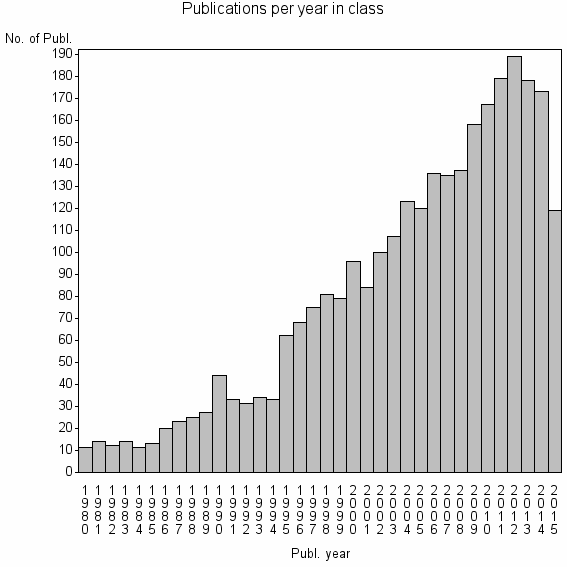
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 869 | 2911 | 52.0 | 83% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | HEART DEVELOPMENT | Author keyword | 193 | 44% | 11% | 334 |
| 2 | SECOND HEART FIELD | Author keyword | 148 | 90% | 2% | 64 |
| 3 | NKX2 5 | Author keyword | 73 | 71% | 2% | 60 |
| 4 | EPICARDIUM | Author keyword | 72 | 41% | 5% | 135 |
| 5 | PROEPICARDIUM | Author keyword | 65 | 93% | 1% | 25 |
| 6 | SECONDARY HEART FIELD | Author keyword | 64 | 86% | 1% | 32 |
| 7 | CARDIOGENESIS | Author keyword | 60 | 39% | 4% | 122 |
| 8 | CARDIAC DEVELOPMENT | Author keyword | 57 | 30% | 5% | 158 |
| 9 | OUTFLOW TRACT | Author keyword | 52 | 50% | 3% | 76 |
| 10 | NKX25 | Author keyword | 50 | 57% | 2% | 60 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | HEART DEVELOPMENT | 193 | 44% | 11% | 334 | Search HEART+DEVELOPMENT | Search HEART+DEVELOPMENT |
| 2 | SECOND HEART FIELD | 148 | 90% | 2% | 64 | Search SECOND+HEART+FIELD | Search SECOND+HEART+FIELD |
| 3 | NKX2 5 | 73 | 71% | 2% | 60 | Search NKX2+5 | Search NKX2+5 |
| 4 | EPICARDIUM | 72 | 41% | 5% | 135 | Search EPICARDIUM | Search EPICARDIUM |
| 5 | PROEPICARDIUM | 65 | 93% | 1% | 25 | Search PROEPICARDIUM | Search PROEPICARDIUM |
| 6 | SECONDARY HEART FIELD | 64 | 86% | 1% | 32 | Search SECONDARY+HEART+FIELD | Search SECONDARY+HEART+FIELD |
| 7 | CARDIOGENESIS | 60 | 39% | 4% | 122 | Search CARDIOGENESIS | Search CARDIOGENESIS |
| 8 | CARDIAC DEVELOPMENT | 57 | 30% | 5% | 158 | Search CARDIAC+DEVELOPMENT | Search CARDIAC+DEVELOPMENT |
| 9 | OUTFLOW TRACT | 52 | 50% | 3% | 76 | Search OUTFLOW+TRACT | Search OUTFLOW+TRACT |
| 10 | NKX25 | 50 | 57% | 2% | 60 | Search NKX25 | Search NKX25 |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ARTERIAL POLE | 148 | 76% | 4% | 104 |
| 2 | SUBEPICARDIAL MESENCHYME | 132 | 89% | 2% | 59 |
| 3 | AVIAN HEART | 125 | 74% | 3% | 92 |
| 4 | HOLT ORAM SYNDROME | 120 | 40% | 8% | 234 |
| 5 | OUTFLOW TRACT | 93 | 30% | 9% | 257 |
| 6 | CHAMBER FORMATION | 87 | 76% | 2% | 62 |
| 7 | CARDIAC MORPHOGENESIS | 87 | 55% | 4% | 108 |
| 8 | CARDIAC OUTFLOW TRACT | 77 | 60% | 3% | 84 |
| 9 | EMBRYONIC HEART | 72 | 42% | 5% | 131 |
