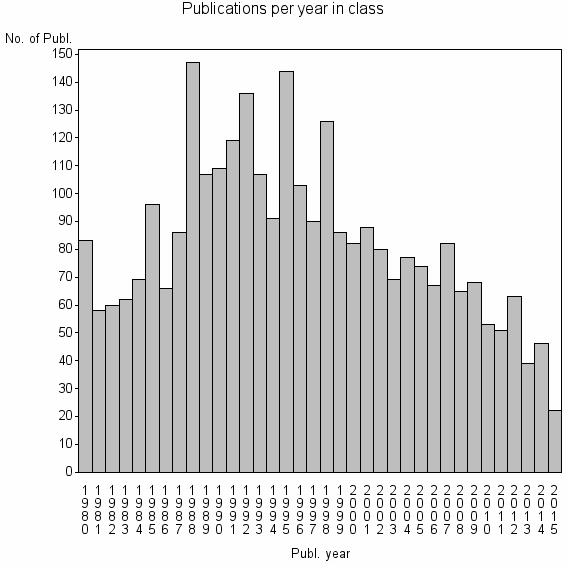
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 798 | 2971 | 34.7 | 67% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 1919 | 5291 | RESTRICTION ENDONUCLEASE//RESTRICTION MODIFICATION//RESTRICTION MODIFICATION SYSTEM |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | RESTRICTION MODIFICATION | Author keyword | 74 | 65% | 2% | 71 |
| 2 | RESTRICTION ENDONUCLEASE | Author keyword | 74 | 44% | 4% | 127 |
| 3 | RESTRICTION MODIFICATION SYSTEM | Author keyword | 48 | 63% | 2% | 49 |
| 4 | ISOSCHIZOMER | Author keyword | 35 | 74% | 1% | 26 |
| 5 | RESTRICTION MODIFICATION SYSTEMS | Author keyword | 33 | 75% | 1% | 24 |
| 6 | DNA PROT INTERACT UNIT | Address | 29 | 52% | 1% | 39 |
| 7 | ANTIRESTRICTION | Author keyword | 27 | 92% | 0% | 11 |
| 8 | SITE SPECIFIC ENDONUCLEASE | Author keyword | 23 | 70% | 1% | 19 |
| 9 | RESTRICTION ENZYME | Author keyword | 17 | 26% | 2% | 56 |
| 10 | RECOGNITION AND CLEAVAGE DOMAINS | Author keyword | 15 | 88% | 0% | 7 |
Web of Science journal categories |
Author Key Words |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MODIFICATION SYSTEM | 245 | 86% | 4% | 126 |
| 2 | MODIFICATION ENZYMES | 184 | 79% | 4% | 116 |
| 3 | MODIFICATION SYSTEMS | 145 | 66% | 5% | 135 |
| 4 | MODIFICATION METHYLASE | 112 | 89% | 2% | 50 |
| 5 | METHYLASE | 87 | 68% | 3% | 77 |
| 6 | TARGET BASE | 85 | 69% | 2% | 73 |
| 7 | REFRACTORY DNA | 76 | 93% | 1% | 28 |
| 8 | PVUII ENDONUCLEASE | 75 | 74% | 2% | 56 |
| 9 | RESTRICTION MODIFICATION SYSTEM | 73 | 52% | 3% | 99 |
| 10 | I RESTRICTION | 69 | 81% | 1% | 42 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Structure and function of type II restriction endonucleases | 2001 | 334 | 175 | 77% |
| Type II restriction endonucleases: structure and mechanism | 2005 | 249 | 191 | 74% |
| BIOLOGY OF DNA RESTRICTION | 1993 | 400 | 172 | 85% |
| RESTRICTION AND MODIFICATION SYSTEMS | 1991 | 446 | 167 | 79% |
| Diverse Functions of Restriction-Modification Systems in Addition to Cellular Defense | 2013 | 28 | 213 | 48% |
| Type I restriction systems: Sophisticated molecular machines (a legacy of Bertani and Weigle) | 2000 | 203 | 150 | 85% |
| Structural and evolutionary classification of Type II restriction enzymes based on theoretical and experimental analyses | 2008 | 50 | 110 | 73% |
| On the Divalent Metal Ion Dependence of DNA Cleavage by Restriction Endonucleases of the EcoRI Family | 2009 | 36 | 82 | 80% |
| The biology of restriction and anti-restriction | 2005 | 127 | 44 | 80% |
| Behavior of restriction-modification systems as selfish mobile elements and their impact on genome evolution | 2001 | 232 | 108 | 56% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | DNA PROT INTERACT UNIT | 29 | 52% | 1.3% | 39 |
| 2 | BIOINFORMAT PROT ENGN | 7 | 20% | 1.1% | 32 |
| 3 | ABT BIOPHYS BIOCHEM VERFAHREN | 6 | 80% | 0.1% | 4 |
| 4 | PROGRAM BIOPHYS BIOCHEM | 5 | 27% | 0.6% | 17 |
| 5 | BIOPHYS S | 5 | 25% | 0.6% | 18 |
| 6 | IBBS BIOPHYS S | 5 | 63% | 0.2% | 5 |
| 7 | BIOL DNA MODIFICAT | 4 | 33% | 0.3% | 10 |
| 8 | INTER PROGRAM BIOCHEM MOL BIOL | 4 | 41% | 0.2% | 7 |
| 9 | SOCIAL GENOME SCI | 3 | 57% | 0.1% | 4 |
| 10 | ABT STRUKTURANAL | 3 | 100% | 0.1% | 3 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000083606 | AIST TSUKUBA 6 10//LEUCINE RESPONSIVE REGULATORY PROTEIN//ASNC |
| 2 | 0.0000072753 | PHOSPHOROTHIOLATE//P 32ORTHOPHOSPHORIC ACID//SYNTHESIS OF P 32 LABELED NUCLEOTIDES |
| 3 | 0.0000057296 | POLYLINKER//STREPTOMYCIN SENSITIVITY//INSERTIONAL INACTIVATION |
| 4 | 0.0000051927 | NARROW ESCAPE//DEN DANS DM2S SERMA LTSD//NANOMED MECH BIOL |
| 5 | 0.0000049342 | PROTEIN SPLICING//INTEIN//HOMING ENDONUCLEASE |
| 6 | 0.0000044298 | MEGAPRIMER//OVERLAP EXTENSION//MULTIPLE SITE DIRECTED MUTAGENESIS |
| 7 | 0.0000042809 | DNA 5 METHYLCYTOSINE//DNA METHYLASE//5 METHYLDCMP DEAMINASE |
| 8 | 0.0000039907 | DYE TERMINATOR//ENERGY TRANSFER DYES//GENOMAT |
| 9 | 0.0000039567 | RETARDATION ANALYSIS//NONSPECIFIC PROTEIN DNA COMPLEXES//GEL CAGE |
| 10 | 0.0000038874 | T7 RNA POLYMERASE//T7 LYSOZYME//PHAGE RNA POLYMERASE |