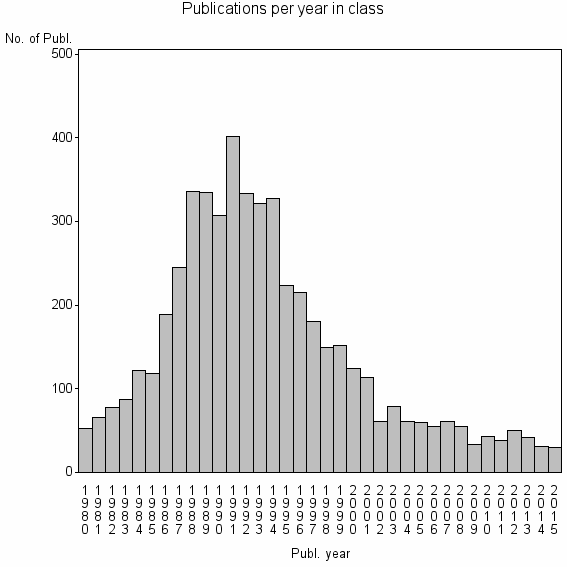
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 1959 | 5166 | 34.0 | 70% |
Classes in level above (level 3) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 96 | 73697 | DNA METHYLATION//FRAGILE X SYNDROME//RETT SYNDROME |
Classes in level below (level 1) |
| ID, lev. below | Publications | Label for level below |
|---|---|---|
| 5742 | 1515 | BIBAC//TAR CLONING//BIBAC LIBRARY |
| 7994 | 1228 | MRC HUMAN ECOGENET UNIT//BIOCHEM UNIV PLEIN 1//BORN BUNGE FDN NEUROL NEUROPATHOL |
| 12630 | 829 | MEIOTIC DRIVE//T HAPLOTYPE//T COMPLEX |
| 14256 | 715 | ANIM GENET BIOINFORMAT//BASIC PROGRAMMAMMALIAN GENET//FREDERICK CANC ILBIONET INC |
| 18232 | 497 | RBM5//HUMAN CHROMOSOME 3//LUCA 15 |
| 20862 | 382 | HAPPY MAPPING//MULTILOCUS ORDERING//EXOGAMIC POPULATIONS |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | RBM5 | Author keyword | 65 | 95% | 0% | 21 |
| 2 | MEIOTIC DRIVE | Author keyword | 42 | 48% | 1% | 64 |
| 3 | HUMAN CHROMOSOME 3 | Author keyword | 21 | 73% | 0% | 16 |
| 4 | BIBAC | Author keyword | 21 | 90% | 0% | 9 |
| 5 | T HAPLOTYPE | Author keyword | 21 | 85% | 0% | 11 |
| 6 | LUCA 15 | Author keyword | 14 | 100% | 0% | 7 |
| 7 | T COMPLEX | Author keyword | 12 | 40% | 0% | 23 |
| 8 | CANC CAUSING GENES SECT | Address | 11 | 78% | 0% | 7 |
| 9 | NOTI MICROARRAYS | Author keyword | 9 | 83% | 0% | 5 |
| 10 | TAR CLONING | Author keyword | 7 | 64% | 0% | 7 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | RBM5 | 65 | 95% | 0% | 21 | Search RBM5 | Search RBM5 |
| 2 | MEIOTIC DRIVE | 42 | 48% | 1% | 64 | Search MEIOTIC+DRIVE | Search MEIOTIC+DRIVE |
| 3 | HUMAN CHROMOSOME 3 | 21 | 73% | 0% | 16 | Search HUMAN+CHROMOSOME+3 | Search HUMAN+CHROMOSOME+3 |
| 4 | BIBAC | 21 | 90% | 0% | 9 | Search BIBAC | Search BIBAC |
| 5 | T HAPLOTYPE | 21 | 85% | 0% | 11 | Search T+HAPLOTYPE | Search T+HAPLOTYPE |
| 6 | LUCA 15 | 14 | 100% | 0% | 7 | Search LUCA+15 | Search LUCA+15 |
| 7 | T COMPLEX | 12 | 40% | 0% | 23 | Search T+COMPLEX | Search T+COMPLEX |
| 8 | NOTI MICROARRAYS | 9 | 83% | 0% | 5 | Search NOTI+MICROARRAYS | Search NOTI+MICROARRAYS |
| 9 | TAR CLONING | 7 | 64% | 0% | 7 | Search TAR+CLONING | Search TAR+CLONING |
| 10 | ELIMINATION TEST | 6 | 80% | 0% | 4 | Search ELIMINATION+TEST | Search ELIMINATION+TEST |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | TERT HAPLOTYPES | 69 | 93% | 1% | 26 |
| 2 | YEAST ARTIFICIAL CHROMOSOMES | 68 | 40% | 3% | 132 |
| 3 | TRANSMISSION RATIO DISTORTION | 56 | 48% | 2% | 87 |
| 4 | CANDIDATE GENE FAMILY | 37 | 100% | 0% | 14 |
| 5 | VIRILITY DEFICIENCY | 35 | 89% | 0% | 16 |
| 6 | ARTIFICIAL CHROMOSOME LIBRARY | 33 | 29% | 2% | 97 |
| 7 | HUMAN CHROMOSOME 3 | 31 | 49% | 1% | 45 |
