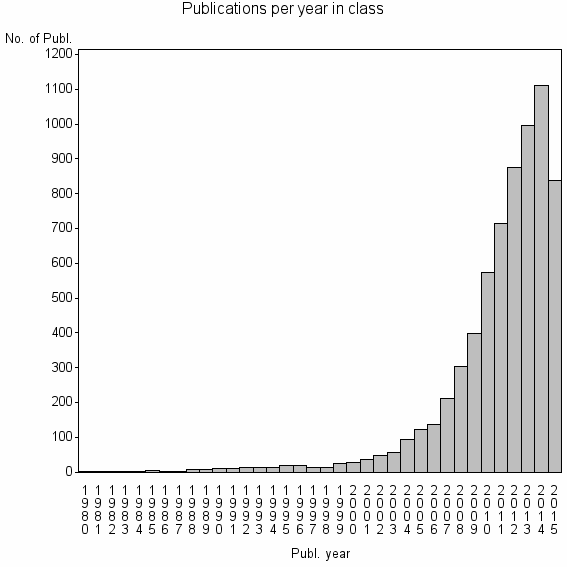
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 1568 | 6715 | 37.4 | 86% |
Classes in level above (level 3) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 159 | 56907 | BIOINFORMATICS//BMC BIOINFORMATICS//MATHEMATICAL & COMPUTATIONAL BIOLOGY |
Classes in level below (level 1) |
| ID, lev. below | Publications | Label for level below |
|---|---|---|
| 1159 | 2714 | GENOME ASSEMBLY//VARIANT CALLING//TARGET ENRICHMENT |
| 5163 | 1594 | COPY NUMBER VARIATION//CCL3L1//PENNCNV |
| 7698 | 1259 | RNA SEQ//DE NOVO ASSEMBLY//TRANSCRIPTOME ASSEMBLY |
| 16697 | 576 | SPLICED LEADER//TRANS SPLICING//CIRCULAR RNA |
| 21583 | 356 | EPIDEMIOL PATHOPHYSIOL ONCOGEN VIRUS UNIT//LYSSAVIRUS DYNAM HOST AD TAT UNIT//VIRAL DISCOVERY |
| 26161 | 216 | MAPPING BY SEQUENCING//RADSEQ//REDUCED REPRESENTATION LIBRARIES |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | NEXT GENERATION SEQUENCING | Author keyword | 75 | 15% | 7% | 449 |
| 2 | RNA SEQ | Author keyword | 50 | 18% | 4% | 251 |
| 3 | COPY NUMBER VARIATION | Author keyword | 34 | 21% | 2% | 145 |
| 4 | DE NOVO ASSEMBLY | Author keyword | 30 | 41% | 1% | 56 |
| 5 | SPLICED LEADER | Author keyword | 21 | 46% | 0% | 33 |
| 6 | TRANS SPLICING | Author keyword | 20 | 28% | 1% | 61 |
| 7 | GENOME ASSEMBLY | Author keyword | 20 | 38% | 1% | 41 |
| 8 | CCL3L1 | Author keyword | 19 | 76% | 0% | 13 |
| 9 | VARIANT CALLING | Author keyword | 16 | 65% | 0% | 15 |
| 10 | PENNCNV | Author keyword | 13 | 80% | 0% | 8 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | NEXT GENERATION SEQUENCING | 75 | 15% | 7% | 449 | Search NEXT+GENERATION+SEQUENCING | Search NEXT+GENERATION+SEQUENCING |
| 2 | RNA SEQ | 50 | 18% | 4% | 251 | Search RNA+SEQ | Search RNA+SEQ |
| 3 | COPY NUMBER VARIATION | 34 | 21% | 2% | 145 | Search COPY+NUMBER+VARIATION | Search COPY+NUMBER+VARIATION |
| 4 | DE NOVO ASSEMBLY | 30 | 41% | 1% | 56 | Search DE+NOVO+ASSEMBLY | Search DE+NOVO+ASSEMBLY |
| 5 | SPLICED LEADER | 21 | 46% | 0% | 33 | Search SPLICED+LEADER | Search SPLICED+LEADER |
| 6 | TRANS SPLICING | 20 | 28% | 1% | 61 | Search TRANS+SPLICING | Search TRANS+SPLICING |
| 7 | GENOME ASSEMBLY | 20 | 38% | 1% | 41 | Search GENOME+ASSEMBLY | Search GENOME+ASSEMBLY |
| 8 | CCL3L1 | 19 | 76% | 0% | 13 | Search CCL3L1 | Search CCL3L1 |
| 9 | VARIANT CALLING | 16 | 65% | 0% | 15 | Search VARIANT+CALLING | Search VARIANT+CALLING |
| 10 | PENNCNV | 13 | 80% | 0% | 8 | Search PENNCNV | Search PENNCNV |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | STRUCTURAL VARIATION | 218 | 40% | 6% | 428 |
| 2 | ARRAY CGH DATA | 176 | 66% | 2% | 163 |
| 3 | READS | 88 | 53% | 2% | 115 |
| 4 | BURROWS WHEELER TRANSFORM | 86 | 50% | 2% | 125 |
| 5 | READ ALIGNMENT | 82 | 44% | 2% | 141 |
| 6 | SHORT DNA SEQUENCES | 71 | 71% | 1% | 58 |
