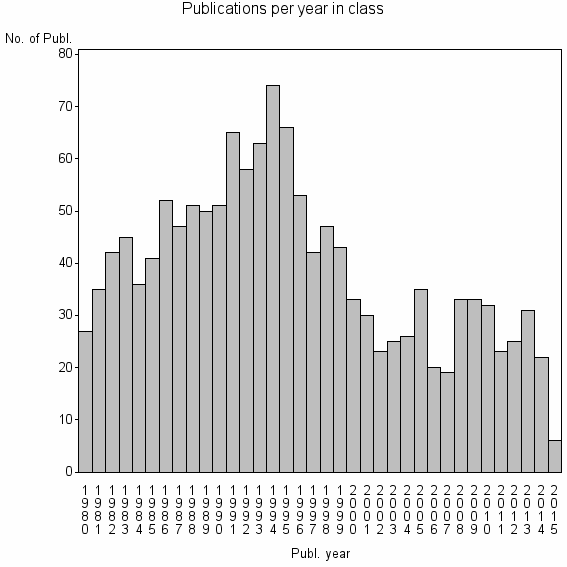
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 6532 | 1404 | 38.7 | 67% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 501 | 14604 | RNA POLYMERASE//SITE SPECIFIC RECOMBINATION//H NS |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | TRP REPRESSOR | Author keyword | 42 | 76% | 2% | 29 |
| 2 | LAC REPRESSOR | Author keyword | 31 | 48% | 3% | 48 |
| 3 | LACTOSE REPRESSOR | Author keyword | 26 | 100% | 1% | 11 |
| 4 | TRP OPERATOR | Author keyword | 8 | 75% | 0% | 6 |
| 5 | GAL REPRESSOR | Author keyword | 8 | 100% | 0% | 5 |
| 6 | LAMBDA REPRESSOR | Author keyword | 7 | 36% | 1% | 15 |
| 7 | DNA LOOPING | Author keyword | 6 | 22% | 2% | 22 |
| 8 | LACTOSE REPRESSOR PROTEIN | Author keyword | 6 | 100% | 0% | 4 |
| 9 | INTEGRAT BIOL MODELING | Address | 5 | 60% | 0% | 6 |
| 10 | LAC OPERATOR | Author keyword | 5 | 38% | 1% | 11 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | TRP REPRESSOR | 42 | 76% | 2% | 29 | Search TRP+REPRESSOR | Search TRP+REPRESSOR |
| 2 | LAC REPRESSOR | 31 | 48% | 3% | 48 | Search LAC+REPRESSOR | Search LAC+REPRESSOR |
| 3 | LACTOSE REPRESSOR | 26 | 100% | 1% | 11 | Search LACTOSE+REPRESSOR | Search LACTOSE+REPRESSOR |
| 4 | TRP OPERATOR | 8 | 75% | 0% | 6 | Search TRP+OPERATOR | Search TRP+OPERATOR |
| 5 | GAL REPRESSOR | 8 | 100% | 0% | 5 | Search GAL+REPRESSOR | Search GAL+REPRESSOR |
| 6 | LAMBDA REPRESSOR | 7 | 36% | 1% | 15 | Search LAMBDA+REPRESSOR | Search LAMBDA+REPRESSOR |
| 7 | DNA LOOPING | 6 | 22% | 2% | 22 | Search DNA+LOOPING | Search DNA+LOOPING |
| 8 | LACTOSE REPRESSOR PROTEIN | 6 | 100% | 0% | 4 | Search LACTOSE+REPRESSOR+PROTEIN | Search LACTOSE+REPRESSOR+PROTEIN |
| 9 | LAC OPERATOR | 5 | 38% | 1% | 11 | Search LAC+OPERATOR | Search LAC+OPERATOR |
| 10 | DNA HELICAL REPEAT | 4 | 75% | 0% | 3 | Search DNA+HELICAL+REPEAT | Search DNA+HELICAL+REPEAT |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | APOREPRESSOR | 99 | 80% | 4% | 61 |
| 2 | LACTOSE REPRESSOR | 70 | 82% | 3% | 41 |
| 3 | HEADPIECE | 38 | 76% | 2% | 26 |
| 4 | LAC REPRESSOR | 34 | 20% | 11% | 156 |
| 5 | OPERATOR DNA | 34 | 47% | 4% | 53 |
| 6 | CI REPRESSOR | 32 | 65% | 2% | 30 |
| 7 | GAL REPRESSOR | 30 | 79% | 1% | 19 |
| 8 | HINGE HELIX | 29 | 88% | 1% | 14 |
| 9 | PHAGE REPRESSOR | 26 | 80% | 1% | 16 |
