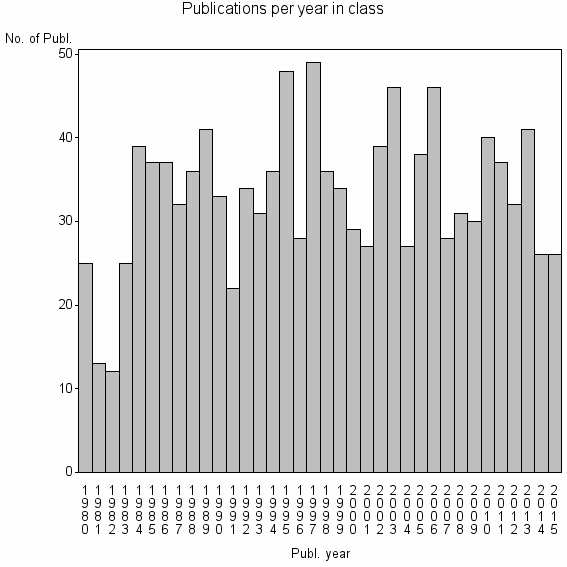
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 8371 | 1191 | 37.6 | 78% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 501 | 14604 | RNA POLYMERASE//SITE SPECIFIC RECOMBINATION//H NS |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | SITE SPECIFIC RECOMBINATION | Author keyword | 72 | 36% | 14% | 163 |
| 2 | SERINE RECOMBINASE | Author keyword | 47 | 88% | 2% | 22 |
| 3 | PHI C31 INTEGRASE | Author keyword | 26 | 54% | 3% | 34 |
| 4 | TYROSINE RECOMBINASE | Author keyword | 15 | 53% | 2% | 20 |
| 5 | LAMBDA INTEGRASE | Author keyword | 15 | 71% | 1% | 12 |
| 6 | XER | Author keyword | 15 | 71% | 1% | 12 |
| 7 | SITE SPECIFIC DNA RECOMBINATION | Author keyword | 15 | 73% | 1% | 11 |
| 8 | PHAGE INTEGRASE | Author keyword | 13 | 80% | 1% | 8 |
| 9 | BACTERIOPHAGE HK022 | Author keyword | 10 | 73% | 1% | 8 |
| 10 | RESOLVASE | Author keyword | 8 | 35% | 2% | 19 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | SITE SPECIFIC RECOMBINATION | 72 | 36% | 14% | 163 | Search SITE+SPECIFIC+RECOMBINATION | Search SITE+SPECIFIC+RECOMBINATION |
| 2 | SERINE RECOMBINASE | 47 | 88% | 2% | 22 | Search SERINE+RECOMBINASE | Search SERINE+RECOMBINASE |
| 3 | PHI C31 INTEGRASE | 26 | 54% | 3% | 34 | Search PHI+C31+INTEGRASE | Search PHI+C31+INTEGRASE |
| 4 | TYROSINE RECOMBINASE | 15 | 53% | 2% | 20 | Search TYROSINE+RECOMBINASE | Search TYROSINE+RECOMBINASE |
| 5 | LAMBDA INTEGRASE | 15 | 71% | 1% | 12 | Search LAMBDA+INTEGRASE | Search LAMBDA+INTEGRASE |
| 6 | XER | 15 | 71% | 1% | 12 | Search XER | Search XER |
| 7 | SITE SPECIFIC DNA RECOMBINATION | 15 | 73% | 1% | 11 | Search SITE+SPECIFIC+DNA+RECOMBINATION | Search SITE+SPECIFIC+DNA+RECOMBINATION |
| 8 | PHAGE INTEGRASE | 13 | 80% | 1% | 8 | Search PHAGE+INTEGRASE | Search PHAGE+INTEGRASE |
| 9 | BACTERIOPHAGE HK022 | 10 | 73% | 1% | 8 | Search BACTERIOPHAGE+HK022 | Search BACTERIOPHAGE+HK022 |
| 10 | RESOLVASE | 8 | 35% | 2% | 19 | Search RESOLVASE | Search RESOLVASE |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | GAMMA DELTA RESOLVASE | 133 | 75% | 8% | 95 |
| 2 | TN3 RESOLVASE | 117 | 72% | 8% | 92 |
| 3 | STEP ARREST MUTANTS | 93 | 100% | 2% | 28 |
| 4 | EXCISIVE RECOMBINATION | 64 | 100% | 2% | 21 |
| 5 | PHI C31 INTEGRASE | 61 | 55% | 6% | 76 |
| 6 | INTEGRASE FAMILY | 60 | 60% | 6% | 66 |
| 7 | LAMBDA INTEGRASE | 59 | 63% | 5% | 60 |
| 8 | SITE SPECIFIC RECOMBINATION | 54 | 14% | 29% | 347 |