| 7 | SHORT READ ALIGNMENT | 68 | 51% | 1% | 94 |
| 8 | SEGMENTAL DUPLICATIONS | 67 | 30% | 3% | 192 |
| 9 | COPY NUMBER VARIATION | 66 | 19% | 5% | 314 |
| 10 | CIRCULAR BINARY SEGMENTATION | 61 | 59% | 1% | 68 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references |
% act. ref. to same field |
|---|---|---|---|---|
| APPLICATIONS OF NEXT-GENERATION SEQUENCING Sequencing technologies - the next generation | 2010 | 1743 | 91 | 59% |
| RNA-Seq: a revolutionary tool for transcriptomics | 2009 | 2552 | 36 | 44% |
| Sequencing depth and coverage: key considerations in genomic analyses | 2014 | 56 | 96 | 51% |
| Field guide to next-generation DNA sequencers | 2011 | 294 | 12 | 92% |
| Detecting and characterizing circular RNAs | 2014 | 39 | 48 | 58% |
| Next-generation transcriptome assembly | 2011 | 285 | 66 | 76% |
| Genome-wide genetic marker discovery and genotyping using next-generation sequencing | 2011 | 445 | 73 | 51% |
| Genotype and SNP calling from next-generation sequencing data | 2011 | 298 | 50 | 68% |
| Repetitive DNA and next-generation sequencing: computational challenges and solutions | 2012 | 153 | 72 | 72% |
| Solving the molecular diagnostic testing conundrum for Mendelian disorders in the era of next-generation sequencing: single-gene, gene panel, or exome/genome sequencing | 2015 | 6 | 37 | 43% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | SIMONS QUANTITAT BIOL | 10 | 57% | 0.2% | 12 |
| 2 | HKU BGI BIOINFORMAT ALGORITHMS CORE TECHNOL | 7 | 67% | 0.1% | 6 |
| 3 | STANLEY COGNIT GENOM | 5 | 37% | 0.2% | 11 |
| 4 | PROGRAM COMPUTAT BIOL BIOINFORMAT | 5 | 12% | 0.6% | 38 |
| 5 | DIAGNOST GENET UNIT | 4 | 38% | 0.1% | 8 |
| 6 | INTEGRATED GENOME POLYMORPHISM | 4 | 27% | 0.2% | 12 |
| 7 | SHANDONG PROV ANIM DIS CONTROL BREEDING | 4 | 46% | 0.1% | 6 |
| 8 | HLTH SOLUT GRP | 3 | 43% | 0.1% | 6 |
| 9 | BINA TECHNOL | 3 | 100% | 0.0% | 3 |
| 10 | CHAIR BIOINFORMAT GRP | 3 | 100% | 0.0% | 3 |
Related classes at same level (level 2) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000017956 | GEMININ//DNA REPLICATION//CDC6 |
| 2 | 0.0000015818 | REFERENCE GENES//REFERENCE GENE//GENORM |
| 3 | 0.0000015541 | GENETIC EPIDEMIOLOGY//GENOMIC SELECTION//BMC GENETICS |
| 4 | 0.0000014844 | BICLUSTERING//PROGRAM TRANSLAT IMMUNOVIROL BIODEF//MAYO VACCINE GRP |
| 5 | 0.0000014178 | BIOINFORMATICS//BMC BIOINFORMATICS//GENOME RESEARCH |
| 6 | 0.0000014158 | SEQUENCING BY HYBRIDIZATION//MUTATION DETECTION//TILLING |
| 7 | 0.0000012609 | CHROMATIN SUPRAORGANIZATION//ZNF703//NUCLEAR PHENOTYPES |
| 8 | 0.0000009193 | LANDSCAPE GENETICS//INBREEDING DEPRESSION//COALESCENT |
| 9 | 0.0000008489 | HUMAN ENDOGENOUS RETROVIRUS//ADAR//ENDOGENOUS RETROVIRUS |
| 10 | 0.0000008430 | WOLF HIRSCHHORN SYNDROME//HOLOPROSENCEPHALY//ANNALES DE GENETIQUE |