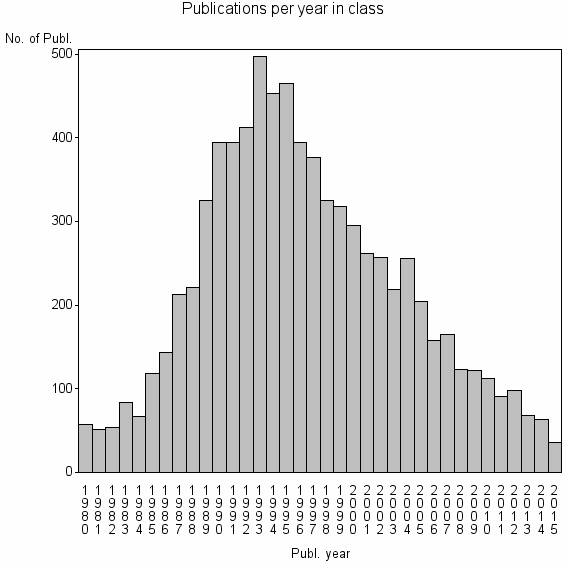
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 1339 | 7890 | 24.7 | 75% |
Classes in level above (level 3) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 159 | 56907 | BIOINFORMATICS//BMC BIOINFORMATICS//MATHEMATICAL & COMPUTATIONAL BIOLOGY |
Classes in level below (level 1) |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | BIOTECHNIQUES | Journal | 96 | 12% | 9% | 730 |
| 2 | PCR-METHODS AND APPLICATIONS | Journal | 20 | 32% | 1% | 52 |
| 3 | DIFFERENTIAL DISPLAY | Author keyword | 15 | 12% | 2% | 120 |
| 4 | SERIAL ANALYSIS OF GENE EXPRESSION | Author keyword | 14 | 30% | 1% | 41 |
| 5 | SAGE | Author keyword | 13 | 14% | 1% | 85 |
| 6 | MKIAA | Author keyword | 13 | 80% | 0% | 8 |
| 7 | MRNA DIFFERENTIAL DISPLAY | Author keyword | 12 | 25% | 1% | 42 |
| 8 | ANIM NUTR FEED YUNNAN PROV | Address | 11 | 54% | 0% | 14 |
| 9 | DIGOXIGENIN | Author keyword | 10 | 18% | 1% | 52 |
| 10 | DYE TERMINATOR | Author keyword | 9 | 83% | 0% | 5 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | DIFFERENTIAL DISPLAY | 15 | 12% | 2% | 120 | Search DIFFERENTIAL+DISPLAY | Search DIFFERENTIAL+DISPLAY |
| 2 | SERIAL ANALYSIS OF GENE EXPRESSION | 14 | 30% | 1% | 41 | Search SERIAL+ANALYSIS+OF+GENE+EXPRESSION | Search SERIAL+ANALYSIS+OF+GENE+EXPRESSION |
| 3 | SAGE | 13 | 14% | 1% | 85 | Search SAGE | Search SAGE |
| 4 | MKIAA | 13 | 80% | 0% | 8 | Search MKIAA | Search MKIAA |
| 5 | MRNA DIFFERENTIAL DISPLAY | 12 | 25% | 1% | 42 | Search MRNA+DIFFERENTIAL+DISPLAY | Search MRNA+DIFFERENTIAL+DISPLAY |
| 6 | DIGOXIGENIN | 10 | 18% | 1% | 52 | Search DIGOXIGENIN | Search DIGOXIGENIN |
| 7 | DYE TERMINATOR | 9 | 83% | 0% | 5 | Search DYE+TERMINATOR | Search DYE+TERMINATOR |
| 8 | GENOME WALKING | 7 | 29% | 0% | 22 | Search GENOME+WALKING | Search GENOME+WALKING |
| 9 | SERIAL ANALYSIS OF GENE EXPRESSION SAGE | 6 | 34% | 0% | 15 | Search SERIAL+ANALYSIS+OF+GENE+EXPRESSION+SAGE | Search SERIAL+ANALYSIS+OF+GENE+EXPRESSION+SAGE |
| 10 | FULL LENGTH CDNA | 6 | 20% | 0% | 27 | Search FULL+LENGTH+CDNA | Search FULL+LENGTH+CDNA |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | SERIAL ANALYSIS | 84 | 28% | 3% | 257 |
| 2 | CROSS COMBINATION | 23 | 74% | 0% | 17 |
| 3 | CHEMI LUMINESCENT | 17 | 42% | 0% | 32 |
| 4 | PHOTOACTIVATABLE BIOTIN | 17 | 100% | 0% | 8 |
| 5 | CAP TRAPPER | 16 | 56% | 0% | 20 |
| 6 | GENOMIC WALKING | 15 | 67% | 0% | 14 |
| 7 | BIOTINYLATED OLIGONUCLEOTIDE PROBES | 15 | 62% | 0% | 16 |
| 8 | TRAPPER SELECTED CDNAS | 15 | 88% | 0% | 7 |
| 9 | HAPTEN DIGOXIGENIN | 14 | 100% | 0% | 7 |
| 10 | MODULAR PRIMERS | 14 | 100% | 0% | 7 |
Journals |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | BIOTECHNIQUES | 96 | 12% | 9% | 730 |
