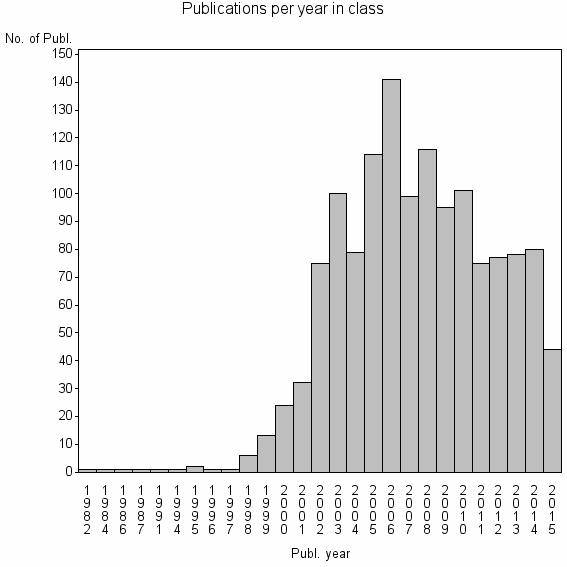
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 6858 | 1359 | 31.4 | 65% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 1650 | 6377 | BICLUSTERING//PROGRAM TRANSLAT IMMUNOVIROL BIODEF//MAYO VACCINE GRP |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | BICLUSTERING | Author keyword | 36 | 35% | 6% | 84 |
| 2 | GENE EXPRESSION DATA | Author keyword | 11 | 16% | 4% | 59 |
| 3 | GENE EXPRESSION DATA ANALYSIS | Author keyword | 9 | 40% | 1% | 17 |
| 4 | ORDER PRESERVING SUBMATRIX | Author keyword | 6 | 80% | 0% | 4 |
| 5 | TIME COURSE GENE EXPRESSION | Author keyword | 5 | 63% | 0% | 5 |
| 6 | GENE EXPRESSION DATA CLUSTERING | Author keyword | 4 | 67% | 0% | 4 |
| 7 | GENE EXPRESSION MICROARRAY DATA | Author keyword | 4 | 67% | 0% | 4 |
| 8 | PLAID MODEL | Author keyword | 4 | 75% | 0% | 3 |
| 9 | BIOLOGICAL SIGNIFICANCE TEST | Author keyword | 3 | 100% | 0% | 3 |
| 10 | TIME COURSE MICROARRAY | Author keyword | 3 | 50% | 0% | 4 |
Web of Science journal categories |
Author Key Words |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | GENE EXPRESSION DATA | 27 | 11% | 17% | 227 |
| 2 | COURSE MICROARRAY EXPERIMENTS | 22 | 58% | 2% | 25 |
| 3 | CYCLE REGULATED GENES | 10 | 34% | 2% | 24 |
| 4 | BICLUSTERING ALGORITHMS | 7 | 67% | 0% | 6 |
| 5 | TRANSCRIPTIONAL PROGRAM | 5 | 15% | 2% | 28 |
| 6 | CLUSTERING ANALYSIS | 4 | 19% | 1% | 17 |
| 7 | CLASS DISCOVERY | 3 | 13% | 2% | 22 |
| 8 | C MEANS METHOD | 1 | 50% | 0% | 2 |
| 9 | COMPUTATIONAL CLUSTER VALIDATION | 1 | 50% | 0% | 2 |
| 10 | PERIODICALLY EXPRESSED GENES | 1 | 50% | 0% | 2 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| STUDY DESIGNS Studying and modelling dynamic biological processes using time-series gene expression data | 2012 | 55 | 103 | 28% |
| Computational cluster validation in post-genomic data analysis | 2005 | 286 | 37 | 46% |
| A roadmap of clustering algorithms: finding a match for a biomedical application | 2009 | 47 | 89 | 31% |
| Analyzing time series gene expression data | 2004 | 194 | 37 | 54% |
| Analysis of time-series gene expression data: Methods, challenges, and opportunities | 2007 | 46 | 100 | 28% |
| Clustering of high throughput gene expression data | 2012 | 5 | 97 | 39% |
| Towards knowledge-based gene expression data mining | 2007 | 26 | 48 | 29% |
| Clustering microarray data | 2006 | 21 | 20 | 50% |
| Analyzing microarray data using cluster analysis | 2003 | 72 | 14 | 36% |
| Basic microarray analysis: grouping and feature reduction | 2001 | 95 | 12 | 42% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | LATICE | 3 | 60% | 0.2% | 3 |
| 2 | GRP GENE COMPUTAT | 2 | 67% | 0.1% | 2 |
| 3 | FICH UNL | 2 | 50% | 0.2% | 3 |
| 4 | HIGHER SCI TECHNOL TUNIS | 2 | 43% | 0.2% | 3 |
| 5 | ONCOL PHYSIOL BIOPHYS | 1 | 38% | 0.2% | 3 |
| 6 | DEV INFORMAT SYST CIDISI | 1 | 50% | 0.1% | 2 |
| 7 | INVEST INGN SISTEMAS INFORMAC | 1 | 50% | 0.1% | 2 |
| 8 | KIT LANGUAGE TECHNOL DOCTORATE | 1 | 100% | 0.1% | 2 |
| 9 | NACL TECNOL AGR IB INTA | 1 | 100% | 0.1% | 2 |
| 10 | KNOWLEDGE DISCOVERY BIOINFORMAT GRP KDBIO | 1 | 40% | 0.1% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000224634 | NETWORK INFERENCE//GENE REGULATORY NETWORKS//GENE NETWORK INFERENCE |
| 2 | 0.0000175366 | MICROARRAY IMAGE PROCESSING//GENE FILTERING//MAQC |
| 3 | 0.0000170641 | MISSING VALUE IMPUTATION//MISSING VALUE ESTIMATION//COMPUTAT MATH BIOL |
| 4 | 0.0000157211 | GENE SET ANALYSIS//ENRICHMENT ANALYSIS//FUNCT GENOM NODE |
| 5 | 0.0000145598 | GENE SELECTION//FEATURE SELECTION//CANCER CLASSIFICATION |
| 6 | 0.0000141133 | FUZZY CLUSTERING//CLUSTER VALIDITY//SUBSPACE CLUSTERING |
| 7 | 0.0000090431 | TAISHO FUNCT GENOM//MULTIPHOTON DETECTION//ADAPTER TAGGED COMPETITIVE PCR |
| 8 | 0.0000074610 | CIS MOTIF//ARABIDOPSIS T87 CELL LINE//GENE ATLAS |
| 9 | 0.0000072090 | NONNEGATIVE MATRIX FACTORIZATION//NONNEGATIVE RANK//NON NEGATIVE MATRIX FACTORIZATION |
| 10 | 0.0000067009 | MODEL BASED CLUSTERING//MIXTURE MODELS//NONPARAMETRIC MIXTURE |