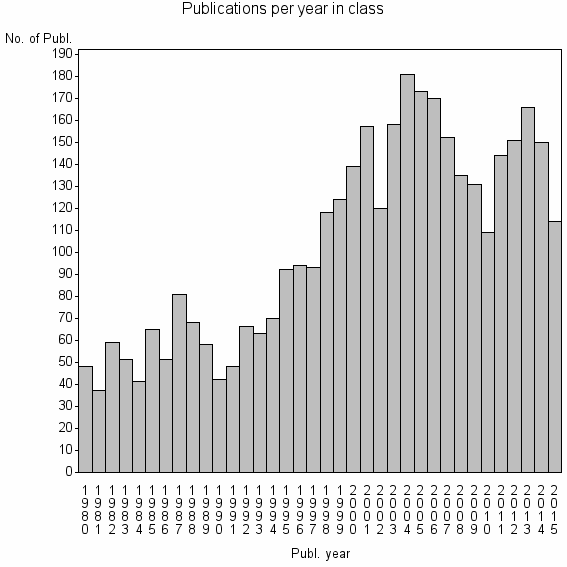
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 2410 | 3719 | 33.8 | 68% |
Classes in level above (level 3) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 295 | 37066 | CORYNEBACTERIUM GLUTAMICUM//ACTA CRYSTALLOGRAPHICA SECTION D-BIOLOGICAL CRYSTALLOGRAPHY//DIHYDRODIPICOLINATE SYNTHASE |
Classes in level below (level 1) |
| ID, lev. below | Publications | Label for level below |
|---|---|---|
| 9662 | 1067 | DIRECTED EVOLUTION//DNA SHUFFLING//EPPCR |
| 15395 | 644 | ASPARTASE//HISTIDASE//HISTIDINE AMMONIA LYASE |
| 17373 | 539 | HYDANTOINASE//D HYDANTOINASE//L N CARBAMOYLASE |
| 18110 | 503 | RIBOKINASE//ADENOSINE KINASE//GLUCONATE DEHYDRATASE |
| 21718 | 350 | AMINOACYLASE//D AMINOACYLASE//L AMINOACYLASE |
| 22938 | 307 | BIOCATALYTIC PROMISCUITY//4 OXALOCROTONATE TAUTOMERASE//TAUTOMERASE SUPERFAMILY |
| 24669 | 255 | MANDELATE RACEMASE//ENOLASE SUPERFAMILY//MANDELATE RACEMASE EC 5122 |
| 35047 | 54 | BIC GRANADA//HERBAL NANOPARTICLES//RAJAH MUTHAIAH MED HOSP |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | HYDANTOINASE | Author keyword | 119 | 88% | 2% | 57 |
| 2 | D HYDANTOINASE | Author keyword | 102 | 88% | 1% | 49 |
| 3 | DIRECTED EVOLUTION | Author keyword | 60 | 22% | 7% | 244 |
| 4 | L N CARBAMOYLASE | Author keyword | 38 | 89% | 0% | 17 |
| 5 | N CARBAMYL D AMINO ACID AMIDOHYDROLASE | Author keyword | 37 | 100% | 0% | 14 |
| 6 | D AMINOACYLASE | Author keyword | 32 | 85% | 0% | 17 |
| 7 | DIHYDROPYRIMIDINASE | Author keyword | 28 | 62% | 1% | 29 |
| 8 | BIOCATALYTIC PROMISCUITY | Author keyword | 27 | 92% | 0% | 11 |
| 9 | N CARBAMOYL D AMINO ACID AMIDOHYDROLASE | Author keyword | 26 | 100% | 0% | 11 |
| 10 | D P HYDROXYPHENYLGLYCINE | Author keyword | 24 | 82% | 0% | 14 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | HYDANTOINASE | 119 | 88% | 2% | 57 | Search HYDANTOINASE | Search HYDANTOINASE |
| 2 | D HYDANTOINASE | 102 | 88% | 1% | 49 | Search D+HYDANTOINASE | Search D+HYDANTOINASE |
| 3 | DIRECTED EVOLUTION | 60 | 22% | 7% | 244 | Search DIRECTED+EVOLUTION | Search DIRECTED+EVOLUTION |
| 4 | L N CARBAMOYLASE | 38 | 89% | 0% | 17 | Search L+N+CARBAMOYLASE | Search L+N+CARBAMOYLASE |
| 5 | N CARBAMYL D AMINO ACID AMIDOHYDROLASE | 37 | 100% | 0% | 14 | Search N+CARBAMYL+D+AMINO+ACID+AMIDOHYDROLASE | Search N+CARBAMYL+D+AMINO+ACID+AMIDOHYDROLASE |
| 6 | D AMINOACYLASE | 32 | 85% | 0% | 17 | Search D+AMINOACYLASE | Search D+AMINOACYLASE |
| 7 | DIHYDROPYRIMIDINASE | 28 | 62% | 1% | 29 | Search DIHYDROPYRIMIDINASE | Search DIHYDROPYRIMIDINASE |
| 8 | BIOCATALYTIC PROMISCUITY | 27 | 92% | 0% | 11 | Search BIOCATALYTIC+PROMISCUITY | Search BIOCATALYTIC+PROMISCUITY |
| 9 | N CARBAMOYL D AMINO ACID AMIDOHYDROLASE | 26 | 100% | 0% | 11 | Search N+CARBAMOYL+D+AMINO+ACID+AMIDOHYDROLASE | Search N+CARBAMOYL+D+AMINO+ACID+AMIDOHYDROLASE |
| 10 | D P HYDROXYPHENYLGLYCINE | 24 | 82% | 0% | 14 | Search D+P+HYDROXYPHENYLGLYCINE | Search D+P+HYDROXYPHENYLGLYCINE |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | D P HYDROXYPHENYLGLYCINE | 99 | 91% | 1% | 41 |