| 2 | PCR-METHODS AND APPLICATIONS | 20 | 32% | 1% | 52 |
| 3 | GENETIC ANALYSIS-BIOMOLECULAR ENGINEERING | 4 | 12% | 0% | 32 |
| 4 | GENE ANALYSIS TECHNIQUES | 2 | 12% | 0% | 12 |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Suppression subtractive hybridization: A versatile method for identifying differentially expressed genes | 1999 | 288 | 20 | 90% |
| Developments in quantitative PCR | 1998 | 215 | 109 | 58% |
| Understanding SAGE data | 2007 | 47 | 47 | 72% |
| Integrative annotation of 21,037 human genes validated by full-length cDNA clones | 2004 | 202 | 96 | 30% |
| Technical aspects of quantitative competitive PCR | 1996 | 219 | 102 | 67% |
| RECENT ADVANCES IN DIFFERENTIAL DISPLAY | 1995 | 210 | 30 | 77% |
| Tag-based approaches for transcriptome research and genome annotation | 2005 | 89 | 51 | 49% |
| RAPID AMPLIFICATION OF COMPLEMENTARY-DNA ENDS FOR GENERATION OF FULL-LENGTH COMPLEMENTARY DNAS - THERMAL RACE | 1993 | 453 | 11 | 45% |
| Serial Analysis of Gene Expression (SAGE): 13 Years of Application in Research | 2008 | 29 | 128 | 73% |
| Comparison and critical evaluation of PCR-mediated methods to walk along the sequence of genomic DNA | 2009 | 15 | 39 | 79% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ANIM NUTR FEED YUNNAN PROV | 11 | 54% | 0.2% | 14 |
| 2 | TAISHO FUNCT GENOM | 7 | 42% | 0.2% | 13 |
| 3 | GENOMAT | 4 | 67% | 0.1% | 4 |
| 4 | IMAGE CONSORTIUM | 4 | 67% | 0.1% | 4 |
| 5 | MOL GERONTOL UNIT | 4 | 67% | 0.1% | 4 |
| 6 | GENOME EXPLORAT GRPTSURUMI KU | 4 | 28% | 0.2% | 13 |
| 7 | TEDA BIOX SYST BIOTECHNOL | 3 | 60% | 0.0% | 3 |
| 8 | GENOME EXPLORAT | 2 | 38% | 0.1% | 5 |
| 9 | ENH | 2 | 21% | 0.1% | 10 |
| 10 | GENOME EXPLORAT GRP | 2 | 12% | 0.2% | 17 |
Related classes at same level (level 2) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000018582 | SEQUENCING BY HYBRIDIZATION//MUTATION DETECTION//TILLING |
| 2 | 0.0000015286 | REFERENCE GENES//REFERENCE GENE//GENORM |
| 3 | 0.0000013707 | RBM5//MEIOTIC DRIVE//HUMAN CHROMOSOME 3 |
| 4 | 0.0000011243 | RESTRICTION ENDONUCLEASE//RESTRICTION MODIFICATION//RESTRICTION MODIFICATION SYSTEM |
| 5 | 0.0000010365 | SINGLE LOCUS PROBES//MINISATELLITE//MISATTRIBUTED PATERNITY |
| 6 | 0.0000009949 | GMO DETECTION//EVENT SPECIFIC//LOOP MEDIATED ISOTHERMAL AMPLIFICATION |
| 7 | 0.0000009841 | BICLUSTERING//PROGRAM TRANSLAT IMMUNOVIROL BIODEF//MAYO VACCINE GRP |
| 8 | 0.0000009545 | ANTIGEN RETRIEVAL//JOURNAL OF HISTOTECHNOLOGY//BIOTECHNIC & HISTOCHEMISTRY |
| 9 | 0.0000007652 | NEXT GENERATION SEQUENCING//RNA SEQ//COPY NUMBER VARIATION |
| 10 | 0.0000005829 | HUMAN ENDOGENOUS RETROVIRUS//ADAR//ENDOGENOUS RETROVIRUS |