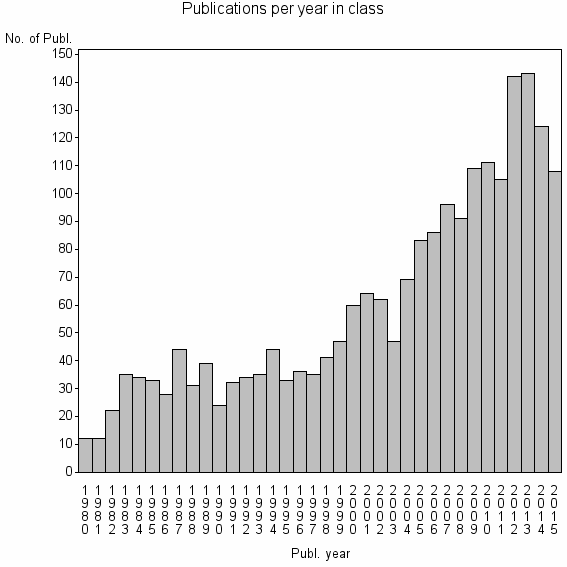
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 2974 | 2151 | 41.6 | 81% |
Classes in level above (level 3) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 204 | 48455 | PROTEIN FOLDING//PROTEINS-STRUCTURE FUNCTION AND BIOINFORMATICS//PROTEIN STRUCTURE PREDICTION |
Classes in level below (level 1) |
| ID, lev. below | Publications | Label for level below |
|---|---|---|
| 10905 | 957 | ISOASPARTATE//PROTEIN L ISOASPARTYL METHYLTRANSFERASE//DEAMIDATION |
| 11432 | 916 | ARGININE METHYLATION//CARM1//PROTEIN ARGININE METHYLTRANSFERASE |
| 23865 | 278 | TOB//BTG1//BTG2 |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | ARGININE METHYLATION | Author keyword | 68 | 59% | 4% | 76 |
| 2 | CARM1 | Author keyword | 51 | 79% | 2% | 33 |
| 3 | ISOASPARTATE | Author keyword | 50 | 72% | 2% | 39 |
| 4 | PROTEIN ARGININE METHYLTRANSFERASE | Author keyword | 48 | 76% | 2% | 34 |
| 5 | PROTEIN ARGININE METHYLATION | Author keyword | 44 | 77% | 1% | 30 |
| 6 | PRMT1 | Author keyword | 40 | 66% | 2% | 38 |
| 7 | PROTEIN L ISOASPARTYL METHYLTRANSFERASE | Author keyword | 38 | 89% | 1% | 17 |
| 8 | PRMT | Author keyword | 35 | 72% | 1% | 28 |
| 9 | DEAMIDATION | Author keyword | 30 | 27% | 4% | 96 |
| 10 | TOB | Author keyword | 29 | 75% | 1% | 21 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | ARGININE METHYLATION | 68 | 59% | 4% | 76 | Search ARGININE+METHYLATION | Search ARGININE+METHYLATION |
| 2 | CARM1 | 51 | 79% | 2% | 33 | Search CARM1 | Search CARM1 |
| 3 | ISOASPARTATE | 50 | 72% | 2% | 39 | Search ISOASPARTATE | Search ISOASPARTATE |
| 4 | PROTEIN ARGININE METHYLTRANSFERASE | 48 | 76% | 2% | 34 | Search PROTEIN+ARGININE+METHYLTRANSFERASE | Search PROTEIN+ARGININE+METHYLTRANSFERASE |
| 5 | PROTEIN ARGININE METHYLATION | 44 | 77% | 1% | 30 | Search PROTEIN+ARGININE+METHYLATION | Search PROTEIN+ARGININE+METHYLATION |
| 6 | PRMT1 | 40 | 66% | 2% | 38 | Search PRMT1 | Search PRMT1 |
| 7 | PROTEIN L ISOASPARTYL METHYLTRANSFERASE | 38 | 89% | 1% | 17 | Search PROTEIN+L+ISOASPARTYL+METHYLTRANSFERASE | Search PROTEIN+L+ISOASPARTYL+METHYLTRANSFERASE |
| 8 | PRMT | 35 | 72% | 1% | 28 | Search PRMT | Search PRMT |
| 9 | DEAMIDATION | 30 | 27% | 4% | 96 | Search DEAMIDATION | Search DEAMIDATION |
| 10 | TOB | 29 | 75% | 1% | 21 | Search TOB | Search TOB |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | ASPARTYL RESIDUES | 184 | 83% | 5% | 105 |
| 2 | TOB BTG1 FAMILY | 157 | 100% | 2% | 42 |
| 3 | PRMT1 | 80 | 77% | 3% | 54 |
| 4 | CARBOXYL METHYLTRANSFERASE | 73 | 52% | 5% | 99 |
