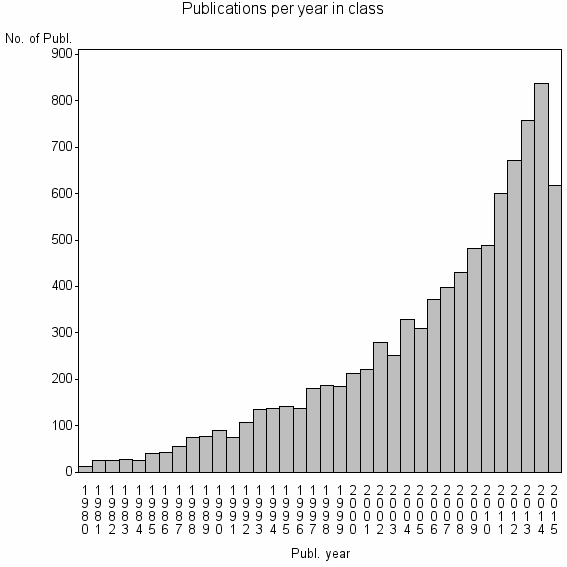
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 1161 | 9031 | 49.3 | 84% |
Classes in level above (level 3) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 4 | 177718 | PLANT SCIENCES//PLANT PHYSIOLOGY//PLANT CELL REPORTS |
Classes in level below (level 1) |
| ID, lev. below | Publications | Label for level below |
|---|---|---|
| 2330 | 2212 | DREB//AP2 ERF//GCC BOX |
| 6495 | 1409 | DEHYDRIN//LEA PROTEIN//DEHYDRINS |
| 6829 | 1362 | HEAT STRESS TRANSCRIPTION FACTOR//THERMOTOLERANCE//HEAT SHOCK TRANSCRIPTION FACTOR |
| 7735 | 1256 | PLANT PROTEOMICS//INPPO//LEAF PROTEOME |
| 8570 | 1172 | SNRK2//ABI3//ABA RECEPTOR |
| 13512 | 768 | MAPKK//MAPKKK//MAPK CASCADE |
| 19957 | 419 | WRKY TRANSCRIPTION FACTOR//WRKY//WRKY PROTEIN |
| 25566 | 230 | U BOX//CELLULAR BIOL MOL GENET//PLANT BIOL GRP PROGRAM |
| 32132 | 112 | SOIL WATER STRESS THRESHOLD//GR H SCI OURCE//SOIL WATER CONSERVAT ECOENVIRONM |
| 33493 | 91 | TSP50//EDF 1//MBF1 |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | DEHYDRIN | Author keyword | 219 | 76% | 2% | 152 |
| 2 | ABIOTIC STRESS | Author keyword | 140 | 23% | 6% | 524 |
| 3 | DREB | Author keyword | 118 | 95% | 0% | 39 |
| 4 | WRKY TRANSCRIPTION FACTOR | Author keyword | 115 | 84% | 1% | 63 |
| 5 | DEHYDRINS | Author keyword | 92 | 72% | 1% | 73 |
| 6 | LEA PROTEIN | Author keyword | 90 | 87% | 0% | 45 |
| 7 | WRKY | Author keyword | 84 | 71% | 1% | 68 |
| 8 | ABSCISIC ACID | Author keyword | 74 | 15% | 5% | 463 |
| 9 | AP2 ERF | Author keyword | 73 | 86% | 0% | 37 |
| 10 | LEA PROTEINS | Author keyword | 72 | 69% | 1% | 62 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | DEHYDRIN | 219 | 76% | 2% | 152 | Search DEHYDRIN | Search DEHYDRIN |
| 2 | ABIOTIC STRESS | 140 | 23% | 6% | 524 | Search ABIOTIC+STRESS | Search ABIOTIC+STRESS |
| 3 | DREB | 118 | 95% | 0% | 39 | Search DREB | Search DREB |
| 4 | WRKY TRANSCRIPTION FACTOR | 115 | 84% | 1% | 63 | Search WRKY+TRANSCRIPTION+FACTOR | Search WRKY+TRANSCRIPTION+FACTOR |
| 5 | DEHYDRINS | 92 | 72% | 1% | 73 | Search DEHYDRINS | Search DEHYDRINS |
| 6 | LEA PROTEIN | 90 | 87% | 0% | 45 | Search LEA+PROTEIN | Search LEA+PROTEIN |
| 7 | WRKY | 84 | 71% | 1% | 68 | Search WRKY | Search WRKY |
| 8 | ABSCISIC ACID | 74 | 15% | 5% | 463 | Search ABSCISIC+ACID | Search ABSCISIC+ACID |
| 9 | AP2 ERF | 73 | 86% | 0% | 37 | Search AP2+ERF | Search AP2+ERF |
| 10 | LEA PROTEINS | 72 | 69% | 1% | 62 | Search LEA+PROTEINS | Search LEA+PROTEINS |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | FREEZING TOLERANCE | 499 | 42% | 10% | 913 |
| 2 | ABSCISIC ACID | 313 | 16% | 19% | 1749 |
| 3 | RESPONSIVE GENE EXPRESSION | 292 | 66% | 3% | 268 |
| 4 | HIGH SALINITY STRESSES | 173 | 68% | 2% | 152 |
