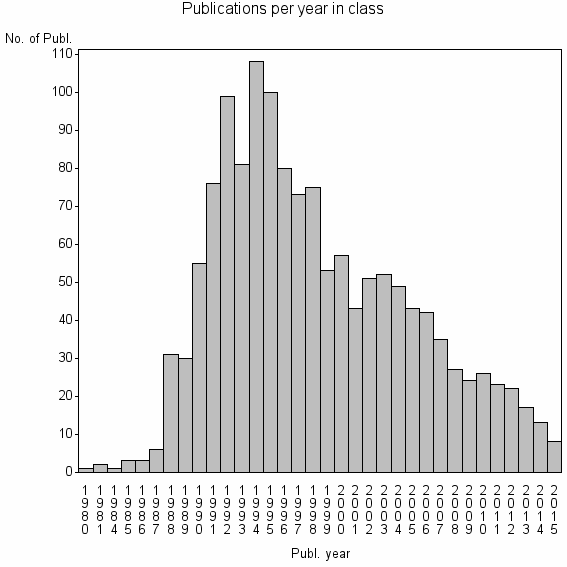
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 6492 | 1409 | 33.8 | 78% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 751 | 11810 | ATOMIC FORCE MICROSCOPE//DISSENY PROGRAMACIO SISTEMES ELECT//LOCAL ANODIC OXIDATION |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | DNA NETWORK | Author keyword | 4 | 75% | 0% | 3 |
| 2 | FOR UNGSZENTRUM IBI STRUCT BIOL 2 | Address | 4 | 75% | 0% | 3 |
| 3 | CRYO AFM | Author keyword | 3 | 100% | 0% | 3 |
| 4 | GIOVANNI ESPOSITO GEN PHYSIOL BIOL CHEM | Address | 3 | 100% | 0% | 3 |
| 5 | URA 5027 | Address | 3 | 50% | 0% | 4 |
| 6 | TAKATSUKA UNIT | Address | 3 | 60% | 0% | 3 |
| 7 | GDCB | Address | 2 | 44% | 0% | 4 |
| 8 | BIOZENTRUM ME MULLER MICROSCOP STRUCT BIOL | Address | 2 | 67% | 0% | 2 |
| 9 | CHANG CONJUNCTIVAL CELLS | Author keyword | 2 | 67% | 0% | 2 |
| 10 | DNA CONTOUR LENGTH | Author keyword | 2 | 67% | 0% | 2 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | DNA NETWORK | 4 | 75% | 0% | 3 | Search DNA+NETWORK | Search DNA+NETWORK |
| 2 | CRYO AFM | 3 | 100% | 0% | 3 | Search CRYO+AFM | Search CRYO+AFM |
| 3 | CHANG CONJUNCTIVAL CELLS | 2 | 67% | 0% | 2 | Search CHANG+CONJUNCTIVAL+CELLS | Search CHANG+CONJUNCTIVAL+CELLS |
| 4 | DNA CONTOUR LENGTH | 2 | 67% | 0% | 2 | Search DNA+CONTOUR+LENGTH | Search DNA+CONTOUR+LENGTH |
| 5 | SEMICONTACT MODE | 2 | 67% | 0% | 2 | Search SEMICONTACT+MODE | Search SEMICONTACT+MODE |
| 6 | SFM | 2 | 11% | 1% | 16 | Search SFM | Search SFM |
| 7 | 11 MERCAPTOUNDECANOIC ACID MUDA | 1 | 100% | 0% | 2 | Search 11+MERCAPTOUNDECANOIC+ACID+MUDA | Search 11+MERCAPTOUNDECANOIC+ACID+MUDA |
| 8 | BAL 31 NUCLEASE | 1 | 50% | 0% | 2 | Search BAL+31+NUCLEASE | Search BAL+31+NUCLEASE |
| 9 | CRUCIFORM CONFORMATION | 1 | 100% | 0% | 2 | Search CRUCIFORM+CONFORMATION | Search CRUCIFORM+CONFORMATION |
| 10 | DNA DEPOSITION | 1 | 50% | 0% | 2 | Search DNA+DEPOSITION | Search DNA+DEPOSITION |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | TUNNELLING MICROSCOPY | 29 | 45% | 3% | 48 |
| 2 | WET INSULATORS | 15 | 82% | 1% | 9 |
| 3 | RECA DNA COMPLEXES | 13 | 61% | 1% | 14 |
| 4 | MOLECULAR RESOLUTION | 8 | 31% | 2% | 22 |
| 5 | INTERCALATION INDUCED CHANGES | 7 | 64% | 0% | 7 |
| 6 | BACTERIAL SURFACE PROTEIN | 5 | 47% | 1% | 8 |
| 7 | MOLECULAR RESOLUTION IMAGES | 5 | 29% | 1% | 14 |
| 8 | AFM SAMPLES | 4 | 75% | 0% | 3 |
| 9 | BACTERIOPHAGE T4 POLYHEADS | 4 | 75% | 0% | 3 |