| 5 | SPONTANEOUS DEGRADATION | 70 | 82% | 2% | 41 |
| 6 | N METHYLTRANSFERASE | 68 | 37% | 7% | 145 |
| 7 | DEAMIDATION | 67 | 30% | 9% | 189 |
| 8 | CARM1 | 62 | 85% | 2% | 33 |
| 9 | ASPARAGINYL RESIDUES | 59 | 63% | 3% | 60 |
| 10 | CHEMICAL PATHWAYS | 58 | 63% | 3% | 59 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| Protein Arginine Methylation in Mammals: Who, What, and Why | 2009 | 444 | 117 | 74% |
| Protein arginine methyltransferases and cancer | 2013 | 78 | 189 | 67% |
| Arginine methylation: An emerging regulator of protein function | 2005 | 525 | 88 | 66% |
| Arginine methyltransferases as novel therapeutic targets for breast cancer | 2015 | 2 | 201 | 66% |
| Histone arginine methylation | 2011 | 81 | 108 | 67% |
| The Mammalian Anti-Proliferative BTG/Tob Protein Family | 2010 | 72 | 75 | 77% |
| BTG2: A rising star of tumor suppressors | 2015 | 1 | 54 | 74% |
| The protein arginine methyltransferase family: an update about function, new perspectives and the physiological role in humans | 2009 | 71 | 111 | 85% |
| Minireview: Protein Arginine Methylation of Nonhistone Proteins in Transcriptional Regulation | 2009 | 93 | 84 | 67% |
| State of the Arg: Protein methylation at arginine comes of age | 2001 | 309 | 18 | 94% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | PROGRAM MOL BASIS DIS | 4 | 75% | 0.1% | 3 |
| 2 | TERRY FOX MOL ONCOL GRP | 4 | 16% | 1.0% | 22 |
| 3 | CELL TRANSFORMAT TORAT PROJECT BK21 | 2 | 67% | 0.1% | 2 |
| 4 | OPEN PROT SCI | 2 | 67% | 0.1% | 2 |
| 5 | TERRY FOX MOL ONCOL | 2 | 67% | 0.1% | 2 |
| 6 | BRAIN KOREA CELL TRANSFORMAT TORAT 21 | 2 | 36% | 0.2% | 4 |
| 7 | BION DESIGN | 2 | 43% | 0.1% | 3 |
| 8 | RADIAT LIFE SCI RADIAT MED SCI | 2 | 33% | 0.2% | 4 |
| 9 | BIOCHEM MOL NEUROBIOL | 1 | 31% | 0.2% | 4 |
| 10 | CHEMORADIOTHER Y | 1 | 18% | 0.3% | 7 |
Related classes at same level (level 2) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000013549 | CHROMATIN//EZH2//NUCLEOSOME |
| 2 | 0.0000010891 | FCRN//NEONATAL FC RECEPTOR//FC GAMMA RECEPTORS |
| 3 | 0.0000008117 | ALPHA CRYSTALLIN//CATARACT//LENS |
| 4 | 0.0000007748 | PRE MRNA SPLICING//RNA LOCALIZATION//SPLICEOSOME |
| 5 | 0.0000007215 | HATCHING ENZYME//MEPRIN//ASTACIN FAMILY |
| 6 | 0.0000006896 | GOOSE TYPE LYSOZYME//G TYPE LYSOZYME//I TYPE LYSOZYME |
| 7 | 0.0000006164 | TRIOSEPHOSPHATE ISOMERASE//PYRUVATE KINASE DEFICIENCY//PYRUVATE KINASE |
| 8 | 0.0000006010 | MOLTEN GLOBULE//APOMYOGLOBIN//PROTEIN FORMULATION |
| 9 | 0.0000005993 | NUCLEOLUS//RIBOSOME BIOGENESIS//NUCLEOLAR ORGANIZER REGIONS |
| 10 | 0.0000005711 | FARNESYLTRANSFERASE//PALMITOYLATION//RLIP76 |