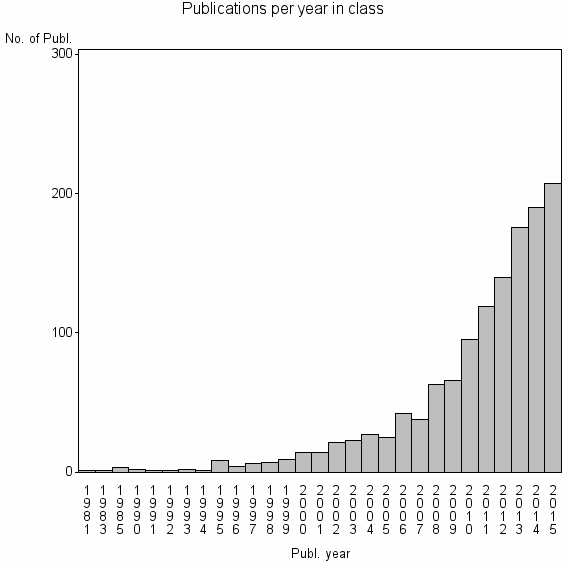
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 7303 | 1306 | 50.1 | 95% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | HIPPO PATHWAY | Author keyword | 148 | 75% | 8% | 106 |
| 2 | YAP | Author keyword | 121 | 59% | 10% | 135 |
| 3 | HIPPO | Author keyword | 100 | 74% | 6% | 75 |
| 4 | HIPPO SIGNALING | Author keyword | 83 | 84% | 4% | 46 |
| 5 | TAZ | Author keyword | 75 | 69% | 5% | 64 |
| 6 | YES ASSOCIATED PROTEIN | Author keyword | 61 | 82% | 3% | 36 |
| 7 | CELL GROWTH PROLIFERAT | Address | 57 | 95% | 1% | 19 |
| 8 | LATS1 | Author keyword | 32 | 78% | 2% | 21 |
| 9 | LATS2 | Author keyword | 30 | 68% | 2% | 26 |
| 10 | YORKIE | Author keyword | 25 | 77% | 1% | 17 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | HIPPO PATHWAY | 148 | 75% | 8% | 106 | Search HIPPO+PATHWAY | Search HIPPO+PATHWAY |
| 2 | YAP | 121 | 59% | 10% | 135 | Search YAP | Search YAP |
| 3 | HIPPO | 100 | 74% | 6% | 75 | Search HIPPO | Search HIPPO |
| 4 | HIPPO SIGNALING | 83 | 84% | 4% | 46 | Search HIPPO+SIGNALING | Search HIPPO+SIGNALING |
| 5 | TAZ | 75 | 69% | 5% | 64 | Search TAZ | Search TAZ |
| 6 | YES ASSOCIATED PROTEIN | 61 | 82% | 3% | 36 | Search YES+ASSOCIATED+PROTEIN | Search YES+ASSOCIATED+PROTEIN |
| 7 | LATS1 | 32 | 78% | 2% | 21 | Search LATS1 | Search LATS1 |
| 8 | LATS2 | 30 | 68% | 2% | 26 | Search LATS2 | Search LATS2 |
| 9 | YORKIE | 25 | 77% | 1% | 17 | Search YORKIE | Search YORKIE |
| 10 | MST1 | 24 | 67% | 2% | 22 | Search MST1 | Search MST1 |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ORGAN SIZE CONTROL | 335 | 78% | 17% | 222 |
| 2 | YES ASSOCIATED PROTEIN | 263 | 67% | 18% | 236 |
| 3 | HIPPO PATHWAY | 194 | 55% | 19% | 243 |
| 4 | TEAD TEF FAMILY | 152 | 78% | 8% | 101 |
| 5 | TUMOR SUPPRESSOR PATHWAY | 134 | 60% | 11% | 146 |
| 6 | HIPPO SIGNALING PATHWAY | 129 | 66% | 9% | 119 |
| 7 | YAP | 113 | 55% | 11% | 143 |
| 8 | PROMOTES APOPTOSIS | 101 | 38% | 16% | 213 |
| 9 | YAP ONCOPROTEIN | 90 | 88% | 3% | 43 |
| 10 | CELL CONTACT INHIBITION | 86 | 90% | 3% | 37 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| The Hippo pathway and human cancer | 2013 | 176 | 129 | 60% |
| The Hippo pathway: regulators and regulations | 2013 | 147 | 202 | 66% |
| The two faces of Hippo: targeting the Hippo pathway for regenerative medicine and cancer treatment | 2014 | 45 | 216 | 86% |
| The Hippo Signaling Pathway in Development and Cancer | 2010 | 459 | 172 | 74% |
| The Hippo pathway in organ size control, tissue regeneration and stem cell self-renewal | 2011 | 234 | 124 | 85% |
| The Hippo pathway effectors TAZ and YAP in development, homeostasis and disease | 2014 | 31 | 148 | 80% |
| The Hippo-YAP pathway in organ size control and tumorigenesis: an updated version | 2010 | 264 | 127 | 83% |
| The mammalian Hippo pathway: regulation and function of YAP1 and TAZ | 2015 | 2 | 239 | 87% |
| The YAP and TAZ transcription co-activators: Key downstream effectors of the mammalian Hippo pathway | 2012 | 84 | 80 | 81% |
| The Hippo signaling pathway in stem cell biology and cancer | 2014 | 25 | 152 | 79% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CELL GROWTH PROLIFERAT | 57 | 95% | 1.5% | 19 |
| 2 | OPTOSIS PROLIFERAT CONTROL | 8 | 46% | 1.0% | 13 |
| 3 | SIGNAL TRANSDUCT PROTE PROFILING | 8 | 62% | 0.6% | 8 |
| 4 | INTER DEGREE PROGRAM CELL DEV BIOL | 6 | 80% | 0.3% | 4 |
| 5 | TUMOUR SUPP SOR SIGNALLING NETWORKS | 6 | 80% | 0.3% | 4 |
| 6 | INT DEV DIS | 3 | 60% | 0.2% | 3 |
| 7 | EQUIPE GENET MOL DIFFERENCIAT | 2 | 67% | 0.2% | 2 |
| 8 | CHEM BIOL SCREENING | 2 | 50% | 0.2% | 3 |
| 9 | PREMED PROGRAMS | 2 | 50% | 0.2% | 3 |
| 10 | CANC DEV CELL BIOL | 2 | 13% | 0.9% | 12 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000199291 | STE20 LIKE KINASE//MAP4K4//STE20 |
| 2 | 0.0000135092 | PUR ALPHA//TEF 1//RTEF 1 |
| 3 | 0.0000090871 | DRONC//DIAP1//REAPER |
| 4 | 0.0000085797 | KIBRA//PKM ZETA//WWC1 |
| 5 | 0.0000085515 | IASPP//ASPP2//53BP2 |
| 6 | 0.0000082683 | RASSF1A//RASSF2//RASSF1 |
| 7 | 0.0000081708 | WWP1//SMURF1//CKIP 1 |
| 8 | 0.0000080146 | PLANAR CELL POLARITY//VANGL2//CONVERGENT EXTENSION |
| 9 | 0.0000073182 | CRUMBS//L2GL//ERBIN |
| 10 | 0.0000061496 | UNPAIRED//BAG OF MARBLES//GERMLINE STEM CELL |