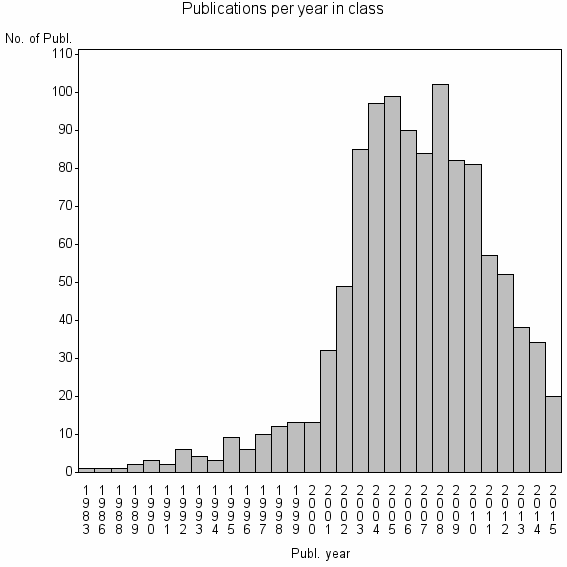
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 9423 | 1088 | 33.5 | 81% |
Classes in level above (level 2) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 2228 | 4315 | SEQUENCING BY HYBRIDIZATION//MUTATION DETECTION//TILLING |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | NONSPECIFIC HYBRIDIZATIONS | Author keyword | 6 | 100% | 0% | 4 |
| 2 | NEAREST NEIGHBOR PROPERTIES | Author keyword | 4 | 75% | 0% | 3 |
| 3 | NUCLEIC ACID OLIGOMERS | Author keyword | 3 | 100% | 0% | 3 |
| 4 | SIMULTANEOUS MULTIPLE DETECTION | Author keyword | 3 | 100% | 0% | 3 |
| 5 | ISIMA LIMOS | Address | 2 | 67% | 0% | 2 |
| 6 | PRIMER DESIGN | Author keyword | 2 | 13% | 1% | 16 |
| 7 | CHEM SENSORS GRP | Address | 2 | 14% | 1% | 13 |
| 8 | DIGITAL CONTENT DESIGN MANAGEMENT | Address | 2 | 36% | 0% | 4 |
| 9 | PROBE SELECTION | Author keyword | 2 | 36% | 0% | 4 |
| 10 | PROBE DESIGN | Author keyword | 2 | 16% | 1% | 10 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | NONSPECIFIC HYBRIDIZATIONS | 6 | 100% | 0% | 4 | Search NONSPECIFIC+HYBRIDIZATIONS | Search NONSPECIFIC+HYBRIDIZATIONS |
| 2 | NEAREST NEIGHBOR PROPERTIES | 4 | 75% | 0% | 3 | Search NEAREST+NEIGHBOR+PROPERTIES | Search NEAREST+NEIGHBOR+PROPERTIES |
| 3 | NUCLEIC ACID OLIGOMERS | 3 | 100% | 0% | 3 | Search NUCLEIC+ACID+OLIGOMERS | Search NUCLEIC+ACID+OLIGOMERS |
| 4 | SIMULTANEOUS MULTIPLE DETECTION | 3 | 100% | 0% | 3 | Search SIMULTANEOUS+MULTIPLE+DETECTION | Search SIMULTANEOUS+MULTIPLE+DETECTION |
| 5 | PRIMER DESIGN | 2 | 13% | 1% | 16 | Search PRIMER+DESIGN | Search PRIMER+DESIGN |
| 6 | PROBE SELECTION | 2 | 36% | 0% | 4 | Search PROBE+SELECTION | Search PROBE+SELECTION |
| 7 | PROBE DESIGN | 2 | 16% | 1% | 10 | Search PROBE+DESIGN | Search PROBE+DESIGN |
| 8 | MICROBIAL DIAGNOSTICS | 2 | 43% | 0% | 3 | Search MICROBIAL+DIAGNOSTICS | Search MICROBIAL+DIAGNOSTICS |
| 9 | CROSS HYBRIDIZATION | 1 | 18% | 1% | 7 | Search CROSS+HYBRIDIZATION | Search CROSS+HYBRIDIZATION |
| 10 | FUNCTIONAL GENE ARRAY | 1 | 38% | 0% | 3 | Search FUNCTIONAL+GENE+ARRAY | Search FUNCTIONAL+GENE+ARRAY |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PROBE DESIGN CRITERIA | 32 | 88% | 1% | 15 |
| 2 | SELECTING SIGNATURE OLIGONUCLEOTIDES | 24 | 91% | 1% | 10 |
| 3 | OLIGONUCLEOTIDE MICROARRAYS | 23 | 23% | 8% | 89 |
| 4 | HYBRIDIZATION ISOTHERMS | 15 | 71% | 1% | 12 |
| 5 | NONEQUILIBRIUM DISSOCIATION APPROACH | 14 | 100% | 1% | 7 |
| 6 | NEAREST NEIGHBOR THERMODYNAMICS | 13 | 23% | 5% | 50 |
| 7 | OPTIMAL DNA OLIGOS | 13 | 80% | 1% | 8 |
| 8 | HOOK CALIBRATION | 10 | 73% | 1% | 8 |
| 9 | PRIMER DESIGN | 9 | 39% | 2% | 18 |
