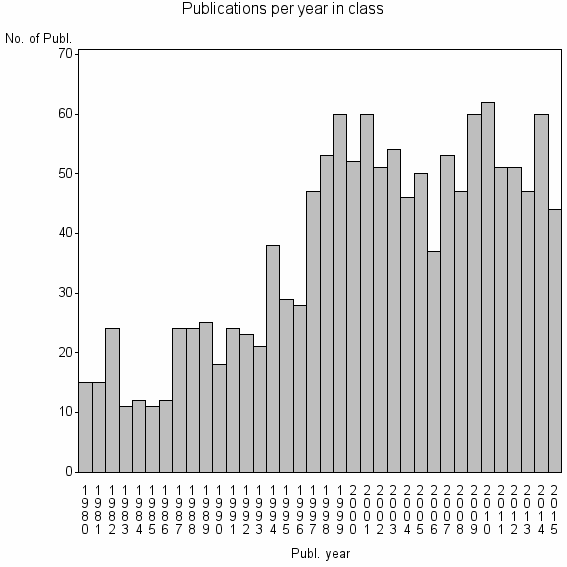
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 7022 | 1339 | 44.3 | 82% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 623 | 13179 | BASE EXCISION REPAIR//DNA GLYCOSYLASE//URACIL DNA GLYCOSYLASE |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | APE1 | Author keyword | 45 | 53% | 4% | 60 |
| 2 | AP ENDONUCLEASE | Author keyword | 39 | 50% | 4% | 56 |
| 3 | APE1 REF 1 | Author keyword | 37 | 81% | 2% | 22 |
| 4 | APURINIC APYRIMIDINIC ENDONUCLEASE | Author keyword | 27 | 83% | 1% | 15 |
| 5 | REF 1 | Author keyword | 26 | 56% | 2% | 31 |
| 6 | BASE EXCISION REPAIR | Author keyword | 21 | 16% | 9% | 123 |
| 7 | ENDONUCLEASE IV | Author keyword | 17 | 79% | 1% | 11 |
| 8 | APE REF 1 | Author keyword | 14 | 59% | 1% | 16 |
| 9 | PEDIATSECT HEMATOL ONCOL | Address | 13 | 67% | 1% | 12 |
| 10 | APEX NUCLEASE | Author keyword | 11 | 100% | 0% | 6 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | APE1 | 45 | 53% | 4% | 60 | Search APE1 | Search APE1 |
| 2 | AP ENDONUCLEASE | 39 | 50% | 4% | 56 | Search AP+ENDONUCLEASE | Search AP+ENDONUCLEASE |
| 3 | APE1 REF 1 | 37 | 81% | 2% | 22 | Search APE1+REF+1 | Search APE1+REF+1 |
| 4 | APURINIC APYRIMIDINIC ENDONUCLEASE | 27 | 83% | 1% | 15 | Search APURINIC+APYRIMIDINIC+ENDONUCLEASE | Search APURINIC+APYRIMIDINIC+ENDONUCLEASE |
| 5 | REF 1 | 26 | 56% | 2% | 31 | Search REF+1 | Search REF+1 |
| 6 | BASE EXCISION REPAIR | 21 | 16% | 9% | 123 | Search BASE+EXCISION+REPAIR | Search BASE+EXCISION+REPAIR |
| 7 | ENDONUCLEASE IV | 17 | 79% | 1% | 11 | Search ENDONUCLEASE+IV | Search ENDONUCLEASE+IV |
| 8 | APE REF 1 | 14 | 59% | 1% | 16 | Search APE+REF+1 | Search APE+REF+1 |
| 9 | APEX NUCLEASE | 11 | 100% | 0% | 6 | Search APEX+NUCLEASE | Search APEX+NUCLEASE |
| 10 | ABASIC SITES | 9 | 30% | 2% | 25 | Search ABASIC+SITES | Search ABASIC+SITES |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | APURINIC APYRIMIDINIC ENDONUCLEASE | 191 | 69% | 12% | 162 |
| 2 | HUMAN APURINIC ENDONUCLEASE | 154 | 73% | 9% | 116 |
| 3 | HUMAN APURINIC APYRIMIDINIC ENDONUCLEASE | 116 | 68% | 8% | 102 |
| 4 | REF 1 | 96 | 71% | 6% | 78 |
| 5 | APE1 REF 1 | 79 | 76% | 4% | 56 |
| 6 | COLI EXONUCLEASE III | 54 | 73% | 3% | 41 |
| 7 | NEGATIVE GENE REGULATION | 53 | 95% | 1% | 18 |
| 8 | DNA REPAIR ENZYME | 52 | 48% | 6% | 80 |
| 9 | EXONUCLEASE III | 50 | 36% | 8% | 111 |