| 5 | GCC BOX | 151 | 79% | 1% | 97 |
| 6 | COLD ACCLIMATION | 146 | 26% | 5% | 493 |
| 7 | ABIOTIC STRESS | 146 | 24% | 6% | 523 |
| 8 | ETHYLENE RESPONSIVE ELEMENT | 125 | 71% | 1% | 100 |
| 9 | LENGTH CDNA MICROARRAY | 123 | 60% | 1% | 134 |
| 10 | LEA PROTEINS | 123 | 64% | 1% | 119 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Abscisic Acid: Emergence of a Core Signaling Network | 2010 | 494 | 176 | 59% |
| Transcriptional regulatory networks in cellular responses and tolerance to dehydration and cold stresses | 2006 | 828 | 131 | 86% |
| AP2/ERF family transcription factors in plant abiotic stress responses | 2012 | 144 | 118 | 74% |
| Plant cold acclimation: Freezing tolerance genes and regulatory mechanisms | 1999 | 1345 | 115 | 71% |
| NAC transcription factors in plant abiotic stress responses | 2012 | 124 | 55 | 80% |
| The transcriptional regulatory network in the drought response and its crosstalk in abiotic stress responses including drought, cold, and heat | 2014 | 18 | 74 | 92% |
| WRKY transcription factors | 2010 | 358 | 96 | 63% |
| Mitogen-activated protein kinase cascades in signaling plant growth and development | 2015 | 4 | 91 | 49% |
| NAC proteins: regulation and role in stress tolerance | 2012 | 99 | 90 | 81% |
| Tolerance to drought and salt stress in plants: unraveling the signaling networks | 2014 | 35 | 117 | 41% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | GENE DISCOVERY GRP | 19 | 39% | 0.4% | 38 |
| 2 | BIOL OURCES POSTHARVEST | 16 | 48% | 0.3% | 25 |
| 3 | PLANT MOL PHYSIOL | 15 | 20% | 0.8% | 70 |
| 4 | GR H SCI OURCE | 12 | 86% | 0.1% | 6 |
| 5 | ABT MOL PHYTOPATHOL | 11 | 100% | 0.1% | 6 |
| 6 | MOE SYST BIOL BIOINFORMAT | 11 | 100% | 0.1% | 6 |
| 7 | SIGNALING PATHWAY UNIT | 9 | 39% | 0.2% | 19 |
| 8 | BIOL GENET IMPROVEMENT HORT CROPS NW RE | 9 | 83% | 0.1% | 5 |
| 9 | DORMANCY AD TAT UNIT | 9 | 83% | 0.1% | 5 |
| 10 | MOL CELLULAR BREEDING GRP | 9 | 83% | 0.1% | 5 |
Related classes at same level (level 2) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000024613 | SALT TOLERANCE//SALT STRESS//GLYCINEBETAINE |
| 2 | 0.0000019375 | GUARD CELL//GUARD CELLS//NYCTINASTY |
| 3 | 0.0000017450 | CYTOKININ//CYTOKININS//CYTOKININ OXIDASE DEHYDROGENASE |
| 4 | 0.0000017303 | AERENCHYMA//WATERLOGGING//SUBMERGENCE |
| 5 | 0.0000017066 | LEAF SENESCENCE//CHLOROPHYLLASE//CHLOROPHYLL DEGRADATION |
| 6 | 0.0000015833 | MADS BOX GENES//MADS BOX//FLOWER DEVELOPMENT |
| 7 | 0.0000015810 | PHYSIOLOGICAL AND MOLECULAR PLANT PATHOLOGY//LATE BLIGHT//PHYTOPHTHORA INFESTANS |
| 8 | 0.0000015759 | JASMONIC ACID//HYDROPEROXIDE LYASE//ALLENE OXIDE SYNTHASE |
| 9 | 0.0000015380 | PLANT DEFENSIN//BETA AMYLASE//LIMIT DEXTRINASE |
| 10 | 0.0000014897 | ROOT GROWTH POTENTIAL//SEEDLING QUALITY//DEEP SUPERCOOLING |