| 2 | 5 SUBSTITUTED HYDANTOINS | 91 | 84% | 1% | 49 |
| 3 | IN VITRO RECOMBINATION | 75 | 57% | 2% | 90 |
| 4 | THERMOSTABLE D HYDANTOINASE | 65 | 95% | 1% | 21 |
| 5 | DIRECTED EVOLUTION | 54 | 13% | 10% | 382 |
| 6 | AMINO ACID AMIDOHYDROLASE | 53 | 63% | 1% | 54 |
| 7 | 3 CHLOROACRYLIC ACID | 53 | 95% | 0% | 18 |
| 8 | D HYDANTOINASE | 52 | 72% | 1% | 41 |
| 9 | BIOCATALYTIC PROMISCUITY | 48 | 91% | 1% | 20 |
| 10 | ENOLASE SUPERFAMILY | 45 | 53% | 2% | 60 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Enzyme promiscuity: mechanism and applications | 2007 | 201 | 46 | 52% |
| Divergent Evolution in Enolase Superfamily: Strategies for Assigning Functions | 2012 | 36 | 34 | 79% |
| Directed evolution drives the next generation of biocatalysts | 2009 | 254 | 85 | 42% |
| Addressing the numbers problem in directed evolution | 2008 | 127 | 96 | 58% |
| Strategies for the discovery and engineering of enzymes for biocatalysis | 2013 | 36 | 47 | 51% |
| Hydrolase-catalyzed asymmetric carbon-carbon bond formation in organic synthesis | 2015 | 2 | 79 | 58% |
| Advances in laboratory evolution of enzymes | 2008 | 133 | 74 | 50% |
| Laboratory Evolution of Stereoselective Enzymes: A Prolific Source of Catalysts for Asymmetric Reactions | 2011 | 112 | 282 | 34% |
| Biocatalytic Promiscuity | 2011 | 74 | 81 | 49% |
| Status of protein engineering for biocatalysts: how to design an industrially useful biocatalyst | 2011 | 68 | 41 | 51% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | GRONINGEN PHARM | 9 | 23% | 0.9% | 33 |
| 2 | QUIM FIS BIOQUIM QUIM INORGAN | 7 | 31% | 0.5% | 19 |
| 3 | CHEM CHEM ENGN 21041 | 6 | 80% | 0.1% | 4 |
| 4 | BIOTECHNOL ENZYME CATALYSIS | 6 | 18% | 0.8% | 28 |
| 5 | FENSKE 112 | 4 | 67% | 0.1% | 4 |
| 6 | HEBEI FERMENTAT ENGN | 3 | 57% | 0.1% | 4 |
| 7 | ALLGEMEINE MIKROBIOL | 3 | 13% | 0.6% | 22 |
| 8 | DPTO QUIM FIS BIOQUIM QUIM INORGAN | 3 | 50% | 0.1% | 4 |
| 9 | TECH CHEM BIOTECHNOL | 3 | 20% | 0.3% | 11 |
| 10 | INVEST BIOTECNOL AGROALIMENTARIA | 2 | 32% | 0.2% | 6 |
Related classes at same level (level 2) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000017949 | ASYMMETRIC REDUCTION//BAEYER VILLIGER OXIDATION//OMEGA TRANSAMINASE |
| 2 | 0.0000015087 | LIPASE//LIPASES//CUTINASE |
| 3 | 0.0000013874 | ALANINE RACEMASE//TYROSINE PHENOL LYASE//MURD |
| 4 | 0.0000012143 | 3 ISOPROPYLMALATE DEHYDROGENASE//GLUTAMATE DEHYDROGENASE//ISOCITRATE DEHYDROGENASE |
| 5 | 0.0000011193 | L ARABINOSE ISOMERASE//RARE SUGAR//D PSICOSE |
| 6 | 0.0000010861 | PHAGE DISPLAY//CATALYTIC ANTIBODIES//CATALYTIC ANTIBODY |
| 7 | 0.0000010511 | CORYNEBACTERIUM GLUTAMICUM//SHIKIMATE PATHWAY//SHIKIMATE KINASE |
| 8 | 0.0000009974 | CELL FREE PROTEIN SYNTHESIS//MEMBRANE PROTEIN//RIBOSOME DISPLAY |
| 9 | 0.0000009807 | PENICILLIN ACYLASE//PENICILLIN G ACYLASE//LACTOSE HYDROLYSIS |
| 10 | 0.0000008878 | LESCH NYHAN SYNDROME//TIAZOFURIN//LESCH NYHAN DISEASE |