| 10 | APYRIMIDINIC ENDONUCLEASE | 48 | 91% | 1% | 20 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Human Apurinic/Apyrimidinic Endonuclease 1 | 2014 | 17 | 224 | 75% |
| The Many Functions of APE1/Ref-1: Not Only a DNA Repair Enzyme | 2009 | 122 | 150 | 61% |
| Transcriptional Regulatory Functions of Mammalian AP-Endonuclease (APE1/Ref-1), an Essential Multifunctional Protein | 2009 | 77 | 156 | 64% |
| The intracellular localization of APE1/Ref-1: More than a passive phenomenon? | 2005 | 211 | 142 | 63% |
| Going APE over ref-1 | 2000 | 371 | 149 | 69% |
| Human AP endonuclease 1 (APE1): From mechanistic insights to druggable target in cancer | 2010 | 49 | 140 | 66% |
| Roles of base excision repair subpathways in correcting oxidized abasic sites in DNA | 2006 | 86 | 71 | 46% |
| Abasic sites in DNA: repair and biological consequences in Saccharomyces cerevisiae | 2004 | 217 | 76 | 42% |
| Small molecule inhibitors of DNA repair nuclease activities of APE1 | 2010 | 28 | 77 | 77% |
| The DNA base excision repair protein Ape1/Ref-1 as a therapeutic and chemopreventive target | 2007 | 104 | 109 | 57% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PEDIATSECT HEMATOL ONCOL | 13 | 67% | 0.9% | 12 |
| 2 | GRP REPARAT ADN | 5 | 31% | 1.0% | 14 |
| 3 | BIOCHEM BIOPHYS BIOL SCI | 4 | 75% | 0.2% | 3 |
| 4 | GRP REPARAT ADNUMR 8200 | 4 | 75% | 0.2% | 3 |
| 5 | EMERGING LIFE SCI TECHNOL | 3 | 100% | 0.2% | 3 |
| 6 | SECT HEMATOL ONCOLHERMAN B WELLS PEDIAT | 3 | 60% | 0.2% | 3 |
| 7 | DNA REPAIR UNIT | 2 | 32% | 0.4% | 6 |
| 8 | CANCEROL GUSTAVE ROUSSYGRP REPARAT ADN | 2 | 67% | 0.1% | 2 |
| 9 | CNRSUMR 8200 | 2 | 67% | 0.1% | 2 |
| 10 | BIOTECHNOL BIOMED LEIPZIG | 1 | 100% | 0.1% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000269192 | DNA GLYCOSYLASE//8 OXOGUANINE//BASE EXCISION REPAIR |
| 2 | 0.0000161885 | URACIL DNA GLYCOSYLASE//DUTPASE//DUTP PYROPHOSPHATASE |
| 3 | 0.0000146648 | ABASIC SITE//REGULATORY BIOORGAN CHEM//MISMATCH BINDING LIGAND |
| 4 | 0.0000119131 | DNA POLYMERASE//DNA POLYMERASE BETA//DNA POLYMERASE LAMBDA |
| 5 | 0.0000098323 | AOA2//ARSACS//APRATAXIN |
| 6 | 0.0000096339 | SPIROIMINODIHYDANTOIN//LES ACIDES NUCL//GUANIDINOHYDANTOIN |
| 7 | 0.0000083347 | FEN1//BRAUN S//CREAT INITIAT CELL CYCLE CONTROL |
| 8 | 0.0000074171 | XRCC1//XRCC3//XRCC1 GENE |
| 9 | 0.0000071245 | ADV AGING BRAIN//DNA GAP REPAIR//DNA DAMAGE SUSCEPTIBILITY |
| 10 | 0.0000067158 | ENDONUCLEASE V//S CAROLINA EXPT STN//OXANINE |