| 10 | BIOLOGICAL MACROMOLECULAR STRUCTURES | 4 | 75% | 0% | 3 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| BIOMOLECULAR IMAGING WITH THE ATOMIC-FORCE MICROSCOPE | 1994 | 442 | 102 | 69% |
| Imaging of nucleic acids with atomic force microscopy | 2011 | 23 | 97 | 48% |
| Surface biology of DNA by atomic force microscopy | 2001 | 180 | 141 | 37% |
| Preparation of DNA and nucleoprotein samples for AFM imaging | 2011 | 16 | 97 | 55% |
| Atomic force microscopy of biological samples | 2010 | 50 | 146 | 28% |
| An overview of the biophysical applications of atomic force microscopy | 2004 | 106 | 178 | 32% |
| Single-molecule studies of DNA transcription using atomic force microscopy | 2012 | 7 | 70 | 44% |
| AFM: a versatile tool in biophysics | 2005 | 136 | 244 | 22% |
| Properties of biomolecules measured from atomic force microscope images: A review | 1997 | 151 | 44 | 59% |
| DNA structure and dynamics - An atomic force microscopy study | 2004 | 37 | 101 | 50% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | FOR UNGSZENTRUM IBI STRUCT BIOL 2 | 4 | 75% | 0.2% | 3 |
| 2 | GIOVANNI ESPOSITO GEN PHYSIOL BIOL CHEM | 3 | 100% | 0.2% | 3 |
| 3 | URA 5027 | 3 | 50% | 0.3% | 4 |
| 4 | TAKATSUKA UNIT | 3 | 60% | 0.2% | 3 |
| 5 | GDCB | 2 | 44% | 0.3% | 4 |
| 6 | BIOZENTRUM ME MULLER MICROSCOP STRUCT BIOL | 2 | 67% | 0.1% | 2 |
| 7 | EA 3637 | 2 | 67% | 0.1% | 2 |
| 8 | IMMUNOCHIM PATOL MOL | 2 | 67% | 0.1% | 2 |
| 9 | UMR CNRS IGR UPS 8126 | 1 | 100% | 0.1% | 2 |
| 10 | BIOX LIFE SCI | 1 | 10% | 0.7% | 10 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000201434 | TIP CHARACTERIZATION//TIP CHARACTERIZER//SIDEWALL MEASUREMENT |
| 2 | 0.0000157906 | ORIENTING SUBSTRATES//ASET SUMITOMO CHEM//PTFE FILMS |
| 3 | 0.0000128006 | UNITE CHIM INTER ES//SINGLE CELL FORCE SPECTROSCOPY//BACTERIAL INTERACTION FORCES |
| 4 | 0.0000127741 | QUANTUM SCI GRP//AVADH BHATIA PHYS//GARCIA MRSEC POLYMERS ENGINEERED INTER ES |
| 5 | 0.0000126217 | CELL MECHANICS//MICROPIPETTE ASPIRATION//CELL STIFFNESS |
| 6 | 0.0000124055 | LOCAL TUNNELING BARRIER HEIGHT//LOCAL TUNNELING BARRIER HEIGHT LBH//I S CHARACTERISTICS |
| 7 | 0.0000118385 | DISSENY PROGRAMACIO SISTEMES ELECT//FUTURE ENERGY IFES//QPLUS |
| 8 | 0.0000111975 | CHEM NANOSCI S//MOLECULAR COMBING//DNA TEMPLATE |
| 9 | 0.0000099971 | SINGLE MOLECULE FORCE SPECTROSCOPY//MECHANICAL UNFOLDING//FORCE SPECTROSCOPY |
| 10 | 0.0000073280 | CARBON NANOTUBE PROBES//CARBON NANOTUBE TIPS//HIGH DENSITY INFORMATION STORAGE |