| 9 | BXB1 INTEGRATION | 52 | 100% | 2% | 18 |
| 10 | GENOMIC INTEGRATION | 49 | 57% | 5% | 59 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Mechanisms of site-specific recombination | 2006 | 259 | 136 | 78% |
| Phage integrases: Biology and applications | 2004 | 186 | 88 | 86% |
| Large serine recombinase domain structure and attachment site binding | 2013 | 6 | 65 | 78% |
| A structural view of Cre-loxP site-specific recombination | 2001 | 128 | 59 | 85% |
| Serine recombinases as tools for genome engineering | 2011 | 18 | 49 | 86% |
| Diversity in the serine recombinases | 2002 | 163 | 54 | 63% |
| Challenging a Paradigm: the Role of DNA Homology in Tyrosine Recombinase Reactions | 2009 | 24 | 79 | 72% |
| Therapeutic Applications of the PhiC31 Integrase System | 2011 | 18 | 43 | 79% |
| DYNAMIC, STRUCTURAL, AND REGULATORY ASPECTS OF LAMBDA-SITE-SPECIFIC RECOMBINATION | 1989 | 440 | 137 | 66% |
| Similarities and differences among 105 members of the Int family of site-specific recombinases | 1998 | 310 | 149 | 36% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BIOCHEM MICROBIOL UNIT | 3 | 57% | 0.3% | 4 |
| 2 | SHANGHAI EMBRYO REPROD ENGN | 2 | 67% | 0.2% | 2 |
| 3 | MED EMBRYO MOL BIOL | 1 | 33% | 0.2% | 2 |
| 4 | BIOL MOL CELL BIOL BIOCHEM | 1 | 50% | 0.1% | 1 |
| 5 | CHEM LIFE SCI B 103 | 1 | 50% | 0.1% | 1 |
| 6 | GRP BIOINFORMAT ECO SYST BIOL | 1 | 50% | 0.1% | 1 |
| 7 | MOL EVOLUTIONARY GENET MUELLER | 1 | 50% | 0.1% | 1 |
| 8 | NEUROL SERV GERIATR | 1 | 50% | 0.1% | 1 |
| 9 | PITTSBURGH BACTERIOL | 1 | 50% | 0.1% | 1 |
| 10 | SHANGHAI MOST HLTH DIS GEN | 1 | 25% | 0.2% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000214733 | CRE RECOMBINASE//LOXP//ROSA26 |
| 2 | 0.0000130714 | DNA TRANSPOSITION//PHAGE MU//TN10 |
| 3 | 0.0000130312 | SKIRBALL PROGRAM MOL PATHOGENESIS//BACTERIOPHAGE P2//RETRON |
| 4 | 0.0000107910 | TN916//CONJUGATIVE TRANSPOSON//CONJUGATIVE TRANSPOSONS |
| 5 | 0.0000092553 | PARB//DNA SEGREGATION//FTSK |
| 6 | 0.0000092286 | H NS//HISTONE LIKE PROTEIN//BACTERIAL CHROMATIN |
| 7 | 0.0000090094 | BLOOMINGTON DROSOPHILA STOCK//ENDS OUT TARGETING//ENDS OUT |
| 8 | 0.0000081272 | MARKER FREE TRANSGENIC PLANTS//MARKER FREE//MANNOSE SELECTION |
| 9 | 0.0000081143 | REX EXCLUSION//BACTERIOPHAGE LAMBDA LAMBDA//CI REXA REXB OPERON |
| 10 | 0.0000069372 | MYCOBACTERIOPHAGE L1//RECOMBINANT BCG VACCINE//RECOMBINANT BCG |