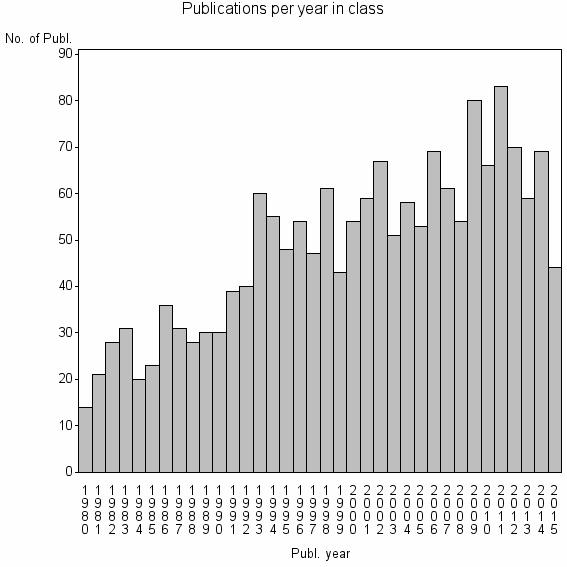
Class information for: |
Basic class information |
| ID | Publications | Average number of references |
Avg. shr. active ref. in WoS |
|---|---|---|---|
| 3134 | 1736 | 43.3 | 77% |
Classes in level above (level 3) |
| ID, lev. above |
Publications | Label for level above |
|---|---|---|
| 96 | 73697 | DNA METHYLATION//FRAGILE X SYNDROME//RETT SYNDROME |
Classes in level below (level 1) |
| ID, lev. below | Publications | Label for level below |
|---|---|---|
| 6846 | 1360 | XIST//X CHROMOSOME INACTIVATION//TSIX |
| 27534 | 186 | TUMOR CLONALITY//CLONAL REMISSION//M27 BETA |
| 33773 | 85 | ABT VIRALE TRANSFORMAT MECHANISMEN//AUSTRALIAN MEDICINAL PLANTS//CELLULAR AUTONOMY |
| 35094 | 53 | HCCR 1//HUMAN CERVICAL CANCER ONCOGENE//HCCR |
| 35123 | 52 | CDK2AP1//DOC 1//P12CDK2 AP1 |
Terms with highest relevance score |
| Rank | Term | Type of term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|---|
| 1 | XIST | Author keyword | 100 | 66% | 5% | 93 |
| 2 | X CHROMOSOME INACTIVATION | Author keyword | 73 | 38% | 9% | 150 |
| 3 | TSIX | Author keyword | 51 | 91% | 1% | 21 |
| 4 | X INACTIVATION | Author keyword | 36 | 28% | 6% | 110 |
| 5 | CDK2AP1 | Author keyword | 20 | 100% | 1% | 9 |
| 6 | X INACTIVAT GRP | Address | 20 | 100% | 1% | 9 |
| 7 | UNITE GENET MOL MURINE | Address | 20 | 52% | 2% | 27 |
| 8 | XIST RNA | Author keyword | 19 | 76% | 1% | 13 |
| 9 | MAMMALIAN DEV EPIGENET GRP | Address | 19 | 74% | 1% | 14 |
| 10 | MED ICSTM | Address | 12 | 86% | 0% | 6 |
Web of Science journal categories |
Author Key Words |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
LCSH search | Wikipedia search |
|---|---|---|---|---|---|---|---|
| 1 | XIST | 100 | 66% | 5% | 93 | Search XIST | Search XIST |
| 2 | X CHROMOSOME INACTIVATION | 73 | 38% | 9% | 150 | Search X+CHROMOSOME+INACTIVATION | Search X+CHROMOSOME+INACTIVATION |
| 3 | TSIX | 51 | 91% | 1% | 21 | Search TSIX | Search TSIX |
| 4 | X INACTIVATION | 36 | 28% | 6% | 110 | Search X+INACTIVATION | Search X+INACTIVATION |
| 5 | CDK2AP1 | 20 | 100% | 1% | 9 | Search CDK2AP1 | Search CDK2AP1 |
| 6 | XIST RNA | 19 | 76% | 1% | 13 | Search XIST+RNA | Search XIST+RNA |
| 7 | X INACTIVE SPECIFIC TRANSCRIPT | 12 | 86% | 0% | 6 | Search X+INACTIVE+SPECIFIC+TRANSCRIPT | Search X+INACTIVE+SPECIFIC+TRANSCRIPT |
| 8 | DOSAGE COMPENSATION | 10 | 19% | 3% | 45 | Search DOSAGE+COMPENSATION | Search DOSAGE+COMPENSATION |
| 9 | HCCR 1 | 6 | 100% | 0% | 4 | Search HCCR+1 | Search HCCR+1 |
| 10 | X INACTIVATION CENTRE | 6 | 100% | 0% | 4 | Search X+INACTIVATION+CENTRE | Search X+INACTIVATION+CENTRE |
Key Words Plus |
| Rank | Web of Science journal category | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | XIST RNA | 123 | 59% | 8% | 138 |
| 2 | TSIX | 114 | 78% | 4% | 75 |
| 3 | CHROMOSOME INACTIVATION | 102 | 38% | 13% | 217 |
| 4 | MEDIATES INITIATION | 76 | 100% | 1% | 24 |
| 5 | TSIX TRANSCRIPTION | 76 | 86% | 2% | 38 |