| 10 | SEQUENCE DEPENDENT PROPERTIES | 8 | 75% | 1% | 6 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Oligonucleotide microarrays in microbial diagnostics | 2004 | 150 | 59 | 47% |
| The thermodynamics of DNA structural motifs | 2004 | 448 | 88 | 18% |
| Physicochemical perspectives on DNA microarray and biosensor technologies | 2005 | 133 | 54 | 43% |
| Microarrays for bacterial detection and microbial community analysis | 2003 | 187 | 30 | 47% |
| Microarray applications in microbial ecology research | 2006 | 88 | 113 | 41% |
| On the hybridization isotherms of DNA microarrays: the Langmuir model and its extensions | 2006 | 71 | 59 | 47% |
| Comparing microarrays and next-generation sequencing technologies for microbial ecology research | 2010 | 47 | 89 | 27% |
| Highly parallel microbial diagnostics using oligonucleotide microarrays | 2006 | 76 | 118 | 42% |
| Challenges and opportunities for pathogen detection using DNA microarrays | 2005 | 94 | 59 | 41% |
| Development of functional gene microarrays for microbial community analysis | 2012 | 6 | 48 | 63% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ISIMA LIMOS | 2 | 67% | 0.2% | 2 |
| 2 | CHEM SENSORS GRP | 2 | 14% | 1.2% | 13 |
| 3 | DIGITAL CONTENT DESIGN MANAGEMENT | 2 | 36% | 0.4% | 4 |
| 4 | PHD MICROBIOL PROGRAM | 1 | 100% | 0.2% | 2 |
| 5 | PL GENE SENSOR | 1 | 100% | 0.2% | 2 |
| 6 | PL GENSENSOR TECHNOL | 1 | 100% | 0.2% | 2 |
| 7 | PROGRAM BIOMATH BIOSTAT | 1 | 50% | 0.2% | 2 |
| 8 | TMAS2 | 1 | 100% | 0.2% | 2 |
| 9 | TRANSPORT MODELING BIOANALYT SEPARAT SCI GRP | 1 | 100% | 0.2% | 2 |
| 10 | WWTF CHAIR BIOINFORMAT | 1 | 50% | 0.2% | 2 |
Related classes at same level (level 1) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000249467 | SEQUENCING BY HYBRIDIZATION//DNA SEQUENCING WITH ERRORS//GAPPED PROBES |
| 2 | 0.0000155996 | EPIDEMIOL PATHOPHYSIOL ONCOGEN VIRUS UNIT//LYSSAVIRUS DYNAM HOST AD TAT UNIT//VIRAL DISCOVERY |
| 3 | 0.0000123076 | MICROARRAY IMAGE PROCESSING//GENE FILTERING//MAQC |
| 4 | 0.0000101440 | POLYNUCLEOTIDE PROBES//SOCIAL ENVIRONM SYST ENGN//CATALYZED REPORTER DEPOSITION |
| 5 | 0.0000096538 | HIGH RESOLUTION MELTING//HIGH RESOLUTION MELTING ANALYSIS HRM//HRMA |
| 6 | 0.0000075015 | ACTIVATED BIOMASS//ENVIRONM MODELING GENOM//ENVIRONM GENOM UNIT |
| 7 | 0.0000069966 | SURFACE MEDIATED POLYMERIZATION//VISIBLE LIGHT PHOTOPOLYMERIZATION//ACCESSIBILITY PREDICTION |
| 8 | 0.0000067332 | DNA DYNAMICS//PEYRARD BISHOP MODEL//DNA DENATURATION |
| 9 | 0.0000056127 | DNA POOLING//DNA POOL//PYROSEQUENCING TECHNOLOGY |
| 10 | 0.0000052273 | DNA COMPUTING//SPLICING SYSTEMS//ADLEMAN LIPTON MODEL |