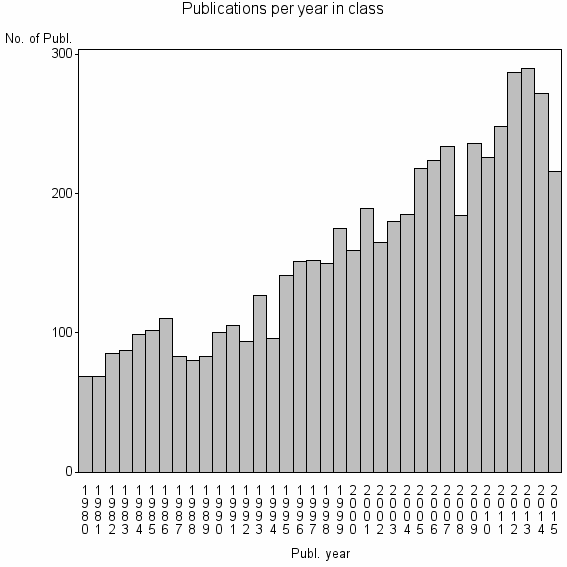
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 1825 | 5671 | 43.6 | 82% |
Classes in level above (level 3) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 32 | 105649 | VIROLOGY//AIDS RESEARCH AND HUMAN RETROVIRUSES//HIV 1 |
Classes in level below (level 1) |
| ID, lev. below | Publications | Label for level below |
|---|---|---|
| 1929 | 2352 | BIOL COMPUTAT VISUALIZAT//SINE//NON LTR RETROTRANSPOSON |
| 4941 | 1630 | HUMAN ENDOGENOUS RETROVIRUS//ENDOGENOUS RETROVIRUS//HERV W |
| 12283 | 853 | ADAR//RNA EDITING//ADAR2 |
| 20313 | 403 | VL30//INTRACISTERNAL A PARTICLE//EARLY TRANSPOSON |
| 22197 | 333 | RT ASSAY//VIRUS FILTRATION//VIRAL CLEARANCE |
| 32948 | 100 | P54NRB//POLYPYRIMIDINE TRACT BINDING PROTEIN ASSOCIATED SPLICING FACTOR//MATRIN 3 |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | HUMAN ENDOGENOUS RETROVIRUS | Author keyword | 139 | 84% | 1% | 75 |
| 2 | ADAR | Author keyword | 127 | 78% | 1% | 83 |
| 3 | ENDOGENOUS RETROVIRUS | Author keyword | 118 | 62% | 2% | 123 |
| 4 | UNITE RETROVIRUS ENDOGENES ELEMENTS RETROIDES E | Address | 91 | 83% | 1% | 52 |
| 5 | HERV W | Author keyword | 82 | 92% | 1% | 33 |
| 6 | SYNCYTIN | Author keyword | 68 | 80% | 1% | 43 |
| 7 | HERV | Author keyword | 66 | 74% | 1% | 50 |
| 8 | RNA EDITING | Author keyword | 66 | 25% | 4% | 234 |
| 9 | BIOL COMPUTAT VISUALIZAT | Address | 58 | 62% | 1% | 60 |
| 10 | RETROTRANSPOSITION | Author keyword | 58 | 50% | 1% | 83 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | HUMAN ENDOGENOUS RETROVIRUS | 139 | 84% | 1% | 75 | Search HUMAN+ENDOGENOUS+RETROVIRUS | Search HUMAN+ENDOGENOUS+RETROVIRUS |
| 2 | ADAR | 127 | 78% | 1% | 83 | Search ADAR | Search ADAR |
| 3 | ENDOGENOUS RETROVIRUS | 118 | 62% | 2% | 123 | Search ENDOGENOUS+RETROVIRUS | Search ENDOGENOUS+RETROVIRUS |
| 4 | HERV W | 82 | 92% | 1% | 33 | Search HERV+W | Search HERV+W |
| 5 | SYNCYTIN | 68 | 80% | 1% | 43 | Search SYNCYTIN | Search SYNCYTIN |
| 6 | HERV | 66 | 74% | 1% | 50 | Search HERV | Search HERV |
| 7 | RNA EDITING | 66 | 25% | 4% | 234 | Search RNA+EDITING | Search RNA+EDITING |
| 8 | RETROTRANSPOSITION | 58 | 50% | 1% | 83 | Search RETROTRANSPOSITION | Search RETROTRANSPOSITION |
| 9 | HERV K | 58 | 83% | 1% | 33 | Search HERV+K | Search HERV+K |
| 10 | SINE | 54 | 44% | 2% | 92 | Search SINE | Search SINE |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | HERV K | 309 | 93% | 2% | 117 |
| 2 | HERV W | 277 | 88% | 2% | 129 |
| 3 | L1 RETROTRANSPOSITION | 173 | 56% | 4% | 211 |
| 4 | HUMAN TRANSPOSABLE ELEMENT | 160 | 79% | 2% | 102 |
| 5 | ADENOSINE DEAMINASES | 158 | 79% | 2% | 100 |