| 10 | ANTERIOR ENDODERM | 63 | 64% | 2% | 61 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Cellular Origin and Developmental Program of Coronary Angiogenesis | 2015 | 4 | 124 | 69% |
| Molecular Regulation of Cardiomyocyte Differentiation | 2015 | 3 | 156 | 70% |
| Building the mammalian heart from two sources of myocardial cells | 2005 | 473 | 76 | 82% |
| How TO MAKE A HEART: THE ORIGIN AND REGULATION OF CARDIAC PROGENITOR CELLS | 2010 | 126 | 171 | 82% |
| Gene regulatory networks in the evolution and development of the heart | 2006 | 411 | 49 | 51% |
| Signaling Pathways Controlling Second Heart Field Development | 2009 | 92 | 104 | 84% |
| Zebrafish as a model to study cardiac development and human cardiac disease | 2011 | 96 | 84 | 55% |
| Genetics of Congenital Heart Disease The Glass Half Empty | 2013 | 66 | 151 | 31% |
| Mechanisms of T-box gene function in the developing heart | 2011 | 67 | 87 | 71% |
| Myocardial Lineage Development | 2010 | 69 | 206 | 80% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CARDIOVASC DEV BIOL | 15 | 34% | 1.2% | 36 |
| 2 | ANAT EMBRYOL PHYSIOL | 15 | 30% | 1.4% | 41 |
| 3 | PUBL CENT | 7 | 57% | 0.3% | 8 |
| 4 | CARDIOVASC DEV | 7 | 23% | 0.9% | 26 |
| 5 | NEONATOLNEONATAL PERINATAL | 6 | 80% | 0.1% | 4 |
| 6 | DEV BIOL MARSEILLES LUMINY | 6 | 58% | 0.2% | 7 |
| 7 | KIMMEL BIOL MEDSKIRBALL BIOMOL MED | 6 | 53% | 0.3% | 8 |
| 8 | EXPT MOL CARDIOL GRP | 6 | 19% | 1.0% | 28 |
| 9 | NEONATAL PERINATAL | 5 | 31% | 0.5% | 15 |
| 10 | CARDIOVASC MORPHOGENESIS | 5 | 54% | 0.2% | 7 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000208006 | HOLT ORAM SYNDROME//TBX3//TBX2 |
| 2 | 0.0000187750 | CARDIAC LOOPING//LEFT ATRIAL LIGATION//EMBRYONIC CIRCULATION |
| 3 | 0.0000122752 | CARDIAC DIFFERENTIATION//STEM CELL REGENERAT MED CONSORTIUM//SOHNIS FAMILY CARDIAC ELE OPHYSIOL RE |
| 4 | 0.0000098327 | BVES//STAHLMAN CARDIOVASC S//LEK1 |
| 5 | 0.0000093612 | NODOSE PLACODE//MYOCARDIAL LAYERS//EXTRA LOAD |
| 6 | 0.0000086462 | GATA 1//GATA 2//EDAG |
| 7 | 0.0000085478 | LONE AF//4Q25//LONE ATRIAL FIBRILLATION |
| 8 | 0.0000084325 | LEFT RIGHT ASYMMETRY//LEFT RIGHT PATTERNING//NODAL FLOW |
| 9 | 0.0000063577 | CITED2//CITED//MELANOCYTE SPECIFIC GENE MSG1 |
| 10 | 0.0000062732 | ARID3B//JUMONJI//AT RICH INTERACTION DOMAIN |