| 8 | HOMOZYGOUS DELETION REGION | 30 | 47% | 1% | 48 |
| 9 | MEIOTIC DRIVE | 29 | 25% | 2% | 98 |
| 10 | T T COMPLEX | 27 | 72% | 0% | 21 |
Journals |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | GENETIC ANALYSIS-BIOMOLECULAR ENGINEERING | 4 | 12% | 1% | 31 |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| SEGREGATION DISTORTERS | 1991 | 391 | 89 | 71% |
| THE PECULIAR JOURNEY OF A SELFISH CHROMOSOME - MOUSE T-HAPLOTYPES AND MEIOTIC DRIVE | 1993 | 171 | 24 | 92% |
| Transmission ratio distortion in mice | 2003 | 70 | 55 | 93% |
| Sex chromosome meiotic drive | 2001 | 141 | 75 | 56% |
| CHEATERS SOMETIMES PROSPER - DISTORTION OF MENDELIAN SEGREGATION BY MEIOTIC DRIVE | 1993 | 126 | 29 | 90% |
| The role of meiotic drive in hybrid male sterility | 2010 | 29 | 70 | 43% |
| The 630-kb lung cancer homozygous deletion region on human chromosome 3p21.3: Identification and evaluation of the resident candidate tumor suppressor genes | 2000 | 346 | 117 | 18% |
| Mapping and sequencing complex genomes: Let's get physical! | 2004 | 48 | 49 | 69% |
| Putting the brake on drive: meiotic drive of t haplotypes in natural populations of mice | 1998 | 34 | 29 | 86% |
| Closing the (Ran)GAP on segregation distortion in Drosophila | 2003 | 45 | 24 | 58% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | CANC CAUSING GENES SECT | 11 | 78% | 0.1% | 7 |
| 2 | MRC HUMAN ECOGENET UNIT | 6 | 100% | 0.1% | 4 |
| 3 | ANIM GENET BIOINFORMAT | 4 | 56% | 0.1% | 5 |
| 4 | SRSNR | 3 | 60% | 0.1% | 3 |
| 5 | ARIZONA GENOM COMPUTAT | 3 | 32% | 0.1% | 7 |
| 6 | DEV TEAM MAMMALIAN CELLULAR DYNAM | 2 | 67% | 0.0% | 2 |
| 7 | BIOL GENET EVOLUCAO | 2 | 43% | 0.1% | 3 |
| 8 | ARIZONA GENOM | 1 | 10% | 0.3% | 13 |
| 9 | BASIC PROGRAMMAMMALIAN GENET | 1 | 50% | 0.0% | 2 |
| 10 | FO TRY FRUIT | 1 | 50% | 0.0% | 2 |
Related classes at same level (level 2) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000021908 | SINGLE LOCUS PROBES//MINISATELLITE//MISATTRIBUTED PATERNITY |
| 2 | 0.0000018855 | CHROMATIN SUPRAORGANIZATION//ZNF703//NUCLEAR PHENOTYPES |
| 3 | 0.0000013707 | BIOTECHNIQUES//PCR-METHODS AND APPLICATIONS//DIFFERENTIAL DISPLAY |
| 4 | 0.0000011519 | WOLF HIRSCHHORN SYNDROME//HOLOPROSENCEPHALY//ANNALES DE GENETIQUE |
| 5 | 0.0000011365 | XIST//X CHROMOSOME INACTIVATION//TSIX |
| 6 | 0.0000011103 | ANIMAL GENETICS//GENET CELLULAIRE//ANIM CYTOGENET GENE M PING |
| 7 | 0.0000010916 | HERITAGE MED 275//JCR LA CP RAT//SA GENE |
| 8 | 0.0000010542 | FRAGILE X SYNDROME//FRAGILE X//FMR1 |
| 9 | 0.0000009400 | RETINOBLASTOMA//CHEMOREDUCTION//OPHTHALM ONCOL SERV |
| 10 | 0.0000009038 | KABUKI SYNDROME//KABUKI MAKE UP SYNDROME//TRICHORHINOPHALANGEAL SYNDROME |