| 6 | DOC 1 | 56 | 100% | 1% | 19 |
| 7 | REPEAT HYPOTHESIS | 54 | 64% | 3% | 54 |
| 8 | XIST GENE | 49 | 41% | 5% | 92 |
| 9 | FEMALE MAMMALS | 49 | 64% | 3% | 47 |
| 10 | CONTROLLING ELEMENTS | 48 | 74% | 2% | 35 |
Journals |
Reviews |
| Title | Publ. year | Cit. | Active references | % act. ref. to same field |
|---|---|---|---|---|
| X-Inactivation, Imprinting, and Long Noncoding RNAs in Health and Disease | 2013 | 114 | 134 | 49% |
| Regulation of X-chromosome inactivation by the X-inactivation centre | 2011 | 91 | 124 | 91% |
| Gene silencing in X-chromosome inactivation: advances in understanding facultative heterochromatin formation | 2011 | 116 | 119 | 67% |
| X-chromosome inactivation: Counting, choice and initiation | 2001 | 352 | 63 | 78% |
| Lessons from X-chromosome inactivation: long ncRNA as guides and tethers to the epigenome | 2009 | 121 | 109 | 62% |
| X Chromosome Dosage Compensation: How Mammals Keep the Balance | 2008 | 190 | 240 | 63% |
| New and Xisting regulatory mechanisms of X chromosome inactivation | 2012 | 31 | 73 | 84% |
| Dosage compensation in mammals: fine-tuning the expression of the X chromosome | 2006 | 235 | 172 | 70% |
| Noncoding RNAs and Epigenetic Mechanisms During X-Chromosome Inactivation | 2014 | 8 | 114 | 82% |
| Recent advances in X-chromosome inactivation | 2004 | 179 | 48 | 81% |
Address terms |
| Rank | Address term | Relevance score (tfidf) |
Class's shr. of term's tot. occurrences |
Shr. of publ. in class containing term |
Num. of publ. in class |
|---|---|---|---|---|---|
| 1 | X INACTIVAT GRP | 20 | 100% | 0.5% | 9 |
| 2 | UNITE GENET MOL MURINE | 20 | 52% | 1.6% | 27 |
| 3 | MAMMALIAN DEV EPIGENET GRP | 19 | 74% | 0.8% | 14 |
| 4 | MED ICSTM | 12 | 86% | 0.3% | 6 |
| 5 | HOWARD HUGHES MED MOL BIOL GENET | 11 | 78% | 0.4% | 7 |
| 6 | UMR EPIGENET DESTIN CELLULAIRE 7216 | 8 | 100% | 0.3% | 5 |
| 7 | MOL EPIGENET GRP | 7 | 40% | 0.8% | 14 |
| 8 | HOWARD HUGHES MED MOL BIOL | 5 | 63% | 0.3% | 5 |
| 9 | CLIN SCI X INACTIVAT GRP | 4 | 75% | 0.2% | 3 |
| 10 | CLIN SCIICSM | 3 | 100% | 0.2% | 3 |
Related classes at same level (level 2) |
| Rank | Relatedness score | Related classes |
|---|---|---|
| 1 | 0.0000057872 | GENOMIC IMPRINTING//H19//BECKWITH WIEDEMANN SYNDROME |
| 2 | 0.0000026267 | RETT SYNDROME//MECP2//MECP2 GENE |
| 3 | 0.0000025534 | TURNER SYNDROME//SRY//TURNERS SYNDROME |
| 4 | 0.0000019068 | CHROMOSOMA//SYNAPTONEMAL COMPLEX//MEIOSIS |
| 5 | 0.0000018506 | DIOECIOUS PLANT//PLANT DEV GENET//SILENE LATIFOLIA |
| 6 | 0.0000018188 | DNA METHYLATION//EPIGENETICS//HISTONE DEACETYLASE INHIBITOR |
| 7 | 0.0000016314 | G6PD DEFICIENCY//GLUCOSE 6 PHOSPHATE DEHYDROGENASE DEFICIENCY//GLUCOSE 6 PHOSPHATE DEHYDROGENASE |
| 8 | 0.0000016091 | MICROBIAL BIOTECHNOL CELL BIOL//INTERPHASE CHROMATIN//BARLEY CHROMOSOME |
| 9 | 0.0000014972 | EMBRYONAL CARCINOMA//TERATOCARCINOMA//HNT |
| 10 | 0.0000012719 | GEMININ//DNA REPLICATION//CDC6 |