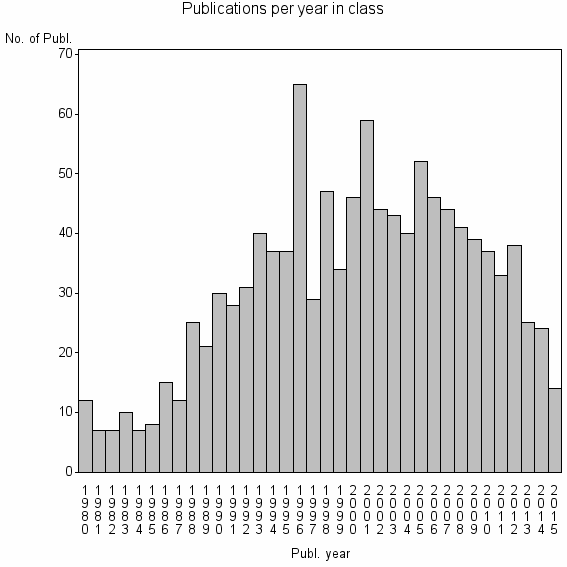
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 9027 | 1127 | 39.2 | 83% |
Classes in level above (level 2) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | RNASE P | Author keyword | 310 | 79% | 18% | 200 |
| 2 | RIBONUCLEASE P | Author keyword | 161 | 93% | 6% | 62 |
| 3 | CARTILAGE HAIR HYPOPLASIA | Author keyword | 100 | 90% | 4% | 44 |
| 4 | RNASE MRP | Author keyword | 71 | 84% | 3% | 38 |
| 5 | TRNA PROCESSING | Author keyword | 54 | 59% | 5% | 60 |
| 6 | TRNASE Z | Author keyword | 53 | 95% | 2% | 18 |
| 7 | C5 PROTEIN | Author keyword | 52 | 100% | 2% | 18 |
| 8 | PROGRAM COMPARAT BIOCHEM | Address | 49 | 78% | 3% | 32 |
| 9 | TRNA PRECURSORS | Author keyword | 32 | 85% | 2% | 17 |
| 10 | M1 RNA | Author keyword | 31 | 74% | 2% | 23 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | RNASE P | 310 | 79% | 18% | 200 | Search RNASE+P | Search RNASE+P |
| 2 | RIBONUCLEASE P | 161 | 93% | 6% | 62 | Search RIBONUCLEASE+P | Search RIBONUCLEASE+P |
| 3 | CARTILAGE HAIR HYPOPLASIA | 100 | 90% | 4% | 44 | Search CARTILAGE+HAIR+HYPOPLASIA | Search CARTILAGE+HAIR+HYPOPLASIA |
| 4 | RNASE MRP | 71 | 84% | 3% | 38 | Search RNASE+MRP | Search RNASE+MRP |
| 5 | TRNA PROCESSING | 54 | 59% | 5% | 60 | Search TRNA+PROCESSING | Search TRNA+PROCESSING |
| 6 | TRNASE Z | 53 | 95% | 2% | 18 | Search TRNASE+Z | Search TRNASE+Z |
| 7 | C5 PROTEIN | 52 | 100% | 2% | 18 | Search C5+PROTEIN | Search C5+PROTEIN |
| 8 | TRNA PRECURSORS | 32 | 85% | 2% | 17 | Search TRNA+PRECURSORS | Search TRNA+PRECURSORS |
| 9 | M1 RNA | 31 | 74% | 2% | 23 | Search M1+RNA | Search M1+RNA |
| 10 | EXTERNAL GUIDE SEQUENCE | 31 | 92% | 1% | 12 | Search EXTERNAL+GUIDE+SEQUENCE | Search EXTERNAL+GUIDE+SEQUENCE |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | M1 RNA | 316 | 90% | 12% | 139 |
| 2 | RIBONUCLEASE P | 259 | 55% | 28% | 321 |
| 3 | PRECURSOR TRANSFER RNA | 144 | 93% | 5% | 54 |
| 4 | 45 S RNA | 103 | 97% | 3% | 30 |
| 5 | HUMAN RIBONUCLEASE P | 93 | 88% | 4% | 44 |
| 6 | 3 PROCESSING ENDORIBONUCLEASE | 73 | 89% | 3% | 33 |
| 7 | RIBONUCLEOPROTEIN ENZYME | 72 | 83% | 4% | 40 |
| 8 | PRE TRANSFER RNA | 58 | 72% | 4% | 46 |
| 9 | BACTERIAL RIBONUCLEASE P | 44 | 63% | 4% | 45 |