| 6 | ORF1 PROTEIN | 149 | 88% | 1% | 69 |
| 7 | INSERTIONAL POLYMORPHISMS | 140 | 92% | 1% | 56 |
| 8 | HUMAN ENDOGENOUS RETROVIRUS | 132 | 65% | 2% | 126 |
| 9 | HIGH FREQUENCY RETROTRANSPOSITION | 131 | 88% | 1% | 61 |
| 10 | HUMAN L1 RETROTRANSPOSITION | 122 | 87% | 1% | 60 |
Journals |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | MOBILE DNA | 10 | 27% | 1% | 32 |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| The impact of retrotransposons on human genome evolution | 2009 | 388 | 156 | 85% |
| Functions and Regulation of RNA Editing by ADAR Deaminases | 2010 | 251 | 149 | 81% |
| Mobile elements: Drivers of genome evolution | 2004 | 813 | 69 | 43% |
| Retrotransposons revisited: The restraint and rehabilitation of parasites | 2008 | 223 | 133 | 67% |
| Initial sequencing and analysis of the human genome | 2001 | 10496 | 369 | 11% |
| LINE-1 Elements in Structural Variation and Disease | 2011 | 82 | 252 | 72% |
| Alu repeats and human genomic diversity | 2002 | 617 | 113 | 65% |
| From ancestral infectious retroviruses to bona fide cellular genes: Role of the captured syncytins in placentation | 2012 | 56 | 45 | 78% |
| RNA editing by adenosine deaminases that act on RNA | 2002 | 557 | 114 | 65% |
| Endogenous Retroviruses and the Development of Cancer | 2014 | 12 | 78 | 68% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | UNITE RETROVIRUS ENDOGENES ELEMENTS RETROIDES E | 91 | 83% | 0.9% | 52 |
| 2 | BIOL COMPUTAT VISUALIZAT | 58 | 62% | 1.1% | 60 |
| 3 | UMR 8122 | 43 | 68% | 0.7% | 38 |
| 4 | RNA EDITING | 30 | 100% | 0.2% | 12 |
| 5 | PBBRC | 30 | 79% | 0.3% | 19 |
| 6 | EUKARYOT GENOME EVOLUT | 23 | 100% | 0.2% | 10 |
| 7 | UNITE MIXTE RECH 8122 | 17 | 100% | 0.1% | 8 |
| 8 | UMR 1573 | 13 | 71% | 0.2% | 10 |
| 9 | POWIC | 12 | 52% | 0.3% | 17 |
| 10 | SIGNAL TRANSDUCTSAKYO KU | 11 | 60% | 0.2% | 12 |
Related classes at same level (level 2) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000010859 | MADS BOX GENES//MADS BOX//FLOWER DEVELOPMENT |
| 2 | 0.0000010449 | APOBEC3G//INTEGRASE//HIV 1 INTEGRASE |
| 3 | 0.0000010186 | CHROMOSOMA//SYNAPTONEMAL COMPLEX//MEIOSIS |
| 4 | 0.0000009110 | HUMAN HYALINE CARTILAGE//JITTER FREE//BLUE CHROMOPHORES |
| 5 | 0.0000008489 | NEXT GENERATION SEQUENCING//RNA SEQ//COPY NUMBER VARIATION |
| 6 | 0.0000008280 | FELINE IMMUNODEFICIENCY VIRUS//CAPRINE ARTHRITIS ENCEPHALITIS VIRUS//FIV |
| 7 | 0.0000008165 | RBM5//MEIOTIC DRIVE//HUMAN CHROMOSOME 3 |
| 8 | 0.0000007638 | BIOINFORMATICS//BMC BIOINFORMATICS//GENOME RESEARCH |
| 9 | 0.0000007377 | MAMMARY GLAND//MOUSE MAMMARY TUMOR VIRUS//JOURNAL OF MAMMARY GLAND BIOLOGY AND NEOPLASIA |
| 10 | 0.0000007032 | PIRNA//RNA INTERFERENCE//PIWI |