SP SCORE | 15 | 82% | 1% | 9 |
| 8 | LINEAR SPACE | 15 | 34% | 3% | 37 |
| 9 | STATISTICAL MULTIPLE ALIGNMENT | 15 | 88% | 0% | 7 |
| 10 | BALIBASE | 14 | 59% | 1% | 16 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Recent developments in the MAFFT multiple sequence alignment program | 2008 | 1326 | 101 | 60% |
| Multiple sequence alignment | 2006 | 116 | 35 | 66% |
| Multiple sequence alignment for phylogenetic purposes | 2006 | 68 | 316 | 49% |
| Upcoming challenges for multiple sequence alignment methods in the high-throughput era | 2009 | 64 | 59 | 64% |
| Multiple sequence alignments | 2005 | 47 | 40 | 73% |
| Multiple sequence alignment: In pursuit of homologous DNA positions | 2007 | 57 | 88 | 44% |
| PROTEIN MULTIPLE SEQUENCE ALIGNMENT AND FLEXIBLE PATTERN-MATCHING | 1990 | 153 | 18 | 67% |
| Quality assessment of multiple alignment programs | 2002 | 89 | 14 | 71% |
| Multiple protein sequence alignment | 2008 | 24 | 42 | 64% |
| Comparison of genomic DNA sequences: solved and unsolved problems | 2001 | 74 | 51 | 43% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | INTERDISCIPLINARY STUDIES STRUCT FORMAT | 6 | 71% | 0.3% | 5 |
| 2 | DEP LENGUAJES CIENCIAS COMPUTAC | 4 | 75% | 0.2% | 3 |
| 3 | COMPARAT BIOINFORMAT GRP | 3 | 57% | 0.3% | 4 |
| 4 | INFORMAT INTERDISZIPLINA | 2 | 67% | 0.1% | 2 |
| 5 | EDUC TECHNOL COMP | 1 | 100% | 0.1% | 2 |
| 6 | GENET BIOL MOL CELLULAIRE BIOL GEN | 1 | 50% | 0.1% | 2 |
| 7 | HUMAN GENET IGH | 1 | 50% | 0.1% | 2 |
| 8 | STRATEG TRAINING PROGRAM BIOINFORMAT | 1 | 100% | 0.1% | 2 |
| 9 | DEP ESTADIST | 1 | 30% | 0.2% | 3 |
| 10 | UR888 | 1 | 23% | 0.2% | 3 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000164965 | PROTEIN STRUCTURE PREDICTION//FOLD RECOGNITION//CASP |
| 2 | 0.0000138177 | LONGEST COMMON SUBSEQUENCE//CONSTRAINED LONGEST COMMON SUBSEQUENCE//SHORTEST COMMON SUPERSTRING |
| 3 | 0.0000111350 | IROQUOIS//CONSERVED NONCODING ELEMENTS//ENERGY ENVIRONM BIOL COMP |
| 4 | 0.0000104762 | GENE PREDICTION//GENE FINDING//GENE RECOGNITION |
| 5 | 0.0000103878 | CIRCOS//BIODADOS//ORTHOLOGY |
| 6 | 0.0000100702 | CODON MODEL//HETEROTACHY//PHYLOGENETIC INVARIANTS |
| 7 | 0.0000098949 | GENOME REARRANGEMENTS//SORTING BY REVERSALS//BREAKPOINT GRAPH |
| 8 | 0.0000092117 | AMBIGRAM//DESK TOP MOLECULAR MODELING//PARTIAL DIGEST |
| 9 | 0.0000077588 | K MAXIMUM SUMS PROBLEM//MAXIMUM SUM PROBLEM//MAXIMUM SUM SEGMENT |
| 10 | 0.0000072414 | EMBL OUTSTN//PROT INFORMAT OURCE//HLTH SCI TECHNOL ICIR |