| 10 | C5 PROTEIN | 37 | 100% | 1% | 14 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Of proteins and RNA: The RNase P/MRP family | 2010 | 50 | 253 | 89% |
| Structural Studies of RNase P | 2013 | 11 | 122 | 82% |
| RNase P: interface of the RNA and protein worlds | 2006 | 90 | 66 | 94% |
| Ribonuclease P: The evolution of an ancient RNA enzyme | 2006 | 103 | 192 | 80% |
| Ribonuclease P: Unity and diversity in a tRNA processing ribozyme | 1998 | 303 | 134 | 78% |
| The Making of tRNAs and More - RNase P and tRNase Z | 2009 | 51 | 192 | 67% |
| Bacterial RNase P: a new view of an ancient enzyme | 2006 | 81 | 117 | 84% |
| Unexpected diversity of RNase P, an ancient tRNA processing enzyme: Challenges and prospects | 2010 | 33 | 135 | 83% |
| Of P and Z: Mitochondrial tRNA processing enzymes | 2012 | 20 | 124 | 54% |
| Eukaryotic ribonuclease P: A plurality of ribonucleoprotein enzymes | 2002 | 88 | 157 | 79% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PROGRAM COMPARAT BIOCHEM | 49 | 78% | 2.8% | 32 |
| 2 | PROGRAM INFECT DIS IMMUN | 9 | 35% | 1.8% | 20 |
| 3 | BIOSCI BIOTECHNOL BIOCHEM | 4 | 42% | 0.7% | 8 |
| 4 | JIANGSU MICROBES GENOM | 4 | 75% | 0.3% | 3 |
| 5 | BIOCHEM BIOSCI BIOTECHNOL | 2 | 43% | 0.3% | 3 |
| 6 | SWEGENE BIOINFORMAT | 1 | 40% | 0.2% | 2 |
| 7 | BIOCHEM BIOSCI BIOTECHNOLHIGASHI KU | 1 | 33% | 0.2% | 2 |
| 8 | ARQUEOL RNA | 1 | 50% | 0.1% | 1 |
| 9 | BIOL PHYSIOCHIM | 1 | 50% | 0.1% | 1 |
| 10 | BIOSYNTHESE NEURALER STRUKTUREN | 1 | 50% | 0.1% | 1 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000207722 | TRNA NUCLEOTIDYLTRANSFERASE//CCA ADDING ENZYME//TRNAHIS GUANYLYLTRANSFERASE |
| 2 | 0.0000107878 | GROUP I INTRON//SELF SPLICING//RIBOZYME |
| 3 | 0.0000107140 | RNASE E//RNASE G//POLYNUCLEOTIDE PHOSPHORYLASE |
| 4 | 0.0000094999 | HAMMERHEAD RIBOZYME//HAIRPIN RIBOZYME//RIBOZYME |
| 5 | 0.0000080209 | SCHIMKE IMMUNO OSSEOUS DYSPLASIA//SMARCAL1//SCHIMKE IMMUNOOSSEOUS DYSPLASIA |
| 6 | 0.0000076948 | TRNA SPLICING//POLYNUCLEOTIDE KINASE//SPLICING ENDONUCLEASE |
| 7 | 0.0000064650 | SNORNA//SMALL NUCLEOLAR RNAS//PRE RRNA PROCESSING |
| 8 | 0.0000053719 | MITO PORTER//MITOCHONDRIAL DRUG DELIVERY//MITOCHONDRIAL GENE THERAPY |
| 9 | 0.0000052882 | RNA POLYMERASE III//TFIIIB//SNAPC |
| 10 | 0.0000044880 | RNA SECONDARY STRUCTURE//RNA SECONDARY STRUCTURE PREDICTION//RNA STRUCTURE PREDICTION |