| 10 | COUPLED ENERGETICS | 24 | 91% | 1% | 10 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| The Chemistry of Regulation of Genes and Other Things | 2014 | 7 | 67 | 31% |
| The lactose repressor system: paradigms for regulation, allosteric behavior and protein folding | 2007 | 67 | 97 | 69% |
| The lac repressor | 2005 | 107 | 47 | 72% |
| Allostery in the Lacl/GaIR family: variations on a theme | 2009 | 39 | 66 | 53% |
| THE HELIX-TURN-HELIX DNA-BINDING MOTIF | 1989 | 459 | 32 | 81% |
| A Tale of Two Repressors | 2011 | 9 | 29 | 72% |
| Lactose repressor protein: Functional properties and structure | 1998 | 41 | 151 | 77% |
| DNA RECOGNITION BY PROTEINS WITH THE HELIX-TURN-HELIX MOTIF | 1990 | 503 | 86 | 58% |
| DNA looping in gene regulation: from the assembly of macromolecular complexes to the control of transcriptional noise | 2005 | 84 | 31 | 45% |
| PROTEIN-DNA RECOGNITION | 1984 | 1400 | 45 | 44% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | INTEGRAT BIOL MODELING | 5 | 60% | 0.4% | 6 |
| 2 | WM KECK INTERDISCIPLINARY BIOSCI TRAINING | 3 | 100% | 0.2% | 3 |
| 3 | MODELS LIFE | 3 | 17% | 1.0% | 14 |
| 4 | BIOCHEM CELL BIOCHEM | 1 | 50% | 0.1% | 1 |
| 5 | CNRSUMR 7196 | 1 | 50% | 0.1% | 1 |
| 6 | DOSIMETRIE RAYONNEMENTS | 1 | 50% | 0.1% | 1 |
| 7 | INTERDISCIPLINAIRE RECH ION LASERS | 1 | 50% | 0.1% | 1 |
| 8 | KECK INTERDISCIPLINARY BIOSCI TRAINING | 1 | 50% | 0.1% | 1 |
| 9 | WRIGHT RIEMEN S | 1 | 50% | 0.1% | 1 |
| 10 | MOL BIOMED SCI BIOCHEM | 1 | 14% | 0.3% | 4 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000199176 | CYCLIC AMP RECEPTOR PROTEIN//CATABOLITE ACTIVATOR PROTEIN CAP//CAMP RECEPTOR PROTEIN |
| 2 | 0.0000114046 | RETARDATION ANALYSIS//NONSPECIFIC PROTEIN DNA COMPLEXES//GEL CAGE |
| 3 | 0.0000106796 | NARROW ESCAPE//DEN DANS DM2S SERMA LTSD//NANOMED MECH BIOL |
| 4 | 0.0000084443 | RNA POLYMERASE//ESCHERICHIA COLI RNA POLYMERASE//SIGMA70 |
| 5 | 0.0000072954 | SKIRBALL PROGRAM MOL PATHOGENESIS//BACTERIOPHAGE P2//RETRON |
| 6 | 0.0000069771 | DNA CURVATURE//A TRACT//DNA FLEXIBILITY |
| 7 | 0.0000069454 | REX EXCLUSION//BACTERIOPHAGE LAMBDA LAMBDA//CI REXA REXB OPERON |
| 8 | 0.0000068547 | INTERACTION PROPENSITY//RNA BINDING RESIDUES//COMPUTAT INTELLIGENCE LEARNING DISCOVERY |
| 9 | 0.0000066553 | TYRR//SHIKIMIC ACID PRODUCTION//CIS CIS MUCONIC ACID |
| 10 | 0.0000061031 | CARAB//PYRIMIDINE REGULATION//PYRR |